Obsolete models of DNA structure
In addition to the variety of verified DNA structures, there have been a range of proposed DNA models that have either been disproven, or lack evidence.
Some of these structures were proposed during the 1950s before the structure of the double helix was solved, most famously by Linus Pauling. Non-helical or "side-by-side" models of DNA were proposed in the 1970s to address what appeared at the time to be problems with the topology of circular DNA chromosomes during replication (subsequently resolved via the discovery of enzymes that modify DNA topology).[1] These were also rejected due to accumulating experimental evidence from X-ray crystallography, solution NMR, and atomic force microscopy (of both DNA alone, and bound to DNA-binding proteins). Although localised or transient non-duplex helical structures exist,[2] non-helical models are not currently accepted by the mainstream scientific community.[3] Finally, there exists a persistent set of contemporary fringe theories proposing a range of unsupported models.
Prior to Watson–Crick structure
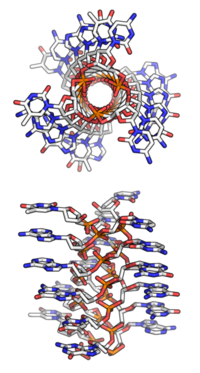
The DNA double helix was discovered in 1953[4] (with further details in 1954[5]) based on X-ray diffraction images of DNA (most notably photo 51, taken by Raymond Gosling and Rosalind Franklin[6]) as well as base-pairing chemical and biochemical information.[7][8] Prior to this, X-ray data being gathered in the 1950s indicated that DNA formed some sort of helix, but it had not yet been discovered what the exact structure of that helix was. There were therefore several proposed structures that were later overturned by the data supporting a DNA duplex. The most famous of these early models was by Linus Pauling and Robert Corey in 1953 in which they proposed a triple helix with the phosphate backbone on the inside, and the nucleotide bases pointing outwards.[9][10] A broadly similar, but detailed structure was also proposed by Bruce Fraser that same year.[11] However, Watson and Crick soon identified several problems with these models:
- Negatively charged phosphates near the axis repel each other, leaving the question of how the three-chain structure stays together.
- In a triple-helix model (specifically Pauling and Corey's model), some of the van der Waals distances appear to be too small.
The initial double helix model discovered, now termed B-form DNA is by far the most common conformation in cells.[12] Two additional rarer helical conformations that also naturally occur were identified in the 1970s: A-form DNA, and Z-form DNA.[13]
Non-helical structure proposals
Before the discovery of topoisomerases
Even once the DNA duplex structure was solved, it was initially an open question whether additional DNA structures were needed to explain its overall topology. there were initially questions about how it might affect DNA replication. In 1963, autoradiographs of the E. coli chromosome demonstrated that it was a single circular molecule that is replicated at a pair of replication forks at which both new DNA strands are being synthesized.[15] The two daughter chromosomes after replication would therefore be topologically linked. The separation of the two linked daughter DNA strands during replication either required DNA to have a net-zero helical twist, or for the strands to be cut, crossed, and rejoined. It was this apparent contradictions that early non-helical models attempted to address until the discovery of topoisomerases in 1970 resolved the problem.[16][17]
In the 1960s and 1970s, a number of structures were hypothesised that would give a net-zero helical twist over the length of the DNA, either by being fully straight throughout or by alternating right-handed and left-handed helical twists.[18][19] For example, in 1969, a linear tetramer structure was hypothesised,[14] and in 1976, a structure with alternating sections of right-handed and left-handed helix was independently proposed by two different groups.[20][21] The alternating twists model was initially presented with the helicity changing every half turn, but later long stretches of each helical direction were later proposed.[22] However, these models suffered from a lack of experimental support.[23] Under torsional stress, a Z-DNA structure can form with opposite twist to B-form DNA, but this is rare within the cellular environment.[24] The discovery of topoisomerases and gyrases, enzymes that can change the linking number of circular nucleic acids and thus "unwind" and "rewind" the replicating bacterial chromosome, solved the topological objections to the B-form DNA helical structure.[25] Indeed, in the absence of these topology-altering enzymes, small circular viral and plasmid DNA are inseparable supporting structure whose strands are topologically locked together.[26]
Non-helical DNA proposals have therefore dropped from mainstream science.[3][16]
Confirmation of helical structure
Initially, there had been questions of whether the solved DNA structures were artefacts of the X-ray crystallography techniques used. However, the structure of DNA was subsequently confirmed in solution via gel electrophoretic methods[27] and later via solution NMR[28] and AFM[29] indicating that the crystallography process did not distort it. The structure of DNA in complex with nucleosomes, helicases, and numerous other DNA binding proteins also supported its biological relevance in vivo.[30]
References
- ^ Stokes, T. D. (1982). "The double helix and the warped zipper—an exemplary tale". Social Studies of Science. 12 (2): 207–240. doi:10.1177/030631282012002002. PMID 11620855. S2CID 29369576.
- ^ Sinden, Richard R. (1994). "Miscellaneous Alternative Conformations of DNA". DNA Structure and Function. Elsevier. pp. 259–286. doi:10.1016/b978-0-08-057173-7.50012-2. ISBN 9780080571737.
- ^ a b Gautham, N. (2004). "Response to "Variety in DNA secondary structure"" (PDF). Current Science. 86 (10): 1352–1353.
However, the discovery of topoisomerases took "the sting" out of the topological objection to the plectonaemic double helix. The more recent solution of the single crystal X-ray structure of the nucleosome core particle showed nearly 150 base pairs of the DNA (i.e. about 15 complete turns), with a structure that is in all essential respects the same as the Watson–Crick model. This dealt a death blow to the idea that other forms of DNA, particularly double helical DNA, exist as anything other than local or transient structures.
- ^ Watson, J. D.; Crick, F. (1953). "A structure for deoxyribose nucleic acid" (PDF). Nature. 171 (4356): 737–8. Bibcode:1953Natur.171..737W. doi:10.1038/171737a0. PMID 13054692. S2CID 4253007.
- ^ Crick, F. & Watson, J. D. (1954). "The Complementary Structure of Deoxyribonucleic Acid" (PDF). Proceedings of the Royal Society of London. 223A (1152): 80–96. Bibcode:1954RSPSA.223...80C. doi:10.1098/rspa.1954.0101. S2CID 45478298.
- ^ "The story behind Photograph 51 | Feature from King's College London". www.kcl.ac.uk. Retrieved 2023-08-29.
- ^ Magasanik B, Vischer E, Doniger R, Elson D, Chargaff E (1950). "The separation and estimation of ribonucleotides in minute quantities". J. Biol. Chem. 186 (1): 37–50. doi:10.1016/S0021-9258(18)56284-0. PMID 14778802.
- ^ Chargaff E (1950). "Chemical specificity of nucleic acids and mechanism of their enzymatic degradation". Experientia. 6 (6): 201–209. doi:10.1007/BF02173653. PMID 15421335. S2CID 2522535.
- ^ Pauling L, Corey RB (February 1953). "A Proposed Structure For The Nucleic Acids". Proceedings of the National Academy of Sciences of the United States of America. 39 (2): 84–97. Bibcode:1953PNAS...39...84P. doi:10.1073/pnas.39.2.84. PMC 1063734. PMID 16578429.
- ^ Pauling L, Corey RB (February 1953). "Structure of the nucleic acids". Nature. 171 (4347): 346. Bibcode:1953Natur.171..346P. doi:10.1038/171346a0. PMID 13036888. S2CID 4151877.
- ^ Fraser B (1953). "The Structure of Deoxyribose Nucleic Acid". Journal of Structural Biology. 145 (3): 184–5. doi:10.1016/j.jsb.2004.01.001. PMID 14997898.
- ^ Richmond TJ, Davey CA (2003). "The structure of DNA in the nucleosome core". Nature. 423 (6936): 145–50. Bibcode:2003Natur.423..145R. doi:10.1038/nature01595. PMID 12736678. S2CID 205209705.
- ^ Sinden, Richard R. (1994). DNA structure and function. San Diego: Academic Press. ISBN 0126457506. OCLC 30109829.
- ^ a b Wu, Tai Te (1969). "Secondary Structures of DNA". Proceedings of the National Academy of Sciences. 63 (2): 400–405. doi:10.1073/pnas.63.2.400. ISSN 0027-8424. PMC 223578. PMID 5257129.
- ^ Cairns, J. (1963). "The bacterial chromosome and its manner of replication as seen by autoradiography". J. Mol. Biol. 6 (3): 208–13. doi:10.1016/s0022-2836(63)80070-4. PMID 14017761.
- ^ a b Biegeleisen K (2002). "Topologically non-linked circular duplex DNA". Bull. Math. Biol. 64 (3): 589–609. CiteSeerX 10.1.1.573.5418. doi:10.1006/bulm.2002.0288. PMID 12094410. S2CID 13269612.
- ^ Wang, J. C. (2002). "Cellular roles of DNA topoisomerases: a molecular perspective". Nature Reviews Molecular Cell Biology. 3 (6): 430–440. doi:10.1038/nrm831. ISSN 1471-0072. PMID 12042765. S2CID 205496065.
- ^ a b Rodley GA, Scobie RS, Bates RH, Lewitt RM (1976). "A possible conformation for double-stranded polynucleotides". Proc. Natl. Acad. Sci. U.S.A. 73 (9): 2959–63. Bibcode:1976PNAS...73.2959R. doi:10.1073/pnas.73.9.2959. PMC 430891. PMID 1067594.
- ^ Sasisekharan, V.; Pattabiraman, N. (1976). "Double standard polynucleotides: Two typical alternative conformations for nucleic acids" (PDF). Curr. Sci. 45: 779–783.
- ^ Sasisekharan, V.; Pattabiraman, N. (1976). "Double stranded polynucleotides: two typical alternative conformations for nucleic acids". Current Science. 45: 779–783.
- ^ Sasisekharan, V.; Pattabiraman, N.; Gupta, G. (1978). "Some implications of an alternative structure for DNA". Proc. Natl. Acad. Sci. U.S.A. 75 (9): 4092–6. Bibcode:1978PNAS...75.4092S. doi:10.1073/pnas.75.9.4092. PMC 336057. PMID 279899.
- ^ Rodley, G. A. (1995). "Reconsideration of some results for linear and circular DNA". Journal of Biosciences. 20 (2): 245–257. doi:10.1007/BF02703272. S2CID 40531350.
- ^ Crick, F. H.; Wang, J. C.; Bauer, W. R. (1979). "Is DNA really a double helix?" (PDF). J. Mol. Biol. 129 (3): 449–457. doi:10.1016/0022-2836(79)90506-0. PMID 458852.
- ^ Rich, Alexander; Zhang, Shuguang (2003). "Z-DNA: the long road to biological function". Nature Reviews Genetics. 4 (7): 566–572. doi:10.1038/nrg1115. ISSN 1471-0064. PMID 12838348. S2CID 835548.
- ^ Keszthelyi, Andrea; Minchell, Nicola; Baxter, Jonathan (2016). "The Causes and Consequences of Topological Stress during DNA Replication". Genes. 7 (12): 134. doi:10.3390/genes7120134. ISSN 2073-4425. PMC 5192510. PMID 28009828.
- ^ Vinograd J, Lebowitz J, Radloff R, Watson R, Laipis P (1965). "The twisted circular form of polyoma viral DNA". Proc. Natl. Acad. Sci. U.S.A. 53 (5): 1104–11. Bibcode:1965PNAS...53.1104V. doi:10.1073/pnas.53.5.1104. PMC 301380. PMID 4287964.
- ^ Wang, J. C. (1979). "Helical repeat of DNA in solution". Proceedings of the National Academy of Sciences. 76 (1): 200–203. Bibcode:1979PNAS...76..200W. doi:10.1073/pnas.76.1.200. ISSN 0027-8424. PMC 382905. PMID 284332.
- ^ Ghosh, Anirban; Kar, Rajiv Kumar; Jana, Jagannath; Saha, Abhijit; Jana, Batakrishna; Krishnamoorthy, Janarthanan; Kumar, Dinesh; Ghosh, Surajit; Chatterjee, Subhrangsu (2014). "Indolicidin Targets Duplex DNA: Structural and Mechanistic Insight through a Combination of Spectroscopy and Microscopy". ChemMedChem. 9 (9): 2052–2058. doi:10.1002/cmdc.201402215. hdl:2027.42/108345. ISSN 1860-7179. PMID 25044630. S2CID 33138700.
- ^ Pyne, Alice; Thompson, Ruth; Leung, Carl; Roy, Debdulal; Hoogenboom, Bart W. (2014). "Single-Molecule Reconstruction of Oligonucleotide Secondary Structure by Atomic Force Microscopy" (PDF). Small. 10 (16): 3257–3261. doi:10.1002/smll.201400265. ISSN 1613-6829. PMID 24740866. S2CID 5050707.
- ^ Morgunova, E.; Taipale, J. (2017). "Structural perspective of cooperative transcription factor binding". Current Opinion in Structural Biology. Protein–nucleic acid interactions • Catalysis and regulation. 47: 1–8. doi:10.1016/j.sbi.2017.03.006. ISSN 0959-440X. PMID 28349863.
