CD4
| CD4, Cluster of differentiation 4, extracellular | |||||||||
|---|---|---|---|---|---|---|---|---|---|
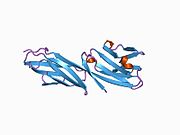
 structure of T cell surface glycoprotein cd4, monoclinic crystal form | |||||||||
| Identifiers | |||||||||
| Symbol | CD4-extrcel | ||||||||
| Pfam | PF09191 | ||||||||
| InterPro | IPR015274 | ||||||||
| SCOP2 | 1cid / SCOPe / SUPFAM | ||||||||
| OPM superfamily | 193 | ||||||||
| OPM protein | 2klu | ||||||||
| CDD | cd07695 | ||||||||
| Membranome | 27 | ||||||||
| |||||||||

In molecular biology, CD4 (cluster of differentiation 4) is a glycoprotein that serves as a co-receptor for the T-cell receptor (TCR). CD4 is found on the surface of immune cells such as helper T cells, monocytes, macrophages, and dendritic cells. It was discovered in the late 1970s and was originally known as leu-3 and T4 (after the OKT4 monoclonal antibody that reacted with it) before being named CD4 in 1984.[5] In humans, the CD4 protein is encoded by the CD4 gene.[6][7]
CD4+ T helper cells are white blood cells that are an essential part of the human immune system. They are often referred to as CD4 cells, T helper cells or T4 cells. They are called helper cells because one of their main roles is to send signals to other types of immune cells, including CD8 killer cells, which then destroy the infectious particle. If CD4 cells become depleted, for example in untreated HIV infection, or following immune suppression prior to a transplant, the body is left vulnerable to a wide range of infections that it would otherwise have been able to fight.
Structure[edit]

Like many cell surface receptors/markers, CD4 is a member of the immunoglobulin superfamily.
It has four immunoglobulin domains (D1 to D4) that are exposed on the extracellular surface of the cell:
- D1 and D3 resemble immunoglobulin variable (IgV) domains.
- D2 and D4 resemble immunoglobulin constant (IgC) domains.
The immunoglobulin variable (IgV) domain of D1 adopts an immunoglobulin-like β-sandwich fold with seven β-strands in 2 β-sheets, in a Greek key topology.[8]
CD4 interacts with the β2-domain of MHC class II molecules through its D1 domain. T cells displaying CD4 molecules (and not CD8) on their surface, therefore, are specific for antigens presented by MHC II and not by MHC class I (they are MHC class II-restricted). MHC class I contains Beta-2 microglobulin.
The short cytoplasmic/intracellular tail (C) of CD4 contains a special sequence of amino acids that allow it to recruit and interact with the tyrosine kinase Lck.
Function[edit]
CD4 is a co-receptor of the T cell receptor (TCR) and assists the latter in communicating with antigen-presenting cells. The TCR complex and CD4 bind to distinct regions of the antigen-presenting MHC class II molecule. The extracellular D1 domain of CD4 binds to the β2 region of MHC class II. The resulting close proximity between the TCR complex and CD4 allows the tyrosine kinase Lck bound to the cytoplasmic tail of CD4[9] to phosphorylate tyrosine residues of immunoreceptor tyrosine activation motifs (ITAMs) on the cytoplasmic domains of CD3[10] to amplify the signal generated by the TCR. Phosphorylated ITAMs on CD3 recruit and activate SH2 domain-containing protein tyrosine kinases (PTK), such as ZAP70, to further mediate downstream signalling through tyrosine phosphorylation. These signals lead to the activation of transcription factors, including NF-κB, NFAT, AP-1, to promote T cell activation.[11]

CD4 is closely related to LAG-3,[12] and together they form an evolutionary conserved system from the level of sharks competing for binding Lck by conserved motifs in their cytoplasmic tails:[13] CD4 through a Cys-X-Cys/His motif[14] and LAG-3 through an immunoreceptor tyrosine-based inhibition motif like (ITIM-like) motif.[13][15][16] LAG-3, which is an inhibitory receptor, is upregulated in activated T cells as a kind of negative feedback loop.
Other interactions[edit]
CD4 has also been shown to interact with SPG21,[17] and Uncoordinated-119 (Unc-119).[18]
Disease[edit]
HIV infection[edit]
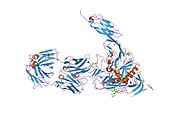
HIV-1 uses CD4 to gain entry into host T-cells and achieves this through its viral envelope protein known as gp120.[19] The binding to CD4 creates a shift in the conformation of gp120 allowing HIV-1 to bind to a co-receptor expressed on the host cell. These co-receptors are chemokine receptors CCR5 or CXCR4. Following a structural change in another viral protein (gp41), HIV inserts a fusion peptide into the host cell that allows the outer membrane of the virus to fuse with the cell membrane.
HIV pathology[edit]
HIV infection leads to a progressive reduction in the number of T cells expressing CD4. Medical professionals refer to the CD4 count to decide when to begin treatment during HIV infection, although recent medical guidelines have changed to recommend treatment at all CD4 counts as soon as HIV is diagnosed. A CD4 count measures the number of T cells expressing CD4. While CD4 counts are not a direct HIV test—e.g. they do not check the presence of viral DNA, or specific antibodies against HIV—they are used to assess the immune system of a patient.[citation needed]
National Institutes of Health guidelines recommend treatment of any HIV-positive individuals, regardless of CD4 count[20] Normal blood values are usually expressed as the number of cells per microliter (μL, or equivalently, cubic millimeter, mm3) of blood, with normal values for CD4 cells being 500–1200 cells/mm3.[21] Patients often undergo treatments when the CD4 counts reach a level of 350 cells per microliter in Europe but usually around 500/μL in the US; people with less than 200 cells per microliter are at high risk of contracting AIDS defined illnesses. Medical professionals also refer to CD4 tests to determine efficacy of treatment.[citation needed]
Viral load testing provides more information about the efficacy for therapy than CD4 counts.[22] For the first 2 years of HIV therapy, CD4 counts may be done every 3–6 months.[22] If a patient's viral load becomes undetectable after 2 years then CD4 counts might not be needed if they are consistently above 500/mm3.[22] If the count remains at 300–500/mm3, then the tests can be done annually.[22] It is not necessary to schedule CD4 counts with viral load tests and the two should be done independently when each is indicated.[22]

Other diseases[edit]
CD4 continues to be expressed in most neoplasms derived from T helper cells. It is therefore possible to use CD4 immunohistochemistry on tissue biopsy samples to identify most forms of peripheral T cell lymphoma and related malignant conditions.[23] The antigen has also been associated with a number of autoimmune diseases such as vitiligo and type I diabetes mellitus.[24]
T-cells play a large part in autoinflammatory diseases.[25] When testing a drug's efficacy or studying diseases, it is helpful to quantify the amount of T-cells on fresh-frozen tissue with CD4+, CD8+, and CD3+ T-cell markers (which stain different markers on a T-cell – giving different results).[26]
See also[edit]
References[edit]
- ^ a b c GRCh38: Ensembl release 89: ENSG00000010610 - Ensembl, May 2017
- ^ a b c GRCm38: Ensembl release 89: ENSMUSG00000023274 - Ensembl, May 2017
- ^ "Human PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ "Mouse PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ Bernard A, Boumsell L, Hill C (1984). "Joint Report of the First International Workshop on Human Leucocyte Differentiation Antigens by the Investigators of the Participating Laboratories". In Bernard A, Boumsell L, Dausset J, Milstein C, Schlossman SF (eds.). Leucocyte typing: human leucocyte differentiation antigens detected by monoclonal antibodies: specification, classification, nomenclature. Berlin: Springer. pp. 45–48. doi:10.1007/978-3-642-68857-7_3. ISBN 0-387-12056-4.
Report on the first international references workshop sponsored by INSERM, WHO and IUIS
- ^ Isobe M, Huebner K, Maddon PJ, Littman DR, Axel R, Croce CM (June 1986). "The gene encoding the T-cell surface protein T4 is located on human chromosome 12". Proceedings of the National Academy of Sciences of the United States of America. 83 (12): 4399–4402. Bibcode:1986PNAS...83.4399I. doi:10.1073/pnas.83.12.4399. PMC 323740. PMID 3086883.
- ^ Ansari-Lari MA, Muzny DM, Lu J, Lu F, Lilley CE, Spanos S, et al. (April 1996). "A gene-rich cluster between the CD4 and triosephosphate isomerase genes at human chromosome 12p13". Genome Research. 6 (4): 314–326. doi:10.1101/gr.6.4.314. PMID 8723724.
- ^ Brady RL, Dodson EJ, Dodson GG, Lange G, Davis SJ, Williams AF, Barclay AN (May 1993). "Crystal structure of domains 3 and 4 of rat CD4: relation to the NH2-terminal domains". Science. 260 (5110): 979–983. Bibcode:1993Sci...260..979B. doi:10.1126/science.8493535. PMID 8493535.
- ^ Rudd CE, Trevillyan JM, Dasgupta JD, Wong LL, Schlossman SF (July 1988). "The CD4 receptor is complexed in detergent lysates to a protein-tyrosine kinase (pp58) from human T lymphocytes". Proceedings of the National Academy of Sciences of the United States of America. 85 (14): 5190–5194. Bibcode:1988PNAS...85.5190R. doi:10.1073/pnas.85.14.5190. PMC 281714. PMID 2455897.
- ^ Barber EK, Dasgupta JD, Schlossman SF, Trevillyan JM, Rudd CE (May 1989). "The CD4 and CD8 antigens are coupled to a protein-tyrosine kinase (p56lck) that phosphorylates the CD3 complex". Proceedings of the National Academy of Sciences of the United States of America. 86 (9): 3277–3281. Bibcode:1989PNAS...86.3277B. doi:10.1073/pnas.86.9.3277. PMC 287114. PMID 2470098.
- ^ Owens JA, Punt J, Stranford SA, Jones PP (2013). Kuby Immunology (7th ed.). New York: W.H. Freeman. pp. 100–101. ISBN 978-14641-3784-6.
- ^ Triebel F, Jitsukawa S, Baixeras E, Roman-Roman S, Genevee C, Viegas-Pequignot E, Hercend T (May 1990). "LAG-3, a novel lymphocyte activation gene closely related to CD4". The Journal of Experimental Medicine. 171 (5): 1393–1405. doi:10.1084/jem.171.5.1393. PMC 2187904. PMID 1692078.
- ^ a b Takizawa F, Hashimoto K, Miyazawa R, Ohta Y, Veríssimo A, Flajnik MF, et al. (2023-12-21). "CD4 and LAG-3 from sharks to humans: related molecules with motifs for opposing functions". Frontiers in Immunology. 14: 1267743. doi:10.3389/fimmu.2023.1267743. PMC 10768021. PMID 38187381.
- ^ Kim PW, Sun ZY, Blacklow SC, Wagner G, Eck MJ (September 2003). "A zinc clasp structure tethers Lck to T cell coreceptors CD4 and CD8". Science. 301 (5640): 1725–1728. Bibcode:2003Sci...301.1725K. doi:10.1126/science.1085643. PMID 14500983. S2CID 32336829.
- ^ Ohashi K, Takizawa F, Tokumaru N, Nakayasu C, Toda H, Fischer U, et al. (August 2010). "A molecule in teleost fish, related with human MHC-encoded G6F, has a cytoplasmic tail with ITAM and marks the surface of thrombocytes and in some fishes also of erythrocytes". Immunogenetics. 62 (8): 543–559. doi:10.1007/s00251-010-0460-1. PMID 20614118. S2CID 12282281.
- ^ Maeda TK, Sugiura D, Okazaki IM, Maruhashi T, Okazaki T (April 2019). "Atypical motifs in the cytoplasmic region of the inhibitory immune co-receptor LAG-3 inhibit T cell activation". The Journal of Biological Chemistry. 294 (15): 6017–6026. doi:10.1074/jbc.RA119.007455. PMC 6463702. PMID 30760527.
- ^ Zeitlmann L, Sirim P, Kremmer E, Kolanus W (March 2001). "Cloning of ACP33 as a novel intracellular ligand of CD4". The Journal of Biological Chemistry. 276 (12): 9123–9132. doi:10.1074/jbc.M009270200. PMID 11113139.
- ^ Gorska MM, Stafford SJ, Cen O, Sur S, Alam R (February 2004). "Unc119, a novel activator of Lck/Fyn, is essential for T cell activation". The Journal of Experimental Medicine. 199 (3): 369–379. doi:10.1084/jem.20030589. PMC 2211793. PMID 14757743.
- ^ Kwong PD, Wyatt R, Robinson J, Sweet RW, Sodroski J, Hendrickson WA (June 1998). "Structure of an HIV gp120 envelope glycoprotein in complex with the CD4 receptor and a neutralizing human antibody". Nature. 393 (6686): 648–659. Bibcode:1998Natur.393..648K. doi:10.1038/31405. PMC 5629912. PMID 9641677.
- ^ "Guidelines for the Use of Antiretroviral Agents in HIV-1-Infected Adults and Adolescents" (PDF). AIDSinfo. U.S. Department of Health & Human Services. 2013-02-13. Archived from the original (PDF) on 2013-10-29. Retrieved 2013-10-24.
- ^ Bofill M, Janossy G, Lee CA, MacDonald-Burns D, Phillips AN, Sabin C, et al. (May 1992). "Laboratory control values for CD4 and CD8 T lymphocytes. Implications for HIV-1 diagnosis". Clinical and Experimental Immunology. 88 (2): 243–252. doi:10.1111/j.1365-2249.1992.tb03068.x. PMC 1554313. PMID 1349272.
- ^ a b c d e HIV Medicine Association (February 2016), "Five Things Physicians and Patients Should Question", Choosing Wisely: an initiative of the ABIM Foundation, HIV Medicine Association, retrieved 9 May 2016
- ^ Cooper K, Leong AS (2003). Manual of diagnostic antibodies for immunohistology. London: Greenwich Medical Media. p. 65. ISBN 1-84110-100-1.
- ^ Zamani M, Tabatabaiefar MA, Mosayyebi S, Mashaghi A, Mansouri P (July 2010). "Possible association of the CD4 gene polymorphism with vitiligo in an Iranian population". Clinical and Experimental Dermatology. 35 (5): 521–524. doi:10.1111/j.1365-2230.2009.03667.x. PMID 19843086. S2CID 10427963.
- ^ Ciccarelli F, De Martinis M, Ginaldi L (2014). "An update on autoinflammatory diseases". Current Medicinal Chemistry. 21 (3): 261–269. doi:10.2174/09298673113206660303. PMC 3905709. PMID 24164192.
- ^ "550280 – BD Biosciences". BD Biosciences. Becton Dickinson.
Further reading[edit]
- Miceli MC, Parnes JR (1993). "Role of CD4 and CD8 in T cell activation and differentiation". Advances in Immunology Volume 53. Vol. 53. pp. 59–122. doi:10.1016/S0065-2776(08)60498-8. ISBN 978-0-12-022453-1. PMID 8512039.
- Geyer M, Fackler OT, Peterlin BM (July 2001). "Structure--function relationships in HIV-1 Nef". EMBO Reports. 2 (7): 580–585. doi:10.1093/embo-reports/kve141. PMC 1083955. PMID 11463741.
- Greenway AL, Holloway G, McPhee DA, Ellis P, Cornall A, Lidman M (April 2003). "HIV-1 Nef control of cell signalling molecules: multiple strategies to promote virus replication". Journal of Biosciences. 28 (3): 323–335. doi:10.1007/BF02970151. PMID 12734410. S2CID 33749514.
- Bénichou S, Benmerah A (January 2003). "[The HIV nef and the Kaposi-sarcoma-associated virus K3/K5 proteins: "parasites"of the endocytosis pathway]". Médecine/Sciences. 19 (1): 100–106. doi:10.1051/medsci/2003191100. PMID 12836198.
- Leavitt SA, SchOn A, Klein JC, Manjappara U, Chaiken IM, Freire E (February 2004). "Interactions of HIV-1 proteins gp120 and Nef with cellular partners define a novel allosteric paradigm". Current Protein & Peptide Science. 5 (1): 1–8. doi:10.2174/1389203043486955. PMID 14965316.
- Tolstrup M, Ostergaard L, Laursen AL, Pedersen SF, Duch M (April 2004). "HIV/SIV escape from immune surveillance: focus on Nef". Current HIV Research. 2 (2): 141–151. doi:10.2174/1570162043484924. PMID 15078178.
- Hout DR, Mulcahy ER, Pacyniak E, Gomez LM, Gomez ML, Stephens EB (July 2004). "Vpu: a multifunctional protein that enhances the pathogenesis of human immunodeficiency virus type 1". Current HIV Research. 2 (3): 255–270. doi:10.2174/1570162043351246. PMID 15279589.
- Joseph AM, Kumar M, Mitra D (January 2005). "Nef: "necessary and enforcing factor" in HIV infection". Current HIV Research. 3 (1): 87–94. doi:10.2174/1570162052773013. PMID 15638726.
- Anderson JL, Hope TJ (April 2004). "HIV accessory proteins and surviving the host cell". Current HIV/AIDS Reports. 1 (1): 47–53. doi:10.1007/s11904-004-0007-x. PMID 16091223. S2CID 34731265.
- Li L, Li HS, Pauza CD, Bukrinsky M, Zhao RY (2006). "Roles of HIV-1 auxiliary proteins in viral pathogenesis and host-pathogen interactions". Cell Research. 15 (11–12): 923–934. doi:10.1038/sj.cr.7290370. PMID 16354571.
- Stove V, Verhasselt B (January 2006). "Modelling thymic HIV-1 Nef effects". Current HIV Research. 4 (1): 57–64. doi:10.2174/157016206775197583. PMID 16454711.
External links[edit]
- CD1+Antigen at the U.S. National Library of Medicine Medical Subject Headings (MeSH)
- Mouse CD Antigen Chart
- Human CD Antigen Chart
- Human Immunodeficiency Virus Glycoprotein 120
- Human CD4 genome location and CD4 gene details page in the UCSC Genome Browser.
- Overview of all the structural information available in the PDB for UniProt: P01730 (T-cell surface glycoprotein CD4) at the PDBe-KB.