Peptidomimetic
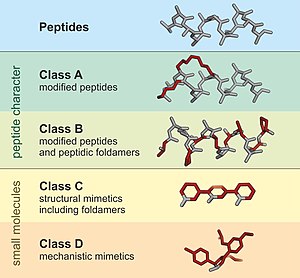
A peptidomimetic is a small protein-like chain designed to mimic a peptide.[1][2] They typically arise either from modification of an existing peptide, or by designing similar systems that mimic peptides, such as peptoids and β-peptides. Irrespective of the approach, the altered chemical structure is designed to advantageously adjust the molecular properties such as stability or biological activity. This can have a role in the development of drug-like compounds from existing peptides. Peptidomimetics can be prepared by cyclization of linear peptides or coupling of stable unnatural amino acids.[3] These modifications involve changes to the peptide that will not occur naturally (such as altered backbones and the incorporation of nonnatural amino acids). Unnatural amino acids can be generated from their native analogs via modifications such as amine alkylation, side chain substitution, structural bond extension cyclization, and isosteric replacements within the amino acid backbone.[3] Based on their similarity with the precursor peptide, peptidomimetics can be grouped into four classes (A – D) where A features the most and D the least similarities. Classes A and B involve peptide-like scaffolds, while classes C and D include small molecules (Figure 1).[1]
Class A peptidomimetics[edit]
This group includes modified peptides that are mainly composed of proteogenic amino acids thereby closely resembling a natural peptide binding epitope.[1] Introduced modifications usually aim to increase the stability of the peptide, its affintiy for a desired binding partner, oral availablity or cell permeability. The design of class A peptidomimetics often involves macrocyclization strategies as for example in stapled peptides.
Class B peptidomimetics[edit]
This class of peptidomimetics encompasses peptides with a large number of non-natural amino acids, major backbone modifications or larger non-natural building fragments that resemble the conformation of a particular peptide binding motif.[1] Examples involve D-peptide and peptidic foldamers such as beta-peptides.
Class C peptidomimetics[edit]
These structural mimetics include molecules that are highly modified when compared to their parent peptide sequence.[4] Usually, a small-molecular scaffold is appyled to project groups in analogy to the bioactive conformation of a peptide.
Class D peptidomimetics[edit]
These mechanistic mimetics do not directly recapitulate the side chains or conformation of a peptide but mimic its mode-of-action.[1] Class D peptidomimetics can be directly designed from a small peptide sequence or identified the screening of compound libraries. For example, Nirmatrelvir is an orally-active small molecule drug derived from lufotrelvir, a modified L-peptide.[5]
-
Lufotrelvir (active form has phosphate group cleaved off)
-
Nirmatrelvir
Uses and examples[edit]
The use of peptides as drugs has some disadvantages because of their bioavailability and biostability. Rapid degradation, poor oral availability, difficult transportation through cell membranes, nonselective receptor binding, and challenging multistep preparation are the major limitations of peptides as active pharmaceutical ingredients.[3] Therefore, small protein-like chains called peptidomimetics could be designed and used to mimic native analogs and conceivably exhibit better pharmacological properties.[3] Many peptidomimetics are utilized as FDA-approved drugs, such as Romidepsin (Istodax), Atazanavir (Reyataz), Saquinavir (Invirase), Oktreotid (Sandostatin), Lanreotide (Somatuline), Plecanatide (Trulance), Ximelagatran (Exanta), Etelcalcetide (Parsabiv), and Bortezomib (Velcade).
Peptidomimetic approaches have been utilized to design small molecules that selectively target cancer cells, an approach known as targeted chemotherapy, by inducing programmed cell death by a process called apoptosis. The following two examples mimic proteins involved in key Protein–protein interactions that reactivate the apoptotic pathway in cancer but do so by distinct mechanisms.[6]
In 2004, Walensky and co-workers reported a stabilized alpha helical peptide that mimics pro-apoptotic BH3-only proteins, such as BID and BAD.[7] This molecule was designed to stabilize the native helical structure by forming a macrocycle between side chains that are not involved in binding. This process, referred to as peptide stapling, uses non-natural amino acids to facilitate macrocyclization by ring-closing olefin metathesis.[8] In this case, a stapled BH3 helix was identified which specifically activates the mitochondrial apoptotic pathway by antagonizing the sequestration of BH3-only proteins by anti-apoptotic proteins (e.g. Bcl-2, see also intrinsic and extrinsic inducers of the apoptosis). This molecule suppressed growth of human leukemia in a mouse xenograft model.[7]
Also in 2004, Harran and co-workers reported a dimeric small molecule that mimics the proapoptotic protein Smac (see mitochondrial regulation in apoptosis).[9] This molecule mimics the N-terminal linear motif Ala-Val-Pro-Ile. Uniquely, the dimeric structure of this peptidomimetic led to a marked increase in activity over an analogous monomer. This binding cooperativity results from the molecule's ability to also mimic the homodimeric structure of Smac, which is functionally important for reactivating caspases.[10] Smac mimetics of this type can sensitize an array of non-small-cell lung cancer cells to conventional chemotherapeutics (e.g. Gemcitabine, Vinorelbine) both in vitro and in mouse xenograft models.[11]
Heterocycles are often used to mimic the amide bond of peptides. Thiazoles, for example, are found in naturally occurring peptides and used by researchers to mimic the amide bond of peptides.[12]
See also[edit]
- Apoptosis
- Beta-peptide
- Cancer
- Clicked peptide polymer
- Depsipeptide
- Expanded genetic code
- Foldamers
- Non-proteinogenic amino acids
- Stapled Peptides
References[edit]
- ^ a b c d e f Pelay-Gimeno M, Glas A, Koch O, Grossmann TN (July 2015). "Structure-Based Design of Inhibitors of Protein-Protein Interactions: Mimicking Peptide Binding Epitopes". Angewandte Chemie. 54 (31): 8896–927. doi:10.1002/anie.201412070. PMC 4557054. PMID 26119925.
- ^ Marshall GR, Ballante F (September 2017). "Limiting Assumptions in the Design of Peptidomimetics". Drug Development Research. 78 (6): 245–267. doi:10.1002/ddr.21406. PMID 28875546. S2CID 5730986.
- ^ a b c d Avan, Ilker; Hall, C. Dennis; Katritzky, Alan R. (22 April 2014). "Peptidomimetics via modifications of amino acids and peptide bonds". Chemical Society Reviews. 43 (10): 3575–3594. doi:10.1039/C3CS60384A. PMID 24626261.
- ^ Orner, Brendan P.; Ernst, Justin T.; Hamilton, Andrew D. (2001-06-01). "Toward Proteomimetics: Terphenyl Derivatives as Structural and Functional Mimics of Extended Regions of an α-Helix". Journal of the American Chemical Society. 123 (22): 5382–5383. doi:10.1021/ja0025548. ISSN 0002-7863. PMID 11457415.
- ^ Owen DR, Allerton CM, Anderson AS, Aschenbrenner L, Avery M, Berritt S, et al. (November 2021). "An oral SARS-CoV-2 Mpro inhibitor clinical candidate for the treatment of COVID-19". Science. 374 (6575): 1586–1593. Bibcode:2021Sci...374.1586O. doi:10.1126/science.abl4784. PMID 34726479. S2CID 240422219.
- ^ Gomari MM, Abkhiz S, Pour TG, Lotfi E, Rostami N, Monfared FN, Ghobari B, Mosavi M, Alipour B, Dokholyan NV (December 2022). "Peptidomimetics in cancer targeting". Mol Med. 28 (1): 146. doi:10.1186/s10020-022-00577-3. PMC 9730693. PMID 36476230.
- ^ a b Walensky LD, Kung AL, Escher I, Malia TJ, Barbuto S, Wright RD, Wagner G, Verdine GL, Korsmeyer SJ (September 2004). "Activation of apoptosis in vivo by a hydrocarbon-stapled BH3 helix". Science. 305 (5689): 1466–70. Bibcode:2004Sci...305.1466W. doi:10.1126/science.1099191. PMC 1360987. PMID 15353804.
- ^ Blackwell HE, Grubbs RH (1998). "Highly Efficient Synthesis of Covalently Cross-Linked Peptide Helices by Ring-Closing Metathesis". Angewandte Chemie International Edition. 37 (23): 3281–3284. doi:10.1002/(SICI)1521-3773(19981217)37:23<3281::AID-ANIE3281>3.0.CO;2-V. PMID 29711420.
- ^ Li L, Thomas RM, Suzuki H, De Brabander JK, Wang X, Harran PG (September 2004). "A small molecule Smac mimic potentiates TRAIL- and TNFalpha-mediated cell death". Science. 305 (5689): 1471–4. Bibcode:2004Sci...305.1471L. doi:10.1126/science.1098231. PMID 15353805. S2CID 58926089.
- ^ Chai J, Du C, Wu JW, Kyin S, Wang X, Shi Y (August 2000). "Structural and biochemical basis of apoptotic activation by Smac/DIABLO". Nature. 406 (6798): 855–62. Bibcode:2000Natur.406..855C. doi:10.1038/35022514. PMID 10972280. S2CID 4385614.
- ^ Greer RM, Peyton M, Larsen JE, Girard L, Xie Y, Gazdar AF, Harran P, Wang L, Brekken RA, Wang X, Minna JD (December 2011). "SMAC mimetic (JP1201) sensitizes non-small cell lung cancers to multiple chemotherapy agents in an IAP-dependent but TNF-α-independent manner". Cancer Research. 71 (24): 7640–8. doi:10.1158/0008-5472.CAN-10-3947. PMC 3382117. PMID 22049529.
- ^ Mak JY, Xu W, Fairlie DP (2015-01-01). Peptidomimetics I (PDF). Topics in Heterocyclic Chemistry. Vol. 48. Springer Berlin Heidelberg. pp. 235–266. doi:10.1007/7081_2015_176. ISBN 978-3-319-49117-2.


