Trajectory inference

Trajectory inference or pseudotemporal ordering is a computational technique used in single-cell transcriptomics to determine the pattern of a dynamic process experienced by cells and then arrange cells based on their progression through the process. Single-cell protocols have much higher levels of noise than bulk RNA-seq,[1] so a common step in a single-cell transcriptomics workflow is the clustering of cells into subgroups.[2] Clustering can contend with this inherent variation by combining the signal from many cells, while allowing for the identification of cell types.[3] However, some differences in gene expression between cells are the result of dynamic processes such as the cell cycle, cell differentiation, or response to an external stimuli. Trajectory inference seeks to characterize such differences by placing cells along a continuous path that represents the evolution of the process rather than dividing cells into discrete clusters.[4] In some methods this is done by projecting cells onto an axis called pseudotime which represents the progression through the process.[5]
Methods
[edit]Since 2015, more than 50 algorithms for trajectory inference have been created.[6] Although the approaches taken are diverse there are some commonalities to the methods. Typically, the steps in the algorithm consist of dimensionality reduction to reduce the complexity of the data, trajectory building to determine the structure of the dynamic process, and projection of the data onto the trajectory so that cells are positioned by their development through the process and cells with similar expression profiles are situated near each other.[6] Trajectory inference algorithms differ in the specific procedure used for dimensionality reduction, the kinds of structures that can be used to represent the dynamic process, and the prior information that is required or can be provided.[2]

Dimensionality reduction
[edit]The data produced by single-cell RNA-seq can consist of thousands of cells each with expression levels recorded across thousands of genes.[7] In order to efficiently process data with such high dimensionality many trajectory inference algorithms employ a dimensionality reduction procedure such as principal component analysis (PCA), independent component analysis (ICA), or t-SNE as their first step.[8] The purpose of this step is to combine many features of the data into a more informative measure of the data.[4] For example, a coordinate resulting from dimensionality reduction could combine expression levels from many genes that are associated with the cell cycle into one value that represents a cell's position in the cell cycle.[8] Such a transformation corresponds to dimensionality reduction in the feature space, but dimensionality reduction can also be applied to the sample space by clustering together groups of similar cells.[1]
Trajectory building
[edit]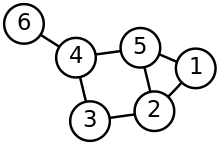
Many methods represent the structure of the dynamic process via a graph-based approach. In such an approach the vertices of the graph correspond to states in the dynamic process, such as cell types in cell differentiation, and the edges between the nodes correspond to transitions between the states.[6] The creation of the trajectory graph can be accomplished using k-nearest neighbors or minimum spanning tree algorithms.[9] The topology of the trajectory refers to the structure of the graph and different algorithms are limited to creation of graph topologies of a particular type such as linear, branching, or cyclic.[4]
Use of prior information
[edit]Some methods require or allow for the input of prior information which is used to guide the creation of the trajectory. The use of prior information can lead to more accurate trajectory determination, but poor priors can lead the algorithm astray or bias results towards expectations.[6] Examples of prior information that can be used in trajectory inference are the selection of start cells that are at the beginning of the trajectory, the number of branches in the trajectory, and the number of end states for the trajectory.[10]
Software
[edit]MARGARET
[edit]MARGARET employs a deep unsupervised metric learning approach for inferring the cellular latent space and cell clusters. The trajectory is modeled using a cluster-connectivity graph to capture complex trajectory topologies. MARGARET utilizes the inferred trajectory for determining terminal states and inferring cell-fate plasticity using a scalable Absorbing Markov chain model.[11]
Monocle
[edit]Monocle first employs a differential expression test to reduce the number of genes then applies independent component analysis for additional dimensionality reduction. To build the trajectory Monocle computes a minimum spanning tree, then finds the longest connected path in that tree. Cells are projected onto the nearest point to them along that path.[5]
p-Creode
[edit]p-Creode finds the most likely path through a density-adjusted k-nearest neighbor graph. Graphs from an ensemble are scored with a graph similarity metric to select the most representative topology. p-Creode has been tested on a range of single-cell platforms, including mass cytometry, multiplex immunofluorescence,[12] and single-cell RNA-seq. No prior information is required.[13]
Slingshot
[edit]Slingshot takes cluster labels as input and then orders these clusters into lineages by the construction of a minimum spanning tree. Paths through the tree are smoothed by fitting simultaneous principal curves and a cell's pseudotime value is determined by its projection onto one or more of these curves. Prior information, such as initial and terminal clusters, is optional.[10]
TSCAN
[edit]TSCAN performs dimensionality reduction using principal component analysis and clusters cells using a mixture model. A minimum spanning tree is calculated using the centers of the clusters and the trajectory is determined as the longest connected path of that tree. TSCAN is an unsupervised algorithm that requires no prior information.[14]
Wanderlust/Wishbone
[edit]Wanderlust was developed for analysis of mass cytometry data, but has been adapted for single-cell transcriptomics applications. A k-nearest neighbors algorithm is used to construct a graph which connects every cell to the cell closest to it with respect to a metric such as Euclidean distance or cosine distance. Wanderlust requires the input of a starting cell as prior information.[15]
Wishbone is built on Wanderlust and allows for a bifurcation in the graph topology, whereas Wanderlust creates a linear graph. Wishbone combines principal component analysis and diffusion maps to achieve dimensionality reduction then also creates a KNN graph.[16]
Waterfall
[edit]Waterfall performs dimensionality reduction via principal component analysis and uses a k-means algorithm to find cell clusters. A minimal spanning tree is built between the centers of the clusters. Waterfall is entirely unsupervised, requiring no prior information, and produces linear trajectories.[17]
References
[edit]- ^ a b Bacher, Rhonda; Kendziorski, Christina (2016-04-07). "Design and computational analysis of single-cell RNA-sequencing experiments". Genome Biology. 17 (1): 63. doi:10.1186/s13059-016-0927-y. ISSN 1474-760X. PMC 4823857. PMID 27052890.
- ^ a b Hwang, Byungjin; Lee, Ji Hyun; Bang, Duhee (2018-08-07). "Single-cell RNA sequencing technologies and bioinformatics pipelines". Experimental & Molecular Medicine. 50 (8): 1–14. doi:10.1038/s12276-018-0071-8. ISSN 2092-6413. PMC 6082860. PMID 30089861.
- ^ Stegle, Oliver; Teichmann, Sarah A.; Marioni, John C. (2015-01-28). "Computational and analytical challenges in single-cell transcriptomics". Nature Reviews Genetics. 16 (3): 133–145. doi:10.1038/nrg3833. ISSN 1471-0056. PMID 25628217. S2CID 205486032.
- ^ a b c Cannoodt, Robrecht; Saelens, Wouter; Saeys, Yvan (2016-10-19). "Computational methods for trajectory inference from single-cell transcriptomics". European Journal of Immunology. 46 (11): 2496–2506. doi:10.1002/eji.201646347. hdl:1854/LU-8510906. ISSN 0014-2980. PMID 27682842. S2CID 19562455.
- ^ a b Trapnell, Cole; Cacchiarelli, Davide; Grimsby, Jonna; Pokharel, Prapti; Li, Shuqiang; Morse, Michael; Lennon, Niall J; Livak, Kenneth J; Mikkelsen, Tarjei S (2014-03-23). "The dynamics and regulators of cell fate decisions are revealed by pseudotemporal ordering of single cells". Nature Biotechnology. 32 (4): 381–386. doi:10.1038/nbt.2859. ISSN 1087-0156. PMC 4122333. PMID 24658644.
- ^ a b c d Saelens, Wouter; Cannoodt, Robrecht; Todorov, Helena; Saeys, Yvan (2019-01-04). "A comparison of single-cell trajectory inference methods". Nature Biotechnology. 37 (5): 547–555. doi:10.1038/s41587-019-0071-9. PMID 30936559. S2CID 89616753.
- ^ Conesa, Ana; Madrigal, Pedro; Tarazona, Sonia; Gomez-Cabrero, David; Cervera, Alejandra; McPherson, Andrew; Szcześniak, Michał Wojciech; Gaffney, Daniel J.; Elo, Laura L. (2016-01-26). "A survey of best practices for RNA-seq data analysis". Genome Biology. 17 (1): 13. doi:10.1186/s13059-016-0881-8. ISSN 1474-760X. PMC 4728800. PMID 26813401.
- ^ a b Yosef, Nir; Regev, Aviv; Wagner, Allon (November 2016). "Revealing the vectors of cellular identity with single-cell genomics". Nature Biotechnology. 34 (11): 1145–1160. doi:10.1038/nbt.3711. ISSN 1546-1696. PMC 5465644. PMID 27824854.
- ^ Cahan, Patrick; Tan, Yuqi; Kumar, Pavithra (2017-01-01). "Understanding development and stem cells using single cell-based analyses of gene expression". Development. 144 (1): 17–32. doi:10.1242/dev.133058. ISSN 1477-9129. PMC 5278625. PMID 28049689.
- ^ a b Street, Kelly; Risso, Davide; Fletcher, Russell B.; Das, Diya; Ngai, John; Yosef, Nir; Purdom, Elizabeth; Dudoit, Sandrine (2018-06-19). "Slingshot: cell lineage and pseudotime inference for single-cell transcriptomics". BMC Genomics. 19 (1): 477. doi:10.1186/s12864-018-4772-0. PMC 6007078. PMID 29914354.
- ^ Pandey, Kushagra; Zafar, Hamim (2022). "Inference of cell state transitions and cell fate plasticity from single-cell with MARGARET". Nucleic Acids Research. 50 (15): e86. doi:10.1093/nar/gkac412. ISSN 0305-1048. PMC 9410915. PMID 35639499.
- ^ Gerdes, M. J.; Sevinsky, C. J.; Sood, A.; Adak, S.; Bello, M. O.; Bordwell, A.; Can, A.; Corwin, A.; Dinn, S. (2013-07-01). "Highly multiplexed single-cell analysis of formalin-fixed, paraffin-embedded cancer tissue". Proceedings of the National Academy of Sciences. 110 (29): 11982–11987. Bibcode:2013PNAS..11011982G. doi:10.1073/pnas.1300136110. ISSN 0027-8424. PMC 3718135. PMID 23818604.
- ^ Lau, Ken S.; Coffey, Robert J.; Gerdes, Michael J.; Liu, Qi; Franklin, Jeffrey L.; Roland, Joseph T.; Ping, Jie; Simmons, Alan J.; McKinley, Eliot T. (2018-01-24). "Unsupervised Trajectory Analysis of Single-Cell RNA-Seq and Imaging Data Reveals Alternative Tuft Cell Origins in the Gut". Cell Systems. 6 (1): 37–51.e9. doi:10.1016/j.cels.2017.10.012. ISSN 2405-4712. PMC 5799016. PMID 29153838.
- ^ Ji, Zhicheng; Ji, Hongkai (2016-05-13). "TSCAN: Pseudo-time reconstruction and evaluation in single-cell RNA-seq analysis". Nucleic Acids Research. 44 (13): e117. doi:10.1093/nar/gkw430. ISSN 0305-1048. PMC 4994863. PMID 27179027.
- ^ Bendall, Sean C.; Davis, Kara L.; Amir, El-ad David; Tadmor, Michelle D.; Simonds, Erin F.; Chen, Tiffany J.; Shenfeld, Daniel K.; Nolan, Garry P.; Pe'Er, Dana (2014-04-24). "Single-Cell Trajectory Detection Uncovers Progression and Regulatory Coordination in Human B Cell Development". Cell. 157 (3): 714–725. doi:10.1016/j.cell.2014.04.005. ISSN 0092-8674. PMC 4045247. PMID 24766814.
- ^ Setty, Manu; Tadmor, Michelle D; Reich-Zeliger, Shlomit; Angel, Omer; Salame, Tomer Meir; Kathail, Pooja; Choi, Kristy; Bendall, Sean; Friedman, Nir (2016-05-02). "Wishbone identifies bifurcating developmental trajectories from single-cell data". Nature Biotechnology. 34 (6): 637–645. doi:10.1038/nbt.3569. ISSN 1087-0156. PMC 4900897. PMID 27136076.
- ^ Shin, Jaehoon; Berg, Daniel A.; Zhu, Yunhua; Shin, Joseph Y.; Song, Juan; Bonaguidi, Michael A.; Enikolopov, Grigori; Nauen, David W.; Christian, Kimberly M.; Ming, Guo-li; Song, Hongjun (2015-09-03). "Single-Cell RNA-Seq with Waterfall Reveals Molecular Cascades underlying Adult Neurogenesis". Cell Stem Cell. 17 (3): 360–372. doi:10.1016/j.stem.2015.07.013. ISSN 1934-5909. PMC 8638014. PMID 26299571.
