Signal transduction
Signal transduction is the process by which a chemical or physical signal is transmitted through a cell as a series of molecular events. Proteins responsible for detecting stimuli are generally termed receptors, although in some cases the term sensor is used.[1] The changes elicited by ligand binding (or signal sensing) in a receptor give rise to a biochemical cascade, which is a chain of biochemical events known as a signaling pathway.
When signaling pathways interact with one another they form networks, which allow cellular responses to be coordinated, often by combinatorial signaling events.[2] At the molecular level, such responses include changes in the transcription or translation of genes, and post-translational and conformational changes in proteins, as well as changes in their location. These molecular events are the basic mechanisms controlling cell growth, proliferation, metabolism and many other processes.[3] In multicellular organisms, signal transduction pathways regulate cell communication in a wide variety of ways.
Each component (or node) of a signaling pathway is classified according to the role it plays with respect to the initial stimulus. Ligands are termed first messengers, while receptors are the signal transducers, which then activate primary effectors. Such effectors are typically proteins and are often linked to second messengers, which can activate secondary effectors, and so on. Depending on the efficiency of the nodes, a signal can be amplified (a concept known as signal gain), so that one signaling molecule can generate a response involving hundreds to millions of molecules.[4] As with other signals, the transduction of biological signals is characterised by delay, noise, signal feedback and feedforward and interference, which can range from negligible to pathological.[5] With the advent of computational biology, the analysis of signaling pathways and networks has become an essential tool to understand cellular functions and disease, including signaling rewiring mechanisms underlying responses to acquired drug resistance.[6]

Stimuli
[edit]
The basis for signal transduction is the transformation of a certain stimulus into a biochemical signal. The nature of such stimuli can vary widely, ranging from extracellular cues, such as the presence of EGF, to intracellular events, such as the DNA damage resulting from replicative telomere attrition.[7] Traditionally, signals that reach the central nervous system are classified as senses. These are transmitted from neuron to neuron in a process called synaptic transmission. Many other intercellular signal relay mechanisms exist in multicellular organisms, such as those that govern embryonic development.[8]
Ligands
[edit]The majority of signal transduction pathways involve the binding of signaling molecules, known as ligands, to receptors that trigger events inside the cell. The binding of a signaling molecule with a receptor causes a change in the conformation of the receptor, known as receptor activation. Most ligands are soluble molecules from the extracellular medium which bind to cell surface receptors. These include growth factors, cytokines and neurotransmitters. Components of the extracellular matrix such as fibronectin and hyaluronan can also bind to such receptors (integrins and CD44, respectively). In addition, some molecules such as steroid hormones are lipid-soluble and thus cross the plasma membrane to reach cytoplasmic or nuclear receptors.[9] In the case of steroid hormone receptors, their stimulation leads to binding to the promoter region of steroid-responsive genes.[10]
Not all classifications of signaling molecules take into account the molecular nature of each class member. For example, odorants belong to a wide range of molecular classes,[11] as do neurotransmitters, which range in size from small molecules such as dopamine[12] to neuropeptides such as endorphins.[13] Moreover, some molecules may fit into more than one class, e.g. epinephrine is a neurotransmitter when secreted by the central nervous system and a hormone when secreted by the adrenal medulla.
Some receptors such as HER2 are capable of ligand-independent activation when overexpressed or mutated. This leads to constitutive activation of the pathway, which may or may not be overturned by compensation mechanisms. In the case of HER2, which acts as a dimerization partner of other EGFRs, constitutive activation leads to hyperproliferation and cancer.[14]
Mechanical forces
[edit]The prevalence of basement membranes in the tissues of Eumetazoans means that most cell types require attachment to survive. This requirement has led to the development of complex mechanotransduction pathways, allowing cells to sense the stiffness of the substratum. Such signaling is mainly orchestrated in focal adhesions, regions where the integrin-bound actin cytoskeleton detects changes and transmits them downstream through YAP1.[15] Calcium-dependent cell adhesion molecules such as cadherins and selectins can also mediate mechanotransduction.[16] Specialised forms of mechanotransduction within the nervous system are responsible for mechanosensation: hearing, touch, proprioception and balance.[17]
Osmolarity
[edit]Cellular and systemic control of osmotic pressure (the difference in osmolarity between the cytosol and the extracellular medium) is critical for homeostasis. There are three ways in which cells can detect osmotic stimuli: as changes in macromolecular crowding, ionic strength, and changes in the properties of the plasma membrane or cytoskeleton (the latter being a form of mechanotransduction).[18] These changes are detected by proteins known as osmosensors or osmoreceptors. In humans, the best characterised osmosensors are transient receptor potential channels present in the primary cilium of human cells.[18][19] In yeast, the HOG pathway has been extensively characterised.[20]
Temperature
[edit]The sensing of temperature in cells is known as thermoception and is primarily mediated by transient receptor potential channels.[21] Additionally, animal cells contain a conserved mechanism to prevent high temperatures from causing cellular damage, the heat-shock response. Such response is triggered when high temperatures cause the dissociation of inactive HSF1 from complexes with heat shock proteins Hsp40/Hsp70 and Hsp90. With help from the ncRNA hsr1, HSF1 then trimerizes, becoming active and upregulating the expression of its target genes.[22] Many other thermosensory mechanisms exist in both prokaryotes and eukaryotes.[21]
Light
[edit]In mammals, light controls the sense of sight and the circadian clock by activating light-sensitive proteins in photoreceptor cells in the eye's retina. In the case of vision, light is detected by rhodopsin in rod and cone cells.[23] In the case of the circadian clock, a different photopigment, melanopsin, is responsible for detecting light in intrinsically photosensitive retinal ganglion cells.[24]
Receptors
[edit]Receptors can be roughly divided into two major classes: intracellular and extracellular receptors.
Extracellular receptors
[edit]Extracellular receptors are integral transmembrane proteins and make up most receptors. They span the plasma membrane of the cell, with one part of the receptor on the outside of the cell and the other on the inside. Signal transduction occurs as a result of a ligand binding to the outside region of the receptor (the ligand does not pass through the membrane). Ligand-receptor binding induces a change in the conformation of the inside part of the receptor, a process sometimes called "receptor activation".[25] This results in either the activation of an enzyme domain of the receptor or the exposure of a binding site for other intracellular signaling proteins within the cell, eventually propagating the signal through the cytoplasm.
In eukaryotic cells, most intracellular proteins activated by a ligand/receptor interaction possess an enzymatic activity; examples include tyrosine kinase and phosphatases. Often such enzymes are covalently linked to the receptor. Some of them create second messengers such as cyclic AMP and IP3, the latter controlling the release of intracellular calcium stores into the cytoplasm. Other activated proteins interact with adaptor proteins that facilitate signaling protein interactions and coordination of signaling complexes necessary to respond to a particular stimulus. Enzymes and adaptor proteins are both responsive to various second messenger molecules.
Many adaptor proteins and enzymes activated as part of signal transduction possess specialized protein domains that bind to specific secondary messenger molecules. For example, calcium ions bind to the EF hand domains of calmodulin, allowing it to bind and activate calmodulin-dependent kinase. PIP3 and other phosphoinositides do the same thing to the Pleckstrin homology domains of proteins such as the kinase protein AKT.
G protein–coupled receptors
[edit]G protein–coupled receptors (GPCRs) are a family of integral transmembrane proteins that possess seven transmembrane domains and are linked to a heterotrimeric G protein. With nearly 800 members, this is the largest family of membrane proteins and receptors in mammals. Counting all animal species, they add up to over 5000.[26] Mammalian GPCRs are classified into 5 major families: rhodopsin-like, secretin-like, metabotropic glutamate, adhesion and frizzled/smoothened, with a few GPCR groups being difficult to classify due to low sequence similarity, e.g. vomeronasal receptors.[26] Other classes exist in eukaryotes, such as the Dictyostelium cyclic AMP receptors and fungal mating pheromone receptors.[26]
Signal transduction by a GPCR begins with an inactive G protein coupled to the receptor; the G protein exists as a heterotrimer consisting of Gα, Gβ, and Gγ subunits.[27] Once the GPCR recognizes a ligand, the conformation of the receptor changes to activate the G protein, causing Gα to bind a molecule of GTP and dissociate from the other two G-protein subunits. The dissociation exposes sites on the subunits that can interact with other molecules.[28] The activated G protein subunits detach from the receptor and initiate signaling from many downstream effector proteins such as phospholipases and ion channels, the latter permitting the release of second messenger molecules.[29] The total strength of signal amplification by a GPCR is determined by the lifetimes of the ligand-receptor complex and receptor-effector protein complex and the deactivation time of the activated receptor and effectors through intrinsic enzymatic activity; e.g. via protein kinase phosphorylation or b-arrestin-dependent internalization.
A study was conducted where a point mutation was inserted into the gene encoding the chemokine receptor CXCR2; mutated cells underwent a malignant transformation due to the expression of CXCR2 in an active conformation despite the absence of chemokine-binding. This meant that chemokine receptors can contribute to cancer development.[30]
Tyrosine, Ser/Thr and Histidine-specific protein kinases
[edit]Receptor tyrosine kinases (RTKs) are transmembrane proteins with an intracellular kinase domain and an extracellular domain that binds ligands; examples include growth factor receptors such as the insulin receptor.[31] To perform signal transduction, RTKs need to form dimers in the plasma membrane;[32] the dimer is stabilized by ligands binding to the receptor. The interaction between the cytoplasmic domains stimulates the autophosphorylation of tyrosine residues within the intracellular kinase domains of the RTKs, causing conformational changes. Subsequent to this, the receptors' kinase domains are activated, initiating phosphorylation signaling cascades of downstream cytoplasmic molecules that facilitate various cellular processes such as cell differentiation and metabolism.[31] Many Ser/Thr and dual-specificity protein kinases are important for signal transduction, either acting downstream of [receptor tyrosine kinases], or as membrane-embedded or cell-soluble versions in their own right. The process of signal transduction involves around 560 known protein kinases and pseudokinases, encoded by the human kinome[33][34]
As is the case with GPCRs, proteins that bind GTP play a major role in signal transduction from the activated RTK into the cell. In this case, the G proteins are members of the Ras, Rho, and Raf families, referred to collectively as small G proteins. They act as molecular switches usually tethered to membranes by isoprenyl groups linked to their carboxyl ends. Upon activation, they assign proteins to specific membrane subdomains where they participate in signaling. Activated RTKs in turn activate small G proteins that activate guanine nucleotide exchange factors such as SOS1. Once activated, these exchange factors can activate more small G proteins, thus amplifying the receptor's initial signal. The mutation of certain RTK genes, as with that of GPCRs, can result in the expression of receptors that exist in a constitutively activated state; such mutated genes may act as oncogenes.[35]
Histidine-specific protein kinases are structurally distinct from other protein kinases and are found in prokaryotes, fungi, and plants as part of a two-component signal transduction mechanism: a phosphate group from ATP is first added to a histidine residue within the kinase, then transferred to an aspartate residue on a receiver domain on a different protein or the kinase itself, thus activating the aspartate residue.[36]
Integrins
[edit]
Integrins are produced by a wide variety of cells; they play a role in cell attachment to other cells and the extracellular matrix and in the transduction of signals from extracellular matrix components such as fibronectin and collagen. Ligand binding to the extracellular domain of integrins changes the protein's conformation, clustering it at the cell membrane to initiate signal transduction. Integrins lack kinase activity; hence, integrin-mediated signal transduction is achieved through a variety of intracellular protein kinases and adaptor molecules, the main coordinator being integrin-linked kinase.[37] As shown in the adjacent picture, cooperative integrin-RTK signaling determines the timing of cellular survival, apoptosis, proliferation, and differentiation.
Important differences exist between integrin-signaling in circulating blood cells and non-circulating cells such as epithelial cells; integrins of circulating cells are normally inactive. For example, cell membrane integrins on circulating leukocytes are maintained in an inactive state to avoid epithelial cell attachment; they are activated only in response to stimuli such as those received at the site of an inflammatory response. In a similar manner, integrins at the cell membrane of circulating platelets are normally kept inactive to avoid thrombosis. Epithelial cells (which are non-circulating) normally have active integrins at their cell membrane, helping maintain their stable adhesion to underlying stromal cells that provide signals to maintain normal functioning.[38]
In plants, there are no bona fide integrin receptors identified to date; nevertheless, several integrin-like proteins were proposed based on structural homology with the metazoan receptors.[39] Plants contain integrin-linked kinases that are very similar in their primary structure with the animal ILKs. In the experimental model plant Arabidopsis thaliana, one of the integrin-linked kinase genes, ILK1, has been shown to be a critical element in the plant immune response to signal molecules from bacterial pathogens and plant sensitivity to salt and osmotic stress.[40] ILK1 protein interacts with the high-affinity potassium transporter HAK5 and with the calcium sensor CML9.[40][41]
Toll-like receptors
[edit]When activated, toll-like receptors (TLRs) take adapter molecules within the cytoplasm of cells in order to propagate a signal. Four adaptor molecules are known to be involved in signaling, which are Myd88, TIRAP, TRIF, and TRAM.[42][43][44] These adapters activate other intracellular molecules such as IRAK1, IRAK4, TBK1, and IKKi that amplify the signal, eventually leading to the induction or suppression of genes that cause certain responses. Thousands of genes are activated by TLR signaling, implying that this method constitutes an important gateway for gene modulation.
Ligand-gated ion channels
[edit]A ligand-gated ion channel, upon binding with a ligand, changes conformation to open a channel in the cell membrane through which ions relaying signals can pass. An example of this mechanism is found in the receiving cell of a neural synapse. The influx of ions that occurs in response to the opening of these channels induces action potentials, such as those that travel along nerves, by depolarizing the membrane of post-synaptic cells, resulting in the opening of voltage-gated ion channels.
An example of an ion allowed into the cell during a ligand-gated ion channel opening is Ca2+; it acts as a second messenger initiating signal transduction cascades and altering the physiology of the responding cell. This results in amplification of the synapse response between synaptic cells by remodelling the dendritic spines involved in the synapse.
Intracellular receptors
[edit]Intracellular receptors, such as nuclear receptors and cytoplasmic receptors, are soluble proteins localized within their respective areas. The typical ligands for nuclear receptors are non-polar hormones like the steroid hormones testosterone and progesterone and derivatives of vitamins A and D. To initiate signal transduction, the ligand must pass through the plasma membrane by passive diffusion. On binding with the receptor, the ligands pass through the nuclear membrane into the nucleus, altering gene expression.
Activated nuclear receptors attach to the DNA at receptor-specific hormone-responsive element (HRE) sequences, located in the promoter region of the genes activated by the hormone-receptor complex. Due to their enabling gene transcription, they are alternatively called inductors of gene expression. All hormones that act by regulation of gene expression have two consequences in their mechanism of action; their effects are produced after a characteristically long period of time and their effects persist for another long period of time, even after their concentration has been reduced to zero, due to a relatively slow turnover of most enzymes and proteins that would either deactivate or terminate ligand binding onto the receptor.
Nucleic receptors have DNA-binding domains containing zinc fingers and a ligand-binding domain; the zinc fingers stabilize DNA binding by holding its phosphate backbone. DNA sequences that match the receptor are usually hexameric repeats of any kind; the sequences are similar but their orientation and distance differentiate them. The ligand-binding domain is additionally responsible for dimerization of nucleic receptors prior to binding and providing structures for transactivation used for communication with the translational apparatus.
Steroid receptors are a subclass of nuclear receptors located primarily within the cytosol. In the absence of steroids, they associate in an aporeceptor complex containing chaperone or heatshock proteins (HSPs). The HSPs are necessary to activate the receptor by assisting the protein to fold in a way such that the signal sequence enabling its passage into the nucleus is accessible. Steroid receptors, on the other hand, may be repressive on gene expression when their transactivation domain is hidden. Receptor activity can be enhanced by phosphorylation of serine residues at their N-terminal as a result of another signal transduction pathway, a process called crosstalk.
Retinoic acid receptors are another subset of nuclear receptors. They can be activated by an endocrine-synthesized ligand that entered the cell by diffusion, a ligand synthesised from a precursor like retinol brought to the cell through the bloodstream or a completely intracellularly synthesised ligand like prostaglandin. These receptors are located in the nucleus and are not accompanied by HSPs. They repress their gene by binding to their specific DNA sequence when no ligand binds to them, and vice versa.
Certain intracellular receptors of the immune system are cytoplasmic receptors; recently identified NOD-like receptors (NLRs) reside in the cytoplasm of some eukaryotic cells and interact with ligands using a leucine-rich repeat (LRR) motif similar to TLRs. Some of these molecules like NOD2 interact with RIP2 kinase that activates NF-κB signaling, whereas others like NALP3 interact with inflammatory caspases and initiate processing of particular cytokines like interleukin-1β.[45][46]
Second messengers
[edit]First messengers are the signaling molecules (hormones, neurotransmitters, and paracrine/autocrine agents) that reach the cell from the extracellular fluid and bind to their specific receptors. Second messengers are the substances that enter the cytoplasm and act within the cell to trigger a response. In essence, second messengers serve as chemical relays from the plasma membrane to the cytoplasm, thus carrying out intracellular signal transduction.
Calcium
[edit]The release of calcium ions from the endoplasmic reticulum into the cytosol results in its binding to signaling proteins that are then activated; it is then sequestered in the smooth endoplasmic reticulum[47] and the mitochondria. Two combined receptor/ion channel proteins control the transport of calcium: the InsP3-receptor that transports calcium upon interaction with inositol triphosphate on its cytosolic side; and the ryanodine receptor named after the alkaloid ryanodine, similar to the InsP3 receptor but having a feedback mechanism that releases more calcium upon binding with it. The nature of calcium in the cytosol means that it is active for only a very short time, meaning its free state concentration is very low and is mostly bound to organelle molecules like calreticulin when inactive.
Calcium is used in many processes including muscle contraction, neurotransmitter release from nerve endings, and cell migration. The three main pathways that lead to its activation are GPCR pathways, RTK pathways, and gated ion channels; it regulates proteins either directly or by binding to an enzyme.
Lipid messengers
[edit]Lipophilic second messenger molecules are derived from lipids residing in cellular membranes; enzymes stimulated by activated receptors activate the lipids by modifying them. Examples include diacylglycerol and ceramide, the former required for the activation of protein kinase C.
Nitric oxide
[edit]Nitric oxide (NO) acts as a second messenger because it is a free radical that can diffuse through the plasma membrane and affect nearby cells. It is synthesised from arginine and oxygen by the NO synthase and works through activation of soluble guanylyl cyclase, which when activated produces another second messenger, cGMP. NO can also act through covalent modification of proteins or their metal co-factors; some have a redox mechanism and are reversible. It is toxic in high concentrations and causes damage during stroke, but is the cause of many other functions like the relaxation of blood vessels, apoptosis, and penile erections.
Redox signaling
[edit]In addition to nitric oxide, other electronically activated species are also signal-transducing agents in a process called redox signaling. Examples include superoxide, hydrogen peroxide, carbon monoxide, and hydrogen sulfide. Redox signaling also includes active modulation of electronic flows in semiconductive biological macromolecules.[48]
Cellular responses
[edit]Gene activations[49] and metabolism alterations[50] are examples of cellular responses to extracellular stimulation that require signal transduction. Gene activation leads to further cellular effects, since the products of responding genes include instigators of activation; transcription factors produced as a result of a signal transduction cascade can activate even more genes. Hence, an initial stimulus can trigger the expression of a large number of genes, leading to physiological events like the increased uptake of glucose from the blood stream[50] and the migration of neutrophils to sites of infection. The set of genes and their activation order to certain stimuli is referred to as a genetic program.[51]
Mammalian cells require stimulation for cell division and survival; in the absence of growth factor, apoptosis ensues. Such requirements for extracellular stimulation are necessary for controlling cell behavior in unicellular and multicellular organisms; signal transduction pathways are perceived to be so central to biological processes that a large number of diseases are attributed to their dysregulation. Three basic signals determine cellular growth:
- Stimulatory (growth factors)
- Transcription dependent response
For example, steroids act directly as transcription factor (gives slow response, as transcription factor must bind DNA, which needs to be transcribed. Produced mRNA needs to be translated, and the produced protein/peptide can undergo posttranslational modification (PTM)) - Transcription independent response
For example, epidermal growth factor (EGF) binds the epidermal growth factor receptor (EGFR), which causes dimerization and autophosphorylation of the EGFR, which in turn activates the intracellular signaling pathway .[52]
- Transcription dependent response
- Inhibitory (cell-cell contact)
- Permissive (cell-matrix interactions)
The combination of these signals is integrated into altered cytoplasmic machinery which leads to altered cell behaviour.
Major pathways
[edit]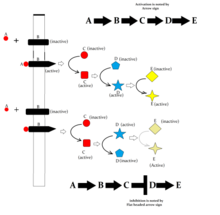
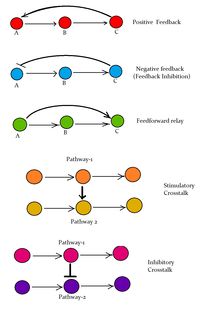
Following are some major signaling pathways, demonstrating how ligands binding to their receptors can affect second messengers and eventually result in altered cellular responses.
- MAPK/ERK pathway: A pathway that couples intracellular responses to the binding of growth factors to cell surface receptors. This pathway is very complex and includes many protein components.[53] In many cell types, activation of this pathway promotes cell division, and many forms of cancer are associated with aberrations in it.[54]
- cAMP-dependent pathway: In humans, cAMP works by activating protein kinase A (PKA, cAMP-dependent protein kinase) (see picture), and, thus, further effects depend mainly on cAMP-dependent protein kinase, which vary based on the type of cell.
- IP3/DAG pathway: PLC cleaves the phospholipid phosphatidylinositol 4,5-bisphosphate (PIP2), yielding diacyl glycerol (DAG) and inositol 1,4,5-triphosphate (IP3). DAG remains bound to the membrane, and IP3 is released as a soluble structure into the cytosol. IP3 then diffuses through the cytosol to bind to IP3 receptors, particular calcium channels in the endoplasmic reticulum (ER). These channels are specific to calcium and allow the passage of only calcium to move through. This causes the cytosolic concentration of Calcium to increase, causing a cascade of intracellular changes and activity.[55] In addition, calcium and DAG together works to activate PKC, which goes on to phosphorylate other molecules, leading to altered cellular activity. End-effects include taste, manic depression, tumor promotion, etc.[55]
History
[edit]
The earliest notion of signal transduction can be traced back to 1855, when Claude Bernard proposed that ductless glands such as the spleen, the thyroid and adrenal glands, were responsible for the release of "internal secretions" with physiological effects.[56] Bernard's "secretions" were later named "hormones" by Ernest Starling in 1905.[57] Together with William Bayliss, Starling had discovered secretin in 1902.[56] Although many other hormones, most notably insulin, were discovered in the following years, the mechanisms remained largely unknown.
The discovery of nerve growth factor by Rita Levi-Montalcini in 1954, and epidermal growth factor by Stanley Cohen in 1962, led to more detailed insights into the molecular basis of cell signaling, in particular growth factors.[58] Their work, together with Earl Wilbur Sutherland's discovery of cyclic AMP in 1956, prompted the redefinition of endocrine signaling to include only signaling from glands, while the terms autocrine and paracrine began to be used.[59] Sutherland was awarded the 1971 Nobel Prize in Physiology or Medicine, while Levi-Montalcini and Cohen shared it in 1986.
In 1970, Martin Rodbell examined the effects of glucagon on a rat's liver cell membrane receptor. He noted that guanosine triphosphate disassociated glucagon from this receptor and stimulated the G-protein, which strongly influenced the cell's metabolism. Thus, he deduced that the G-protein is a transducer that accepts glucagon molecules and affects the cell.[60] For this, he shared the 1994 Nobel Prize in Physiology or Medicine with Alfred G. Gilman. Thus, the characterization of RTKs and GPCRs led to the formulation of the concept of "signal transduction", a word first used in 1972.[61] Some early articles used the terms signal transmission and sensory transduction.[62][63] In 2007, a total of 48,377 scientific papers—including 11,211 review papers—were published on the subject. The term first appeared in a paper's title in 1979.[64][65] Widespread use of the term has been traced to a 1980 review article by Rodbell:[60][66] Research papers focusing on signal transduction first appeared in large numbers in the late 1980s and early 1990s.[46]
Signal transduction in Immunology
[edit]The purpose of this section is to briefly describe some developments in immunology in the 1960s and 1970s, relevant to the initial stages of transmembrane signal transduction, and how they impacted our understanding of immunology, and ultimately of other areas of cell biology.
The relevant events begin with the sequencing of myeloma protein light chains, which are found in abundance in the urine of individuals with multiple myeloma. Biochemical experiments revealed that these so-called Bence Jones proteins consisted of 2 discrete domains –one that varied from one molecule to the next (the V domain) and one that did not (the Fc domain or the Fragment crystallizable region).[67] An analysis of multiple V region sequences by Wu and Kabat [68] identified locations within the V region that were hypervariable and which, they hypothesized, combined in the folded protein to form the antigen recognition site. Thus, within a relatively short time a plausible model was developed for the molecular basis of immunological specificity, and for mediation of biological function through the Fc domain. Crystallization of an IgG molecule soon followed [69] ) confirming the inferences based on sequencing, and providing an understanding of immunological specificity at the highest level of resolution.
The biological significance of these developments was encapsulated in the theory of clonal selection[70] which holds that a B cell has on its surface immunoglobulin receptors whose antigen-binding site is identical to that of antibodies that are secreted by the cell when it encounters an antigen, and more specifically a particular B cell clone secretes antibodies with identical sequences. The final piece of the story, the Fluid mosaic model of the plasma membrane provided all the ingredients for a new model for the initiation of signal transduction; viz, receptor dimerization.
The first hints of this were obtained by Becker et al [71] who demonstrated that the extent to which human basophils—for which bivalent Immunoglobulin E (IgE) functions as a surface receptor – degranulate, depends on the concentration of anti IgE antibodies to which they are exposed, and results in a redistribution of surface molecules, which is absent when monovalent ligand is used. The latter observation was consistent with earlier findings by Fanger et al.[72] These observations tied a biological response to events and structural details of molecules on the cell surface. A preponderance of evidence soon developed that receptor dimerization initiates responses (reviewed in [73]) in a variety of cell types, including B cells.
Such observations led to a number of theoretical (mathematical) developments. The first of these was a simple model proposed by Bell [74] which resolved an apparent paradox: clustering forms stable networks; i.e. binding is essentially irreversible, whereas the affinities of antibodies secreted by B cells increase as the immune response progresses. A theory of the dynamics of cell surface clustering on lymphocyte membranes was developed by DeLisi and Perelson [75] who found the size distribution of clusters as a function of time, and its dependence on the affinity and valence of the ligand. Subsequent theories for basophils and mast cells were developed by Goldstein and Sobotka and their collaborators,[76][77] all aimed at the analysis of dose-response patterns of immune cells and their biological correlates.[78] For a recent review of clustering in immunological systems see.[79]
Ligand binding to cell surface receptors is also critical to motility, a phenomenon that is best understood in single-celled organisms. An example is a detection and response to concentration gradients by bacteria [80]-–the classic mathematical theory appearing in.[81] A recent account can be found in [82]
See also
[edit]- Adaptor protein
- Scaffold protein
- Biosemiotics
- Cell signaling
- Gene regulatory network
- Hormonal imprinting
- Metabolic pathway
- Protein–protein interaction
- Two-component regulatory system
References
[edit]- ^ Bradshaw RA, Dennis EA, eds. (2010). Handbook of Cell Signaling (2nd ed.). Amsterdam, Netherlands: Academic Press. ISBN 9780123741455.
- ^ Papin JA, Hunter T, Palsson BO, Subramaniam S (February 2005). "Reconstruction of cellular signalling networks and analysis of their properties". Nature Reviews. Molecular Cell Biology. 6 (2): 99–111. doi:10.1038/nrm1570. PMID 15654321. S2CID 3065483.
- ^ Krauss G (2008). Biochemistry of Signal Transduction and Regulation. Wiley-VCH. p. 15. ISBN 978-3527313976.
- ^ Reece J, Campbell N (2002). Biology. San Francisco: Benjamin Cummings. ISBN 978-0-8053-6624-2.
- ^ Kolch W, Halasz M, Granovskaya M, Kholodenko BN (September 2015). "The dynamic control of signal transduction networks in cancer cells". Nature Reviews. Cancer. 15 (9): 515–27. doi:10.1038/nrc3983. PMID 26289315. S2CID 35252401.
- ^ Bago R, Sommer E, Castel P, Crafter C, Bailey FP, Shpiro N, Baselga J, Cross D, Eyers PA, Alessi DR (2016) The hVps34-SGK3 pathway alleviates sustained PI3K/Akt inhibition by stimulating mTORC1 and tumour growth. EMBO Journal 35:1902-22
- ^ Smogorzewska A, de Lange T (August 2002). "Different telomere damage signaling pathways in human and mouse cells". The EMBO Journal. 21 (16): 4338–48. doi:10.1093/emboj/cdf433. PMC 126171. PMID 12169636.
- ^ Lawrence PA, Levine M (April 2006). "Mosaic and regulative development: two faces of one coin". Current Biology. 16 (7): R236-9. doi:10.1016/j.cub.2006.03.016. PMID 16581495.
- ^ Beato M, Chávez S, Truss M (April 1996). "Transcriptional regulation by steroid hormones". Steroids. 61 (4): 240–51. doi:10.1016/0039-128X(96)00030-X. PMID 8733009. S2CID 20654561.
- ^ Hammes SR (March 2003). "The further redefining of steroid-mediated signaling". Proceedings of the National Academy of Sciences of the United States of America. 100 (5): 2168–70. Bibcode:2003PNAS..100.2168H. doi:10.1073/pnas.0530224100. PMC 151311. PMID 12606724.
- ^ Ronnett GV, Moon C (2002). "G proteins and olfactory signal transduction". Annual Review of Physiology. 64 (1): 189–222. doi:10.1146/annurev.physiol.64.082701.102219. PMID 11826268.
- ^ Missale C, Nash SR, Robinson SW, Jaber M, Caron MG (January 1998). "Dopamine receptors: from structure to function". Physiological Reviews. 78 (1): 189–225. doi:10.1152/physrev.1998.78.1.189. PMID 9457173.
- ^ Goldstein A (September 1976). "Opioid peptides endorphins in pituitary and brain". Science. 193 (4258): 1081–6. Bibcode:1976Sci...193.1081G. doi:10.1126/science.959823. PMID 959823.
- ^ Koboldt DC, Fulton RS, McLellan MD, Schmidt H, Kalicki-Veizer J, McMichael JF, et al. (The Cancer Genome Atlas Network) (October 2012). "Comprehensive molecular portraits of human breast tumours". Nature. 490 (7418): 61–70. Bibcode:2012Natur.490...61T. doi:10.1038/nature11412. PMC 3465532. PMID 23000897.
- ^ Dupont S, Morsut L, Aragona M, Enzo E, Giulitti S, Cordenonsi M, et al. (June 2011). "Role of YAP/TAZ in mechanotransduction". Nature. 474 (7350): 179–83. doi:10.1038/nature10137. PMID 21654799. S2CID 205225137.
- ^ Ingber DE (May 2006). "Cellular mechanotransduction: putting all the pieces together again". FASEB Journal. 20 (7): 811–27. doi:10.1096/fj.05-5424rev. PMID 16675838. S2CID 21267494.
- ^ Kung C (August 2005). "A possible unifying principle for mechanosensation". Nature. 436 (7051): 647–54. Bibcode:2005Natur.436..647K. doi:10.1038/nature03896. PMID 16079835. S2CID 4374012.
- ^ a b Pedersen SF, Kapus A, Hoffmann EK (September 2011). "Osmosensory mechanisms in cellular and systemic volume regulation". Journal of the American Society of Nephrology. 22 (9): 1587–97. doi:10.1681/ASN.2010121284. PMID 21852585.
- ^ Verbalis JG (December 2007). "How does the brain sense osmolality?". Journal of the American Society of Nephrology. 18 (12): 3056–9. doi:10.1681/ASN.2007070825. PMID 18003769.
- ^ Hohmann S (June 2002). "Osmotic stress signaling and osmoadaptation in yeasts". Microbiology and Molecular Biology Reviews. 66 (2): 300–72. doi:10.1128/MMBR.66.2.300-372.2002. PMC 120784. PMID 12040128.
- ^ a b Sengupta P, Garrity P (April 2013). "Sensing temperature". Current Biology. 23 (8): R304-7. doi:10.1016/j.cub.2013.03.009. PMC 3685181. PMID 23618661.
- ^ Shamovsky I, Ivannikov M, Kandel ES, Gershon D, Nudler E (March 2006). "RNA-mediated response to heat shock in mammalian cells". Nature. 440 (7083): 556–60. Bibcode:2006Natur.440..556S. doi:10.1038/nature04518. PMID 16554823. S2CID 4311262.
- ^ Burns ME, Arshavsky VY (November 2005). "Beyond counting photons: trials and trends in vertebrate visual transduction". Neuron. 48 (3): 387–401. doi:10.1016/j.neuron.2005.10.014. PMID 16269358.
- ^ Berson DM (August 2007). "Phototransduction in ganglion-cell photoreceptors". Pflügers Archiv. 454 (5): 849–55. doi:10.1007/s00424-007-0242-2. PMID 17351786.
- ^ "A molecular model for receptor activation" (PDF).
- ^ a b c Fredriksson R, Schiöth HB (May 2005). "The repertoire of G-protein-coupled receptors in fully sequenced genomes". Molecular Pharmacology. 67 (5): 1414–25. doi:10.1124/mol.104.009001. PMID 15687224. S2CID 7938806.
- ^ Qin K, Dong C, Wu G, Lambert NA (August 2011). "Inactive-state preassembly of G(q)-coupled receptors and G(q) heterotrimers". Nature Chemical Biology. 7 (10): 740–7. doi:10.1038/nchembio.642. PMC 3177959. PMID 21873996.
- ^ Berg JM, Tymoczko JL, Stryer L, Clarke ND (2002). Biochemistry. San Francisco: W.H. Freeman. ISBN 978-0-7167-4954-7.
- ^ Yang W, Xia S (2006). "Mechanisms of regulation and function of G-protein-coupled receptor kinases". World J Gastroenterol. 12 (48): 7753–7. doi:10.3748/wjg.v12.i48.7753. PMC 4087537. PMID 17203515.
- ^ Burger M, Burger JA, Hoch RC, Oades Z, Takamori H, Schraufstatter IU (August 1999). "Point mutation causing constitutive signaling of CXCR2 leads to transforming activity similar to Kaposi's sarcoma herpesvirus-G protein-coupled receptor". Journal of Immunology. 163 (4): 2017–22. doi:10.4049/jimmunol.163.4.2017. PMID 10438939. S2CID 45743458.
- ^ a b Li E, Hristova K (May 2006). "Role of receptor tyrosine kinase transmembrane domains in cell signaling and human pathologies". Biochemistry. 45 (20): 6241–51. doi:10.1021/bi060609y. PMC 4301406. PMID 16700535.
- ^ Schlessinger J (November 1988). "Signal transduction by allosteric receptor oligomerization". Trends in Biochemical Sciences. 13 (11): 443–7. doi:10.1016/0968-0004(88)90219-8. PMID 3075366.
- ^ Manning G, Whyte DB, Martinez R, Hunter T, Sudarsanam S (December 2002). "The protein kinase complement of the human genome". Science. 298 (5600): 1912–34. Bibcode:2002Sci...298.1912M. doi:10.1126/science.1075762. PMID 12471243. S2CID 26554314.
- ^ Reiterer V, Eyers PA, Farhan H (September 2014). "Day of the dead: pseudokinases and pseudophosphatases in physiology and disease". Trends in Cell Biology. 24 (9): 489–505. doi:10.1016/j.tcb.2014.03.008. PMID 24818526.
- ^ Roskoski R (June 2004). "The ErbB/HER receptor protein-tyrosine kinases and cancer". Biochemical and Biophysical Research Communications. 319 (1): 1–11. doi:10.1016/j.bbrc.2004.04.150. PMID 15158434.
- ^ Wolanin PM, Thomason PA, Stock JB (September 2002). "Histidine protein kinases: key signal transducers outside the animal kingdom". Genome Biology. 3 (10): REVIEWS3013. doi:10.1186/gb-2002-3-10-reviews3013. PMC 244915. PMID 12372152.
- ^ a b Hehlgans S, Haase M, Cordes N (January 2007). "Signalling via integrins: implications for cell survival and anticancer strategies". Biochimica et Biophysica Acta (BBA) - Reviews on Cancer. 1775 (1): 163–80. doi:10.1016/j.bbcan.2006.09.001. PMID 17084981.
- ^ Gilcrease MZ (March 2007). "Integrin signaling in epithelial cells". Cancer Letters. 247 (1): 1–25. doi:10.1016/j.canlet.2006.03.031. PMID 16725254.
- ^ Knepper C, Savory EA, Day B (May 2011). "Arabidopsis NDR1 is an integrin-like protein with a role in fluid loss and plasma membrane-cell wall adhesion". Plant Physiology. 156 (1): 286–300. doi:10.1104/pp.110.169656. PMC 3091050. PMID 21398259.
- ^ a b Brauer EK, Ahsan N, Dale R, Kato N, Coluccio AE, Piñeros MA, et al. (June 2016). "The Raf-like Kinase ILK1 and the High Affinity K+ Transporter HAK5 Are Required for Innate Immunity and Abiotic Stress Response". Plant Physiology. 171 (2): 1470–84. doi:10.1104/pp.16.00035. PMC 4902592. PMID 27208244.
- ^ Popescu SC, Popescu GV, Bachan S, Zhang Z, Seay M, Gerstein M, et al. (March 2007). "Differential binding of calmodulin-related proteins to their targets revealed through high-density Arabidopsis protein microarrays". Proceedings of the National Academy of Sciences of the United States of America. 104 (11): 4730–5. Bibcode:2007PNAS..104.4730P. doi:10.1073/pnas.0611615104. PMC 1838668. PMID 17360592.
- ^ Yamamoto M, Sato S, Hemmi H, Hoshino K, Kaisho T, Sanjo H, et al. (August 2003). "Role of adaptor TRIF in the MyD88-independent toll-like receptor signaling pathway". Science. 301 (5633): 640–3. Bibcode:2003Sci...301..640Y. doi:10.1126/science.1087262. PMID 12855817. S2CID 19276476.
- ^ Yamamoto M, Sato S, Hemmi H, Uematsu S, Hoshino K, Kaisho T, et al. (November 2003). "TRAM is specifically involved in the Toll-like receptor 4-mediated MyD88-independent signaling pathway". Nature Immunology. 4 (11): 1144–50. doi:10.1038/ni986. PMID 14556004. S2CID 13016860.
- ^ Yamamoto M, Sato S, Hemmi H, Sanjo H, Uematsu S, Kaisho T, et al. (November 2002). "Essential role for TIRAP in activation of the signalling cascade shared by TLR2 and TLR4". Nature. 420 (6913): 324–9. Bibcode:2002Natur.420..324Y. doi:10.1038/nature01182. PMID 12447441. S2CID 16163262.
- ^ Delbridge LM, O'Riordan MX (February 2007). "Innate recognition of intracellular bacteria". Current Opinion in Immunology. 19 (1): 10–6. doi:10.1016/j.coi.2006.11.005. PMID 17126540.
- ^ a b Vander AJ, Sherman J, Luciano D (1998). Human Physiology (7th ed.). McGraw-Hill. pp. 159–60. ISBN 978-0-07-067065-5.
- ^ Wilson CH, Ali ES, Scrimgeour N, Martin AM, Hua J, Tallis GA, et al. (March 2015). "Steatosis inhibits liver cell store-operated Ca²⁺ entry and reduces ER Ca²⁺ through a protein kinase C-dependent mechanism". The Biochemical Journal. 466 (2): 379–90. doi:10.1042/bj20140881. PMID 25422863.
- ^ Forman HJ (November 2009). "Signal transduction and reactive species". Free Radical Biology & Medicine. 47 (9): 1237–8. doi:10.1016/j.freeradbiomed.2009.09.002. PMID 19735727.
- ^ Lalli E, Sassone-Corsi P (July 1994). "Signal transduction and gene regulation: the nuclear response to cAMP". The Journal of Biological Chemistry. 269 (26): 17359–62. doi:10.1016/S0021-9258(17)32442-0. PMID 8021233.
- ^ a b Rosen OM (September 1987). "After insulin binds". Science. 237 (4821): 1452–8. Bibcode:1987Sci...237.1452R. doi:10.1126/science.2442814. PMID 2442814.
- ^ Massagué J, Gomis RR (May 2006). "The logic of TGFbeta signaling". FEBS Letters. 580 (12): 2811–20. doi:10.1016/j.febslet.2006.04.033. PMID 16678165.
- ^ Sako Y, Minoghchi S, Yanagida T (March 2000). "Single-molecule imaging of EGFR signalling on the surface of living cells". Nature Cell Biology. 2 (3): 168–72. doi:10.1038/35004044. PMID 10707088. S2CID 25515586.
- ^ Orton RJ, Sturm OE, Vyshemirsky V, Calder M, Gilbert DR, Kolch W (December 2005). "Computational modelling of the receptor-tyrosine-kinase-activated MAPK pathway". The Biochemical Journal. 392 (Pt 2): 249–61. doi:10.1042/BJ20050908. PMC 1316260. PMID 16293107.
- ^ Vogelstein B, Kinzler KW (August 2004). "Cancer genes and the pathways they control". Nature Medicine. 10 (8): 789–99. doi:10.1038/nm1087. PMID 15286780. S2CID 205383514.
- ^ a b Alberts B, Lewis J, Raff M, Roberts K, Walter P (2002). Molecular biology of the cell (4th ed.). New York: Garland Science. ISBN 978-0-8153-3218-3.
- ^ a b Bradshaw & Dennis (2010) p. 1.
- ^ Tata JR (June 2005). "One hundred years of hormones". EMBO Reports. 6 (6): 490–6. doi:10.1038/sj.embor.7400444. PMC 1369102. PMID 15940278.
- ^ Cowan WM (March 2001). "Viktor Hamburger and Rita Levi-Montalcini: the path to the discovery of nerve growth factor". Annual Review of Neuroscience. 24 (1): 551–600. doi:10.1146/annurev.neuro.24.1.551. PMID 11283321. S2CID 6747529.
- ^ Bradshaw & Dennis (2010) p. 2.
- ^ a b Rodbell M (March 1980). "The role of hormone receptors and GTP-regulatory proteins in membrane transduction". Nature. 284 (5751): 17–22. Bibcode:1980Natur.284...17R. doi:10.1038/284017a0. PMID 6101906. S2CID 5650340.
- ^ Rensing L (1972). "Periodic geophysical and biological signals as Zeitgeber and exogenous inducers in animal organisms". International Journal of Biometeorology. 16 Suppl: 113–25. PMID 4621276.
- ^ Tonndorf J (September 1975). "Davis-1961 revisited. Signal transmission in the cochlear hair cell-nerve junction". Archives of Otolaryngology. 101 (9): 528–35. doi:10.1001/archotol.1975.00780380006002. PMID 169771.
- ^ Ashcroft SJ, Crossley JR, Crossley PC (March 1976). "The effect of N-acylglucosamines on the biosynthesis and secretion of insulin in the rat". The Biochemical Journal. 154 (3): 701–7. doi:10.1042/bj1540701. PMC 1172772. PMID 782447.
- ^ Hildebrand E (April 1977). "What does Halobacterium tell us about photoreception?". Biophysics of Structure and Mechanism. 3 (1): 69–77. doi:10.1007/BF00536457. PMID 857951. S2CID 62775788.
- ^ Kenny JJ, Martínez-Maza O, Fehniger T, Ashman RF (April 1979). "Lipid synthesis: an indicator of antigen-induced signal transduction in antigen-binding cells". Journal of Immunology. 122 (4): 1278–84. doi:10.4049/jimmunol.122.4.1278. PMID 376714. S2CID 29355685.
- ^ Gomperts BD, Kramer IM, Tatham PE (2002). Signal transduction. Academic Press. ISBN 978-0-12-289631-6.
- ^ Steiner, L A (1996) Immunoglobulin evolution, 30 years on. Glycobiology 6 , 649-656
- ^ Wu, T T, Kabat, E A (1970) An analysis of the sequences of the variable regions of Bence Jones proteins and myeloma light chains and their implications for antibody complementarity. J. Exp. Med. 132: 211-250
- ^ Sarma, V R, Silverton, E W, Davies, D R, Terry W D (1971) The three-dimensional structure at 6 A resolution of a human gamma G1 immunoglobulin molecule, J Biol. Chem. 246 (11) 3752- 9
- ^ Burnet, F M (1976) A modification of Jerne's theory of antibody production using the concept of clonal selection. CA: A Cancer Journal for Clinicians 26 (2) 119–21
- ^ Becker, K E, Ishizaka, T, Metzger, H, Ishizaka, K and Grimley, P M (1973) Surface IgE on Human Basophils during histamine release. J Exp med, 138, 394-408
- ^ Fanger, M W, Hart, D A, Wells, J V, and Nisonoff, A J (1970) Requirement for cross-linkage in the stimulation of transformation of rabbit peripheral lymphocytes by antiglobulin reagents J. Immun., 105, 1484 - 92
- ^ Klemm J D, Schreiber S L, Crabtree G R (1998) Ann. Rev. Immunol. Dimerization as a regulatory mechanism in signal transduction 16: 569-592
- ^ Bell, G I (1974) Model for the binding of multivalent antigens to cells, Nature Lond. 248, 430
- ^ DeLisi, C and Perelson A (1976). The kinetics of aggregation phenomena, J. theor. Biol. 62, 159-210
- ^ Dembo, M and Goldstein, B (1978) Theory of equilibrium binding of symmetric bivalent haptens to cell surface antibody: application to histamine release from basophils. The Journal of Immunology 121 (1), 345-353
- ^ Sobotka, A.K. Dembo, M, Goldstein, B and Lichtenstein, L M, (1979) Antigen-specific desensitization of human basophils The Journal of Immunology, 122 (2) 511-517
- ^ Kagey-Sobotka, A, Dembo, M, Goldstein, B, Metzger, H and Lichtenstein, L M (1981) Qualitative characteristics of histamine release from human basophils by covalently cross-linked IgE. The Journal of Immunology 127 (6), 2285-2291
- ^ How does T cell receptor clustering impact signal transduction? Jesse Goyette, Daniel J. Nieves, Yuanqing Ma, Katharina Gaus Journal of Cell Science 2019 132:jcs226423 doi:10.1242/jcs.226423 Published 11 February 2019
- ^ MacNab, R., and D. E. Koshland, Jr. (1972). The gradient-sensing mechanism in bacterial chemotaxis. Proc. Natl. Acad. Sci. U.S.A. 69:2509-2512
- ^ Berg, H C and Purcell, E M (1977) Physics of chemoreception, Biophys. J 20(2):193-219
- ^ Kirsten Jung, Florian Fabiani, Elisabeth Hoyer, and Jürgen Lassak 2018 Bacterial transmembrane signaling systems and their engineering for biosensing Open Biol. Apr; 8(4): 180023
External links
[edit]- Netpath - A curated resource of signal transduction pathways in humans Archived 2012-09-20 at the Wayback Machine
- Signal Transduction - The Virtual Library of Biochemistry, Molecular Biology and Cell Biology
- TRANSPATH(R) - A database about signal transduction pathways
- Science's STKE - Signal Transduction Knowledge Environment, from the journal Science, published by AAAS.
- Signal+Transduction at the U.S. National Library of Medicine Medical Subject Headings (MeSH)
- UCSD-Nature Signaling Gateway Archived 2013-02-12 at the Wayback Machine, from Nature Publishing Group
- LitInspector Archived 2019-05-11 at the Wayback Machine - Signal transduction pathway mining in PubMed abstracts
- Huaxian Chen, et al. A Cell Based Immunocytochemical Assay For Monitoring Kinase Signaling Pathways And Drug Efficacy (PDF) Archived 2012-02-22 at the Wayback Machine Analytical Biochemistry 338 (2005) 136-142
- www.Redoxsignaling.com
- Signaling PAthway Database Archived 2012-09-17 at the Wayback Machine - Kyushu University
- Cell cycle - Homo sapiens (human) Archived 2012-10-23 at the Wayback Machine - KEGG PATHWAY [1]
- Pathway Interaction Database - NCI
- Literature-curated human signaling network, the largest human signaling network database
