Structure and genome of HIV
The genome and proteins of HIV (human immunodeficiency virus) have been the subject of extensive research since the discovery of the virus in 1983.[1][2] "In the search for the causative agent, it was initially believed that the virus was a form of the Human T-cell leukemia virus (HTLV), which was known at the time to affect the human immune system and cause certain leukemias. However, researchers at the Pasteur Institute in Paris isolated a previously unknown and genetically distinct retrovirus in patients with AIDS which was later named HIV."[3] Each virion comprises a viral envelope and associated matrix enclosing a capsid, which itself encloses two copies of the single-stranded RNA genome and several enzymes. The discovery of the virus itself occurred two years following the report of the first major cases of AIDS-associated illnesses.[4][5]
Structure
[edit]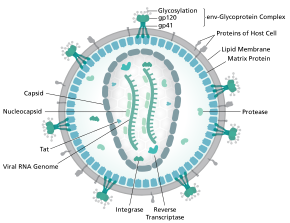


The complete sequence of the HIV-1 genome, extracted from infectious virions, has been solved to single-nucleotide resolution.[6] The HIV genome encodes a small number of viral proteins, invariably establishing cooperative associations among HIV proteins and between HIV and host proteins, to invade host cells and hijack their internal machineries.[7] HIV is different in structure from other retroviruses. The HIV virion is ~100 nm in diameter. Its innermost region consists of a cone-shaped core that includes two copies of the (positive sense) ssRNA genome, the enzymes reverse transcriptase, integrase and protease, some minor proteins, and the major core protein.[8] The genome of human immunodeficiency virus (HIV) encodes 8 viral proteins playing essential roles during the HIV life cycle.[7]
HIV-1 is composed of two copies of noncovalently linked, unspliced, positive-sense single-stranded RNA enclosed by a conical capsid composed of the viral protein p24, typical of lentiviruses.[9][10] The two RNAs are often identical, yet they are not independent, but form a compact dimer within the virion.[11] Several reasons as for why two copies of RNA are packaged rather than just one have been proposed, including probably a combination of these advantages: One advantage is that the two copies of RNA strands are vital in contributing to HIV-1 recombination, which occurs during reverse transcription of viral replication, thus increasing genetic diversity.[11] Another advantage is that having two copies of RNA would allow the reverse transcriptase to switch templates when encountering a break in the viral RNA, thus completing the reverse transcription without loss of genetic information.[11] Yet another reason is that the dimeric nature of the RNA genome of the virus may play a structural role in viral replication.[11] The containment of two copies of single-stranded RNA within a virion but the production of only a single DNA provirus is called pseudodiploidy.[12] The RNA component is 9749 nucleotides long[13][14] and bears a 5’ cap (Gppp), a 3’ poly(A) tail, and many open reading frames (ORFs).[15] Viral structural proteins are encoded by long ORFs, whereas smaller ORFs encode regulators of the viral life cycle: attachment, membrane fusion, replication, and assembly.[15]
The single-strand RNA is tightly bound to p7 nucleocapsid proteins, late assembly protein p6, and enzymes essential to the development of the virion, such as reverse transcriptase and integrase. Lysine tRNA is the primer of the magnesium-dependent reverse transcriptase.[9] The nucleocapsid associates with the genomic RNA (one molecule per hexamer) and protects the RNA from digestion by nucleases. Also enclosed within the virion particle are Vif, Vpr, Nef, and viral protease.[citation needed] The envelope of the virion is formed by a plasma membrane of host cell origin, which is supported by a matrix composed of the viral p17 protein, ensuring the integrity of the virion particle. At the surface of the virion can be found a limited number of the envelope glycoprotein (Env) of HIV, a trimer formed by heterodimers of gp120 and gp41. Env is responsible for binding to its primary host receptor, CD4, and its co-receptor (mainly CCR5 or CXCR4), leading to viral entry into its target cell.[16]
As the only proteins on the surface of the virus, the envelope glycoproteins (gp120 and gp41) are the major targets for HIV vaccine efforts.[17] Over half of the mass of the trimeric envelope spike is N-linked glycans. The density is high as the glycans shield underlying viral protein from neutralisation by antibodies. This is one of the most densely glycosylated molecules known and the density is sufficiently high to prevent the normal maturation process of glycans during biogenesis in the endoplasmic reticulum and Golgi apparatus.[18][19] The majority of the glycans are therefore stalled as immature 'high-mannose' glycans not normally present on secreted or cell surface human glycoproteins.[20] The unusual processing and high density means that almost all broadly neutralising antibodies that have so far been identified (from a subset of patients that have been infected for many months to years) bind to or, are adapted to cope with, these envelope glycans.[21]
The molecular structure of the viral spike has now been determined by X-ray crystallography[22] and cryo-electron microscopy.[23] These advances in structural biology were made possible due to the development of stable recombinant forms of the viral spike by the introduction of an intersubunit disulphide bond and an isoleucine to proline mutation in gp41.[24] The so-called SOSIP trimers not only reproduce the antigenic properties of the native viral spike but also display the same degree of immature glycans as presented on the native virus.[25] Recombinant trimeric viral spikes are promising vaccine candidates as they display less non-neutralising epitopes than recombinant monomeric gp120 which act to suppress the immune response to target epitopes.[26]
Genome organization
[edit]
HIV has several major genes coding for structural proteins that are found in all retroviruses as well as several nonstructural ("accessory") genes unique to HIV.[27] The HIV genome contains nine genes that encode fifteen viral proteins.[28] These are synthesized as polyproteins which produce proteins for virion interior, called Gag, group specific antigen; the viral enzymes (Pol, polymerase) or the glycoproteins of the virion env (envelope).[29] In addition to these, HIV encodes for proteins which have certain regulatory and auxiliary functions as well.[29] HIV-1 has two important regulatory elements: Tat and Rev and few important accessory proteins such as Nef, Vpr, Vif and Vpu which are not essential for replication in certain tissues.[29] The gag gene provides the basic physical infrastructure of the virus, and pol provides the basic mechanism by which retroviruses reproduce, while the others help HIV to enter the host cell and enhance its reproduction. Though they may be altered by mutation, all of these genes except tev exist in all known variants of HIV; see Genetic variability of HIV.[citation needed]
HIV employs a sophisticated system of differential RNA splicing to obtain nine different gene products from a less than 10kb genome.[30] HIV has a 9.2kb unspliced genomic transcript which encodes for gag and pol precursors; a singly spliced, 4.5 kb encoding for env, Vif, Vpr and Vpu and a multiply spliced, 2 kb mRNA encoding for Tat, Rev and Nef.[30]
| Class | Gene name | Primary protein products | Processed protein products |
|---|---|---|---|
| Viral structural proteins | gag | Gag polyprotein | MA, CA, SP1, NC, SP2, P6 |
| pol | Pol polyprotein | RT, RNase H, IN, PR | |
| env | gp160 | gp120, gp41 | |
| Essential regulatory elements | tat | Tat | |
| rev | Rev | ||
| Accessory regulatory proteins | nef | Nef | |
| vpr | Vpr | ||
| vif | Vif | ||
| vpu | Vpu |
Viral structural proteins
[edit]
- gag (group-specific antigen) codes for the precursor gag polyprotein which is processed by viral protease during maturation to MA (matrix protein, p17); CA (capsid protein, p24); SP1 (spacer peptide 1, p2); NC (nucleocapsid protein, p7); SP2 (spacer peptide 2, p1) and P6 protein.[31]
- pol codes for viral enzymes reverse transcriptase (RT) and RNase H, integrase (IN), and HIV protease (PR).[29] HIV protease is required to cleave the precursor Gag polyprotein to produce structural proteins, RT is required to transcribe DNA from RNA template, and IN is necessary to integrate the double-stranded viral DNA into the host genome.[27]
- env (for "envelope") codes for gp160, which is cleaved by a host protease, furin, within the endoplasmic reticulum of the host cell. The post-translational processing produces a surface glycoprotein, gp120 or SU, which attaches to the CD4 receptors present on lymphocytes, and gp41 or TM, which embeds in the viral envelope to enable the virus to attach to and fuse with target cells.[27][31]
Essential regulatory elements
[edit]- tat (HIV trans-activator) plays an important role in regulating the reverse transcription of viral genome RNA, ensuring efficient synthesis of viral mRNAs and regulating the release of virions from infected cells.[29] Tat is expressed as 72-amino acid one-exon Tat as well as the 86–101-amino-acid two-exon Tat, and plays an important role early in HIV infection. Tat (14–15 kDa) binds to the bulged genomic RNA stem-loop secondary structure near the 5' LTR region forming the trans-activation response element (TAR).[9][29]
- rev (regulator of expression of virion proteins): The Rev protein binds to the viral genome via an arginine-rich RNA-binding motif that also acts as a NLS (nuclear localization signals), required for the transport of Rev to the nucleus from cytosol during viral replication.[29] Rev recognizes a complex stem-loop structure of the mRNA env located in the intron separating coding exon of Tat and Rev, known as the HIV Rev response element (RRE).[9][29] Rev is important for the synthesis of major viral proteins and is hence essential for viral replication.[citation needed]
Accessory regulatory proteins
[edit]- vpr (lentivirus protein R): Vpr is a virion-associated, nucleocytoplasmic shuttling regulatory protein.[29] It is believed to play an important role in replication of the virus, specifically, nuclear import of the preintegration complex. Vpr also appears to cause its host cells to arrest their cell cycle in the G2 phase. This arrest activates the host DNA repair machinery which may enable integration of the viral DNA.[9] HIV-2 and SIV encode an additional Vpr related protein called Vpx which functions in association with Vpr.[29]
- vif – Vif is a highly conserved, 23 kDa phosphoprotein important for the infectivity of HIV-1 virions depending on the cell type.[9] HIV-1 has been found to require Vif to synthesize infectious viruses in lymphocytes, macrophages, and certain human cell lines. It does not appear to require Vif for the same process in HeLa cells or COS cells, among others.[29]
- nef – Nef, negative factor, is a N-terminal myristoylated membrane-associated phosphoprotein. It is involved in multiple functions during the replication cycle of the virus. It is believed to play an important role in cell apoptosis and increase virus infectivity.[29]
- vpu (Virus protein U) – Vpu is specific to HIV-1. It is a class I oligomeric integral membrane phosphoprotein with numerous biological functions. Vpu is involved in CD4 degradation involving the ubiquitin proteasome pathway as well as in the successful release of virions from infected cells.[9][29]
- tev: This gene is only present in a few HIV-1 isolates. It is a fusion of parts of the tat, env, and rev genes, and codes for a protein with some of the properties of tat, but little or none of the properties of rev.[32]
RNA secondary structure
[edit]| HIV pol-1 stem loop | |
|---|---|
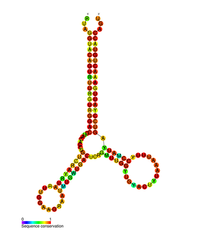 Predicted secondary structure of the HIV pol-1 stem loop | |
| Identifiers | |
| Symbol | pol |
| Rfam | RF01418 |
| Other data | |
| RNA type | Cis-reg |
| PDB structures | PDBe |
Several conserved secondary structure elements have been identified within the HIV RNA genome. The HIV viral RNA structures regulates the progression of reverse transcription. [33] The 5'UTR structure consists of series of stem-loop structures connected by small linkers.[10] These stem-loops (5' to 3') include the trans-activation region (TAR) element, the 5' polyadenylation signal [poly(A)], the PBS, the DIS, the major SD and the ψ hairpin structure located within the 5' end of the genome and the HIV Rev response element (RRE) within the env gene.[10][34][35] Another RNA structure that has been identified is gag stem loop 3 (GSL3), thought to be involved in viral packaging.[36][37] RNA secondary structures have been proposed to affect the HIV life cycle by altering the function of HIV protease and reverse transcriptase, although not all elements identified have been assigned a function.[citation needed]
An RNA secondary structure determined by SHAPE analysis has shown to contain three stem loops and is located between the HIV protease and reverse transcriptase genes. This cis regulatory RNA has been shown to be conserved throughout the HIV family and is thought to influence the viral life cycle.[38]
V3 loop
[edit]The third variable loop or V3 loop is a part or region of the Human Immunodeficiency Virus. The V3 loop of the viron's envelope glycoprotein, gp120, allows it to infect human immune cells by binding to a cytokine receptor on the target human immune cell, such as a CCR5 cell or CXCR4 cell, depending on the strain of HIV.[39] The envelope glycoprotein (Env) gp 120/41 is essential for HIV-1 entry into cells. Env serves as a molecular target of a medicine treating individuals with HIV-1 infection, and a source of immunogen to develop AIDS vaccine. However, the structure of the functional Env trimer has remained elusive.[40]
See also
[edit]References
[edit]- ^ Barré-Sinoussi F, Chermann JC, Rey F, Nugeyre MT, Chamaret S, Gruest J, Dauguet C, Axler-Blin C, Vézinet-Brun F, Rouzioux C, Rozenbaum W, Montagnier L (May 1983). "Isolation of a T-lymphotropic retrovirus from a patient at risk for acquired immune deficiency syndrome (AIDS)". Science. 220 (4599): 868–71. Bibcode:1983Sci...220..868B. doi:10.1126/science.6189183. PMID 6189183. S2CID 390173.
- ^ Gallo RC, Sarin PS, Gelmann EP, Robert-Guroff M, Richardson E, Kalyanaraman VS, Mann D, Sidhu GD, Stahl RE, Zolla-Pazner S, Leibowitch J, Popovic M (May 1983). "Isolation of human T-cell leukemia virus in acquired immune deficiency syndrome (AIDS)". Science. 220 (4599): 865–7. Bibcode:1983Sci...220..865G. doi:10.1126/science.6601823. PMID 6601823.
- ^ Churi C, Ross MW (2015). "Hiv/Aids". In Whelehan P, Bolin A (eds.). The international encyclopedia of human sexuality. Wiley. ISBN 9781405190060. OCLC 949701914.
- ^ Centers for Disease Control (June 1981). "Pneumocystis pneumonia--Los Angeles". MMWR. Morbidity and Mortality Weekly Report. 30 (21): 250–2. PMID 6265753.
- ^ Centers for Disease Control (CDC) (July 1981). "Kaposi's sarcoma and Pneumocystis pneumonia among homosexual men – New York City and California" (PDF). MMWR. Morbidity and Mortality Weekly Report. 30 (25): 305–8. PMID 6789108. Archived from the original on October 22, 2012. Retrieved September 15, 2017.
{{cite journal}}: CS1 maint: unfit URL (link) - ^ Watts JM, Dang KK, Gorelick RJ, Leonard CW, Bess JW, Swanstrom R, Burch CL, Weeks KM (August 2009). "Architecture and secondary structure of an entire HIV-1 RNA genome". Nature. 460 (7256): 711–6. Bibcode:2009Natur.460..711W. doi:10.1038/nature08237. PMC 2724670. PMID 19661910.
- ^ a b Li G, De Clercq E (September 2016). "HIV Genome-Wide Protein Associations: a Review of 30 Years of Research". Microbiology and Molecular Biology Reviews. 80 (3): 679–731. doi:10.1128/MMBR.00065-15. PMC 4981665. PMID 27357278.
- ^ Singleton P, Sainsbury D, eds. (2006). "Hiv". Dictionary of microbiology & molecular biology (3rd ed.). Hoboken, NJ: Wiley. ISBN 9780470035450. OCLC 71223221.
- ^ a b c d e f g Montagnier L (1999). "Human Immunodeficiency Viruses (Retroviridae)". Encyclopedia of Virology (2nd ed.). pp. 763–774.
- ^ a b c Lu K, Heng X, Summers MF (July 2011). "Structural determinants and mechanism of HIV-1 genome packaging". Journal of Molecular Biology. 410 (4): 609–33. doi:10.1016/j.jmb.2011.04.029. PMC 3139105. PMID 21762803.
- ^ a b c d Moore, Michael D.; Hu, Wei Shau (2009). "HIV-1 RNA dimerization: It takes two to tango". AIDS Reviews. 11 (2): 91–102. ISSN 1139-6121. PMC 3056336. PMID 19529749.
- ^ Hwang CK, Svarovskaia ES, Pathak VK (October 2001). "Dynamic copy choice: steady state between murine leukemia virus polymerase and polymerase-dependent RNase H activity determines frequency of in vivo template switching". Proceedings of the National Academy of Sciences of the United States of America. 98 (21): 12209–14. Bibcode:2001PNAS...9812209H. doi:10.1073/pnas.221289898. PMC 59793. PMID 11593039.
- ^ Wain-Hobson S, Sonigo P, Danos O, Cole S, Alizon M (January 1985). "Nucleotide sequence of the AIDS virus, LAV". Cell. 40 (1): 9–17. doi:10.1016/0092-8674(85)90303-4. PMID 2981635. S2CID 33055050.
- ^ Ratner L, Haseltine W, Patarca R, Livak KJ, Starcich B, Josephs SF, Doran ER, Rafalski JA, Whitehorn EA, Baumeister K (1985). "Complete nucleotide sequence of the AIDS virus, HTLV-III". Nature. 313 (6000): 277–84. Bibcode:1985Natur.313..277R. doi:10.1038/313277a0. PMID 2578615. S2CID 4316242.
- ^ a b Castelli JC, Levy A (2002). "HIV (Human Immunodeficiency Virus)". Encyclopedia of Cancer. Vol. 2 (2nd ed.). pp. 407–415.
- ^ Checkly MA, Freed EO (22 July 2011). "HIV-1 envelope glycoprotein biosynthesis, trafficking, and incorporation". Journal of Molecular Biology. 410 (4): 582–608. doi:10.1016/j.jmb.2011.04.042. PMC 3139147. PMID 21762802.
- ^ National Institute of Health (June 17, 1998). "Crystal structure of key HIV protein reveals new prevention, treatment targets" (Press release). Archived from the original on February 19, 2006. Retrieved September 14, 2006.
- ^ Behrens AJ, Vasiljevic S, Pritchard LK, Harvey DJ, Andev RS, Krumm SA, Struwe WB, Cupo A, Kumar A, Zitzmann N, Seabright GE, Kramer HB, Spencer DI, Royle L, Lee JH, Klasse PJ, Burton DR, Wilson IA, Ward AB, Sanders RW, Moore JP, Doores KJ, Crispin M (March 2016). "Composition and Antigenic Effects of Individual Glycan Sites of a Trimeric HIV-1 Envelope Glycoprotein". Cell Reports. 14 (11): 2695–706. doi:10.1016/j.celrep.2016.02.058. PMC 4805854. PMID 26972002.
- ^ Pritchard LK, Spencer DI, Royle L, Bonomelli C, Seabright GE, Behrens AJ, Kulp DW, Menis S, Krumm SA, Dunlop DC, Crispin DJ, Bowden TA, Scanlan CN, Ward AB, Schief WR, Doores KJ, Crispin M (June 2015). "Glycan clustering stabilizes the mannose patch of HIV-1 and preserves vulnerability to broadly neutralizing antibodies". Nature Communications. 6: 7479. Bibcode:2015NatCo...6.7479P. doi:10.1038/ncomms8479. PMC 4500839. PMID 26105115.
- ^ Pritchard LK, Harvey DJ, Bonomelli C, Crispin M, Doores KJ (September 2015). "Cell- and Protein-Directed Glycosylation of Native Cleaved HIV-1 Envelope". Journal of Virology. 89 (17): 8932–44. doi:10.1128/JVI.01190-15. PMC 4524065. PMID 26085151.
- ^ Crispin M, Doores KJ (April 2015). "Targeting host-derived glycans on enveloped viruses for antibody-based vaccine design". Current Opinion in Virology. Viral pathogenesis • Preventive and therapeutic vaccines. 11: 63–9. doi:10.1016/j.coviro.2015.02.002. PMC 4827424. PMID 25747313.
- ^ Julien JP, Cupo A, Sok D, Stanfield RL, Lyumkis D, Deller MC, Klasse PJ, Burton DR, Sanders RW, Moore JP, Ward AB, Wilson IA (December 2013). "Crystal structure of a soluble cleaved HIV-1 envelope trimer". Science. 342 (6165): 1477–83. Bibcode:2013Sci...342.1477J. doi:10.1126/science.1245625. PMC 3886632. PMID 24179159.
- ^ Lyumkis D, Julien JP, de Val N, Cupo A, Potter CS, Klasse PJ, Burton DR, Sanders RW, Moore JP, Carragher B, Wilson IA, Ward AB (December 2013). "Cryo-EM structure of a fully glycosylated soluble cleaved HIV-1 envelope trimer". Science. 342 (6165): 1484–90. Bibcode:2013Sci...342.1484L. doi:10.1126/science.1245627. PMC 3954647. PMID 24179160.
- ^ Sanders RW, Derking R, Cupo A, Julien JP, Yasmeen A, de Val N, Kim HJ, Blattner C, de la Peña AT, Korzun J, Golabek M, de Los Reyes K, Ketas TJ, van Gils MJ, King CR, Wilson IA, Ward AB, Klasse PJ, Moore JP (September 2013). "A next-generation cleaved, soluble HIV-1 Env trimer, BG505 SOSIP.664 gp140, expresses multiple epitopes for broadly neutralizing but not non-neutralizing antibodies". PLOS Pathogens. 9 (9): e1003618. doi:10.1371/journal.ppat.1003618. PMC 3777863. PMID 24068931.
- ^ Pritchard LK, Vasiljevic S, Ozorowski G, Seabright GE, Cupo A, Ringe R, Kim HJ, Sanders RW, Doores KJ, Burton DR, Wilson IA, Ward AB, Moore JP, Crispin M (June 2015). "Structural Constraints Determine the Glycosylation of HIV-1 Envelope Trimers". Cell Reports. 11 (10): 1604–13. doi:10.1016/j.celrep.2015.05.017. PMC 4555872. PMID 26051934.
- ^ de Taeye SW, Ozorowski G, Torrents de la Peña A, Guttman M, Julien JP, van den Kerkhof TL, Burger JA, Pritchard LK, Pugach P, Yasmeen A, Crampton J, Hu J, Bontjer I, Torres JL, Arendt H, DeStefano J, Koff WC, Schuitemaker H, Eggink D, Berkhout B, Dean H, LaBranche C, Crotty S, Crispin M, Montefiori DC, Klasse PJ, Lee KK, Moore JP, Wilson IA, Ward AB, Sanders RW (December 2015). "Immunogenicity of Stabilized HIV-1 Envelope Trimers with Reduced Exposure of Non-neutralizing Epitopes". Cell. 163 (7): 1702–15. doi:10.1016/j.cell.2015.11.056. PMC 4732737. PMID 26687358.
- ^ a b c Mushahwar IK (2007). "Human Immunodeficiency Viruses: Molecular Virology, pathogenesis, diagnosis and treatment". Perspectives in Medical Virology. 13: 75–87. doi:10.1016/S0168-7069(06)13005-0. ISBN 9780444520739.
- ^ Li G, Piampongsant S, Faria NR, Voet A, Pineda-Peña AC, Khouri R, Lemey P, Vandamme AM, Theys K (February 2015). "An integrated map of HIV genome-wide variation from a population perspective". Retrovirology. 12 (1): 18. doi:10.1186/s12977-015-0148-6. PMC 4358901. PMID 25808207.
- ^ a b c d e f g h i j k l m Votteler J, Schubert U (2008). "Human Immunodeficiency Viruses: Molecular Biology". Encyclopedia of Virology (3rd ed.). pp. 517–525.
- ^ a b Feinberg Mark B, Greene Warner C (1992). "Molecular Insights into human immunodeficiency virus type1 pathogenesis". Current Opinion in Immunology. 4 (4): 466–474. doi:10.1016/s0952-7915(06)80041-5. PMID 1356348.
- ^ a b King Steven R (1994). "HIV: Virology and Mechanisms of disease". Annals of Emergency Medicine. 24 (3): 443–449. doi:10.1016/s0196-0644(94)70181-4. PMID 7915889.
- ^ Benko DM, Schwartz S, Pavlakis GN, Felber BK (June 1990). "A novel human immunodeficiency virus type 1 protein, tev, shares sequences with tat, env, and rev proteins". Journal of Virology. 64 (6): 2505–18. doi:10.1128/JVI.64.6.2505-2518.1990. PMC 249426. PMID 2186172.
- ^ Krupkin M, Jackson LN, Ha B, Puglisi EV (Dec 2020). "Advances in understanding the initiation of HIV-1 reverse transcription". Curr Opin Struct Biol. 65: 175–183. doi:10.1016/j.sbi.2020.07.005. PMC 9973426. PMID 32916568. S2CID 221636459.
- ^ Berkhout B (January 1992). "Structural features in TAR RNA of human and simian immunodeficiency viruses: a phylogenetic analysis". Nucleic Acids Research. 20 (1): 27–31. doi:10.1093/nar/20.1.27. PMC 310321. PMID 1738599.
- ^ Paillart JC, Skripkin E, Ehresmann B, Ehresmann C, Marquet R (February 2002). "In vitro evidence for a long range pseudoknot in the 5'-untranslated and matrix coding regions of HIV-1 genomic RNA". The Journal of Biological Chemistry. 277 (8): 5995–6004. doi:10.1074/jbc.M108972200. PMID 11744696.
- ^ Damgaard CK, Andersen ES, Knudsen B, Gorodkin J, Kjems J (February 2004). "RNA interactions in the 5' region of the HIV-1 genome". Journal of Molecular Biology. 336 (2): 369–79. doi:10.1016/j.jmb.2003.12.010. PMID 14757051.
- ^ Rong L, Russell RS, Hu J, Laughrea M, Wainberg MA, Liang C (September 2003). "Deletion of stem-loop 3 is compensated by second-site mutations within the Gag protein of human immunodeficiency virus type 1". Virology. 314 (1): 221–8. doi:10.1016/S0042-6822(03)00405-7. PMID 14517075.
- ^ Wang Q, Barr I, Guo F, Lee C (December 2008). "Evidence of a novel RNA secondary structure in the coding region of HIV-1 pol gene". RNA. 14 (12): 2478–88. doi:10.1261/rna.1252608. PMC 2590956. PMID 18974280.
- ^ "The interactions of the gp120 V3 loop of different HIV-1 strains with the potent anti-HIV human monoclonal antibody 447-52D". Weizmann Institute of Science: Department of Structural Biology. Archived from the original on 2007-07-18. Retrieved 2017-04-18.
- ^ Takeda S, Takizawa M, Miyauchi K, Urano E, Fujino M, Murakami T, Murakami T, Komano J (June 2016). "Conformational properties of the third variable loop of HIV-1AD8 envelope glycoprotein in the liganded conditions". Biochemical and Biophysical Research Communications. 475 (1): 113–8. doi:10.1016/j.bbrc.2016.05.051. PMID 27178216.
External links
[edit]- Rfam entry for HIV pol-1 stem loop
- 3D model of the complete HIV1 virion
- Liu J, Wright ER, Winkler H (2010). "3D Visualization of HIV Virions by Cryoelectron Tomography". Cryo-EM, Part C: Analyses, Interpretation, and Case studies. Methods in Enzymology. Vol. 483. pp. 267–90. doi:10.1016/S0076-6879(10)83014-9. ISBN 9780123849939. PMC 3056484. PMID 20888479.
