Dioxygenase: Difference between revisions
No edit summary |
|||
| Line 18: | Line 18: | ||
| CDD = |
| CDD = |
||
}} |
}} |
||
Aerobic life, from simple single-celled bacteria species to complex eukaryotic organisms, has evolved to depend on the oxidizing power of [[dioxygen]] in various metabolic pathways. From energetic [[ATP]] generation to [[xenobiotic]] degradation, the use of dioxygen as a biological oxidant is widespread and varied in the exact mechanism of its use. Enzymes employ many different schemes to use dioxygen, and this largely depends on the substrate and reaction at hand. In the [[monooxygenase|monooxygenases]], only a single atom of dioxygen is incorporated into a substrate with the other being reduced to a water molecule. The dioxygenases ({{EC number|1.13.11}}) catalyze the oxidation a substrate without the reduction of one oxygen atom from dioxygen into a water molecule. However, this definition is ambiguous because it does not take into account how many substrates are involved in the reaction. The majority of dioxygenases fully incorporate dioxygen into a single substrate, and a variety of cofactor schemes are utilized to achieve this. For example, in the [[α-ketoglutarate]]-dependent enzymes, one atom of dioxygen is incorporated into two substrates, with one always being α-ketoglutarate, and this reaction is brought about by a mononuclear iron center. |
|||
{{Infobox protein family |
|||
| Symbol = Dioxygenase_N |
|||
| Name = Catechol dioxygenase N terminus |
|||
| image = PDB 1s9a EBI.jpg |
|||
| ⚫ | |||
| caption = crystal structure of 4-chlorocatechol 1,2-dioxygenase from rhodococcus opacus 1cp |
|||
| Pfam = PF04444 |
|||
| Pfam_clan = |
|||
| InterPro = IPR007535 |
|||
| SMART = |
|||
| PROSITE = |
|||
| MEROPS = |
|||
| SCOP = 1dlm |
|||
| TCDB = |
|||
| OPM family = |
|||
| OPM protein = |
|||
| CAZy = |
|||
| CDD = |
|||
| ⚫ | |||
In molecular biology, a '''dioxygenase''' is an [[enzyme]] which [[catalyse]]s the incorporation of both [[atom]]s of [[oxygen|molecular oxygen]] into [[substrate (biochemistry)|substrates]] using a variety of [[reaction mechanism]]s. [[Bond cleavage|Cleavage]] of [[aromatic rings]] is one of the most important functions of dioxygenases, which play key roles in the [[Chemical decomposition|degradation]] of [[aromatic compounds]]. The [[Enzyme substrate|substrate]]s of ring-cleavage dioxygenases can be classified into two groups according to the mode of scission of the aromatic ring. Intradiol [[enzyme]]s use a non-[[haem]] Fe(III) to [[bond cleavage|cleave]] the aromatic ring between two [[hydroxyl]] groups (ortho-cleavage), whereas extradiol [[enzymes]] use a non-haem Fe(II) to cleave the aromatic ring between a [[hydroxylated]] [[carbon]] and an adjacent non-hydroxylated carbon (meta-cleavage).<ref name="pmid10730195">{{cite journal | author = Broderick JB | title = Catechol dioxygenases | journal = Essays Biochem. | volume = 34 | issue = | pages = 173–89 | year = 1999 | pmid = 10730195 | doi = | url = }}</ref> These two subfamilies differ in sequence, [[secondary structure|structural]] fold, [[iron]] [[ligands]], and the orientation of second sphere [[active site]] [[amino acid]] [[residue (chemistry)|residues]]. |
|||
==Iron-Containing Enzymes== |
|||
Enzymes that belong to the intradiol family include [[catechol 1,2-dioxygenase]] (1,2-CTD) {{EC number|1.13.11.1}}; [[protocatechuate 3,4-dioxygenase]] (3,4-PCD) {{EC number|1.13.11.3}} and [[chlorocatechol 1,2-dioxygenase]] {{EC number|1.13.11.1}}<ref name="pmid15060064">{{cite journal | author = Ferraroni M, Solyanikova IP, Kolomytseva MP, Scozzafava A, Golovleva L, Briganti F | title = Crystal structure of 4-chlorocatechol 1,2-dioxygenase from the chlorophenol-utilizing gram-positive Rhodococcus opacus 1CP | journal = J. Biol. Chem. | volume = 279 | issue = 26 | pages = 27646–55 |date=June 2004 | pmid = 15060064 | doi = 10.1074/jbc.M401692200 | url = }}</ref> |
|||
The most widely observed cofactor involved in dioxygenation reactions is iron, but the catalytic scheme employed by these iron-containing enzymes is highly diverse. Iron-containing dioxygenases can be subdivided into three classes on the basis of how iron is incorporated into the active site: those utilizing a [[heme]] prosthetic group, those containing a [[Rieske protein|Rieske]] [2Fe-2S] cluster, and those employing a mononuclear iron center. |
|||
===Mononuclear Iron Dioxygenases=== |
|||
The mononuclear iron dioxygenases, or non-[[heme]] iron-dependent dioxygenases as they are also termed, all utilize a single catalytic iron to incorporate either one or both atoms of dioxygen into a substrate. Despite this common oxygenation event, the mononuclear iron dioxygenases are diverse in how dioxygen activation is used to promote certain chemical reactions<ref name=leitg>{{cite journal|last=Leitgeb|first=Stefan|coauthors=Nidetzky, Bernd|title=Structural and functional comparison of 2-His- 1-carboxylate and 3-His metallocentres in non-haem iron(II)-dependent enzymes|journal=Biochemical Society Transactions|date=1 December 2008|volume=36|issue=6|pages=1180|doi=10.1042/BST0361180}}</ref> . For instance, carbon-carbon bond cleavage, fatty acid hydroperoxidation, carbon-sulfur bond cleavage, and thiol oxidation are all reactions catalyzed by mononuclear iron dioxygenases<ref name=leitg/><ref name=abu /><ref name=book/>. |
|||
Most mononculear iron dioxygenases are members of the [[cupin superfamily]] in which the overall domain structure is described as a six-stranded β-barrel fold (or [[Beta_barrel#Jelly_roll|jelly roll]] motif). At the center this barrel structure is a metal ion, most commonly ferrous iron, whose coordination environment is frequently provided by residues in two partially conserved structural motifs: G(X)<sub>5</sub>HXH(X)<sub>3</sub>-<sub>4</sub>E(X)<sub>6</sub>G and G(X)<sub>5</sub>-<sub>7</sub>PXG(X)<sub>2</sub>H(X)<sub>3</sub>N <ref name=four /> <ref>{{cite journal|last=Stipanuk|first=Martha H.|coauthors=Simmons, Chad R.; Andrew Karplus, P.; Dominy, John E.|title=Thiol dioxygenases: unique families of cupin proteins|journal=Amino Acids|date=1 March 2010|volume=41|issue=1|pages=91–102|doi=10.1007/s00726-010-0518-2}}</ref>. |
|||
Enzymes that belong to the extradiol class II family include [[catechol 2,3-dioxygenase]] (2,3-CTD) {{EC number|1.13.11.2}} and [[biphenyl-2,3-diol 1,2-dioxygenase]] (BphC) {{EC number|1.13.11.39}}. |
|||
{{multiple image |
|||
| ⚫ | |||
| ⚫ | |||
{{reflist}} |
|||
| align = right |
|||
{{InterPro content|IPR000627}} |
|||
| image1 = Extradiol Mechanism.png |
|||
| alt1 = Extradiol Ring Cleavage |
|||
| caption1 = Figure 1. Extradiol Ring Cleavage |
|||
| image2 = Intradiol Mechanism.png |
|||
| alt2 = Intradiol Ring Cleavage |
|||
| caption2 = Figure 2. Intradiol Ring Cleavage |
|||
| ⚫ | |||
The [[catechol dioxygenase|catechol dioxygenases]], some of the most well-studied dioxygenase enzymes, use dioxygen to cleave a carbon-carbon bond of an aromatic [[catechol]] ring system<ref name=four /> . Catechol dioxygenases are further classified as being “extradiol” or “intradiol,” and this distinction is based on mechanistic differences in the reactions (figures 1 & 2). Intradiol enzymes cleave the carbon-carbon bond between the two hydroxyl groups. The active ferric center is coordinated by four protein ligands—two [[histidine]] and two [[tyrosine|tyrosinate residues]]—in a trigonal bipyramidal manner with a water molecule occupying the fifth coordination site<ref name=book /> . Once a catecholate substrate binds to the metal center in a [[Denticity|bidentate]] fashion through the deprotonated hydroxyl groups, the ferric iron “activates” the substrate by means of abstracting an electron to produce a [[Radical_(chemistry)|radical]] on the substrate. This then allows for reaction with dioxygen and subsequent intradiol cleavage to occur through a cyclic anhydride intermediate<ref name=abu /> <ref name=four /> . Extradiol members utilize ferrous iron as the active redox state, and this center is commonly coordinated octahedrally through a 2-His-1-Glu motif with labile water ligands occupying empty positions. Once a substrate binds to the ferrous center, this promotes dioxygen binding and subsequent activation<ref name=abu /> <ref name=four /> <ref name=bugg /> . This activated oxygen species then proceeds to react with the substrate ultimately cleaving the carbon-carbon bond adjacent to the hydroxyl groups through the formation of an α-keto lactone intermediate<ref name=book />. |
|||
{{clear}} |
|||
[[Category:Protein domains]] |
|||
[[Category:Oxidoreductases]] |
|||
===Rieske Dioxygenases=== |
|||
The Rieske dioxygenases catalyze the cis-dihydroxylation of arenes to cis-dihydro-diol products. These enzymes are prominently found in soil bacteria such as [[''Pseudomonas'']]<ref name=book>{{cite book|first=Samuel de Visser, Devesh Kumar,|title=Iron-containing enzymes versatile catalysts of hydroxylation reactions in nature|date=2011|publisher=Royal Society of Chemistry|isbn=978-1-84973-298-7}}</ref> , and their reactions constitute the initial step in aromatic hydrocarbon biodegradation<ref name=abu>{{cite journal|last=Abu-Omar|first=Mahdi M.|coauthors=Loaiza, Aristobulo; Hontzeas, Nikos|title=Reaction Mechanisms of Mononuclear Non-Heme Iron Oxygenases|journal=Chemical Reviews|date=June 2005|volume=105|issue=6|pages=2227–2252|doi=10.1021/cr040653o}}</ref> . Rieske dioxygenases are structurally more complex than other dioxygenases due to the need for an efficient electron transfer pathway (figure 2) to mediate the additional, simultaneous two-electron reduction of the aromatic substrate. |
|||
[[File:Rieske Dioxygenases.png|thumb|right|Figure 2. Electron transfer mechanism of Rieske dioxygenases]] |
|||
A catalytically-competent Rieske dioxygenase is comprised of three components: an [[FAD reductase (NADH)|NADH-dependent FAD reductase]], a [[ferredoxin]] with two [2Fe-2S] Rieske clusters, and an α3β3 oxygenase with each α-subunit containing a mononuclear iron center and a [2Fe-2S] Rieske cluster<ref name=abu /> . Within each α-subunit, the iron-sulfur cluster and mononuclear iron center are separated by a distance of some ~43 Å, much too far for efficient [[electron transfer]] to occur. Instead, it is proposed electron transfer is mediated through these two centers in adjacent subunits, that the iron-sulfur cluster of one subunit transfers electrons to the mononuclear iron center of the adjacent subunit which is conveniently separated by ~12 Å. While this distance would appear optimal for efficient electron transfer, replacement of the bridging aspartate residue causes a loss of enzyme function, suggesting that electron transfer instead proceeds through the hydrogen-bonding network held in place by this aspartate residue<ref name=book/>. |
|||
[[File:2B1X Active Site.png|thumb|right|Active site of Rieske dioxygenase (naphthalene 1,2-dioxygenase from ''Rhodococcus sp.'') (PDB 2B1X)]] |
|||
The mechanistic picture for this class of dioxygenases is not yet clear, but there is evidence supporting an iron(III) hydroperoxy intermediate in the reaction pathway<ref name=bugg>{{cite journal|last=Bugg|first=Timothy DH|coauthors=Ramaswamy, S|title=Non-heme iron-dependent dioxygenases: unravelling catalytic mechanisms for complex enzymatic oxidations|journal=Current Opinion in Chemical Biology|date=April 2008|volume=12|issue=2|pages=134–140|doi=10.1016/j.cbpa.2007.12.007}}</ref> . This species could represent the active oxidant, or it could undergo hemolytic O-O bond cleavage to yield an iron(V)-oxo intermediate as the working oxidizing agent<ref name=book /><ref name=bugg />. The Rieske dioxygenase are a powerful class of redox-active enzymes, and reactions such as sulfoxidation, [[desaturation]], and benzylic oxidation have been reported in addition to dioxygenation<ref name=abu /> . |
|||
{{clear}} |
|||
===Heme-Containing Dioxygenases=== |
|||
While most iron-dependent dioxygenases utilize a non-heme iron cofactor, the oxidation of L-(and D-)tryptophan to N-formylkynurenine is catalyzed by either [[tryptophan 2,3-dioxygenase]] (TDO) or [[indoleamine 2,3-dioxygenase]] (IDO), which are heme dioxygenases that utilize iron coordinated by a heme B prosthetic group<ref name=one>{{cite journal|last=Efimov|first=Igor|coauthors=Basran, Jaswir; Thackray, Sarah J.; Handa, Sandeep; Mowat, Christopher G.; Raven, Emma Lloyd|title=Structure and Reaction Mechanism in the Heme Dioxygenases|journal=Biochemistry|date=12 April 2011|volume=50|issue=14|pages=2717–2724|doi=10.1021/bi101732n}}</ref><ref name=sono>{{cite journal|last=Sono|first=M|coauthors=Roach, MP; Coulter, ED; Dawson, JH|title=Heme-Containing Oxygenases.|journal=Chemical reviews|date=1996 Nov 7|volume=96|issue=7|pages=2841-2888|pmid=11848843}}</ref> . While these dioxygenases are of interest in part because they uniquely use heme for catalysis, they are also of interest due to their importance in [[tryptophan]] regulation in the cell, which has numerous physiological implications <ref name=TDO>{{cite journal|last=Thackray|first=Sarah J.|coauthors=Mowat, Christopher G.; Chapman, Stephen K.|title=Exploring the mechanism of tryptophan 2,3-dioxygenase|*=Biochemical Society Transactions|date=1 December 2008|volume=36|issue=6|pages=1120|doi=10.1042/BST0361120}}</ref> . The initial association of the substrate with the dioxygen-iron in the enzyme active site is thought to either proceed via radical or electrophilic addition, requiring either ferrous iron or ferric iron, respectively <ref name=one />. While the exact reaction mechanism for the heme-dependent dioxygenases is still under debate, it is postulated that the reaction proceeds through either a dioxetane or [[Criegee rearrangement|Criegee]] mechanism (figures 4, 5) <ref name=one /> <ref name=TDO />. |
|||
{{multiple image |
|||
| width = 300 |
|||
| align = center |
|||
| image1 = Criegee Rearrangement.png |
|||
| alt1 = Criegee Rearrangement |
|||
| caption1 = Figure 4. Possible Criegee Rearrangement Mechanism by TDO/IDO |
|||
| image2 = Dioxetane Mechanism.png |
|||
| alt2 = Dioxetane Mechanism |
|||
| caption2 = Figure 5. Possible Dioxetane Mechanism by TDO/IDO] |
|||
}} |
|||
{{clear}} |
|||
==Cambialistic Dioxygenases== |
|||
While iron is by far the most prevalent cofactor used for enzymatic dioxygenation, it is not required by all dioxygenases for catalysis. [[Quercetin 2,3-dioxygenase]] (quercetinase, QueD) catalyzes the dioxygenolytic cleavage of quercetin to 2-protocatechuoylphloroglucinolcarboxylic acid and carbon monoxide<ref name=five>{{cite journal|last=Schaab|first=MR|coauthors=Barney, BM; Francisco, WA|title=Kinetic and spectroscopic studies on the quercetin 2,3-dioxygenase from Bacillus subtilis.|journal=Biochemistry|date=2006 Jan 24|volume=45|issue=3|pages=1009-16|pmid=16411777}}</ref>. The most characterized enzyme, from Aspergillus japonicus, requires the presence of copper<ref name=four>{{cite journal|last=Fetzner|first=S.|title=Ring-Cleaving Dioxygenases with a Cupin Fold|journal=Applied and Environmental Microbiology|date=27 January 2012|volume=78|issue=8|pages=2505–2514|doi=10.1128/AEM.07651-11}}</ref>, and bacterial quercetinases have been discovered that are quite promiscuous in their requirements of a metal center, with varying degrees of activity reported with substitution of divalent Mn, Co, Fe, Ni, and Cu<ref name=five/>. (Quercetin, role in metabolism). |
|||
[[Acireductone dioxygenase (iron(II)-requiring)|Acireductone (1,2-dihydroxy-5-(methylthio)pent-1-en-3-one) dioxygenase]] (ARD) is found in both prokaryotes and eukaryotes<ref name=four/><ref name=five/><ref name=onne>{{cite journal|last=Maroney|first=Michael J.|coauthors=Ciurli, Stefano|title=Nonredox Nickel Enzymes|journal=Chemical Reviews|date=26 December 2013|doi=10.1021/cr4004488}}</ref>. ARD enzymes from most species bind ferrous iron and catalyze the oxidation of acireductone to 4-(methylthio)-2-oxobutanoate, the α-keto acid of methionine, and formic acid. However, [[Acireductone dioxygenase (Ni2+-requiring)|ARD]] from [[Klebsiella oxytoca|''Klebsiella oxytoca'']] catalyzes an additional reaction when nickel(II) is bound: it instead produces 3-(methylthio)propionate, formate, and carbon monoxide from the reaction of acireductone with dioxygen. The activity of Fe-ARD is closely interwoven with the methionine salvage pathway, in which the methylthioadenosine product of cellular SAM reactions is eventually converted to acireductone. |
|||
While the exact role of Ni-ARD is not known, it is suspected to help regulate methionine levels by acting as a shunt in the salvage pathway. This ''K. oxytoca'' enzyme represents a unique example whereby the metal ion present dictates which reaction is catalyzed. Interestingly, the quercetinases and ARD enzymes all are members of the [[cupin superfamily]], to which the mononuclear iron enzymes also belong<ref name=seven>{{cite journal|last=Boer|first=Jodi L.|coauthors=Mulrooney, Scott B.; Hausinger, Robert P.|title=Nickel-dependent metalloenzymes|journal=Archives of Biochemistry and Biophysics|date=February 2014|volume=544|pages=142–152|doi=10.1016/j.abb.2013.09.002}}</ref> . The metal coordination scheme for the QueD enzymes is either a 3-His or 3-His-1-Glu with the exact arrangement being organism-specific<ref name=four/>. The ARD enzymes all chelate the catalytic metal (either Ni or Fe) through the 3-His-1-Glu motif <ref name=seven/>. In these dioxygenases, the coordinating ligands are provided by both of the typical cupin motifs. In the ARD enzymes, the metal exists in an [[Octahedral molecular geometry|octahedral arrangement]] with the three histidine residues comprising a facial triad<ref name=onne/>. The bacterial quercetinase metal centers typically have a [[Trigonal bipyramidal molecular geometry|trigonal bipyramidal]] or octahedral coordination environment when there are four protein ligands; the metal centers of the copper-dependent QueD enzymes possesses a distorted tetrahedral geometry in which only the three conserved histidine residues provide coordination ligands<ref name=four/><ref name=five/>. Empty coordination sites in all metal centers are occupied by aqua ligands until these are displaced by the incoming substrate. |
|||
The ability of these dioxygenases to retain activity in the presence of other metal cofactors with wide ranges of redox potentials suggests the metal center does not play an active role in the activation of dioxygen. Rather, it is thought the metal center functions to hold the substrate in the proper geometry for it to react with dioxygen. In this respect, these enzymes are reminiscent of the intradiol catechol dioxygenases whereby the metal centers activate the substrate for subsequent reaction with dioxygen. |
|||
==Cofactor-Independent Dioxygenases== |
|||
Dioxygenases that catalyze reactions without the need for a cofactor are much more rare in nature than those that do require them. Two dioxygenases, [[3-hydroxy-4-oxoquinoline 2,4-dioxygenase|1H-3-hydroxy-4-oxo-quinoline 2,4-dioxygenase]] (QDO) and [[3-hydroxy-4-oxoquinaldine 2,4-dioxygenase|1H-3-hydroxy-4-oxoquinaldine 2,4-dioxygenase]] (HDO), have been shown to require neither an organic or metal cofactor<ref name=eight>{{cite journal|last=S.|first=Fetzner|title=Oxygenases without requirement for cofactors or metal ions|journal=Applied Microbiology and Biotechnology|date=1 November 2002|volume=60|issue=3|pages=243–257|doi=10.1007/s00253-002-1123-4}}</ref>. These enzymes catalyze the degradation of quinolone heterocycles in a manner similar to [[Quercetin 2,3-dioxygenase|quercetin dioxygenase]], but are thought to mediate a radical reaction of a dioxygen molecule with a carbanion on the substrate (figure 5)<ref>{{cite journal|last=Bugg|first=Timothy D.H.|title=Dioxygenase enzymes: catalytic mechanisms and chemical models|journal=Tetrahedron|date=September 2003|volume=59|issue=36|pages=7075–7101|doi=10.1016/S0040-4020(03)00944-X}}</ref>. Both HDO and QDO belong to the [[Alpha/beta hydrolase fold|α/β hydrolase]] superfamily of enzymes, although the cataclytic residues in HDO and QDO do not seem to serve the same function as they do in the rest of the enzymes in the α/β hydrolase superfamily<ref name=eight/>. |
|||
[[File:1H-3-hydroxy-4-oxoquinoline 2,4-dioxygenase Catalytic Mechanism.png|thumb|center|Figure 5. QDO Catalytic Mechanism]] |
|||
==Clinical Significance== |
|||
Due to the degree of diversity in the dioxygenase family, dioxygenases have a wide range of influences in biology: |
|||
- TDO is important for regulating the levels of tryptophan in the body and is expressed in a high number of human tumors<ref>{{cite journal|last=Pilotte|first=L.|coauthors=Larrieu, P.; Stroobant, V.; Colau, D.; Dolusic, E.; Frederick, R.; De Plaen, E.; Uyttenhove, C.; Wouters, J.; Masereel, B.; Van den Eynde, B. J.|title=Reversal of tumoral immune resistance by inhibition of tryptophan 2,3-dioxygenase|journal=Proceedings of the National Academy of Sciences|date=30 January 2012|volume=109|issue=7|pages=2497–2502|doi=10.1073/pnas.1113873109}}</ref> . The other heme iron-dependent dioxygenase, IDO, also has relevance to human health, as it functions in inflammatory responses in the context of certain diseases<ref>{{cite journal|last=Murakami|first=Yuki|coauthors=Hoshi, Masato; Imamura, Yukio; Arioka, Yuko; Yamamoto, Yasuko; Saito, Kuniaki|title=Remarkable Role of Indoleamine 2,3-Dioxygenase and Tryptophan Metabolites in Infectious Diseases: Potential Role in Macrophage-Mediated Inflammatory Diseases|journal=Mediators of Inflammation|date=2013|volume=2013|pages=1–9|doi=doi:10.1155/2013/391984}}</ref> . Since it affects levels of both tryptophan and kynurenine, IDO has also been implicated in influencing systems related to depression in humans<ref>{{cite journal|last=Sublette|first=M. E.|coauthors=Postolache, T. T.|title=Neuroinflammation and Depression: The Role of Indoleamine 2,3-dioxygenase (IDO) as a Molecular Pathway|journal=Psychosomatic Medicine|date=24 August 2012|volume=74|issue=7|pages=668–672|doi=doi:10.1097/PSY.0b013e318268de9f}}</ref> . |
|||
- [[Alkaptonuria]] is a genetic disease that results in a deficiency of [[homogentisate 1,2-dioxygenase]], which is responsible for catalyzing the formation of 4-maleylacetoacetate from homogentisate<ref>{{cite book|last=Voet|first=Donald Voet, Judith G.|title=Biochemistry|date=2011|publisher=John Wiley & Sons|location=Hoboken, NJ|isbn=0470917458|pages=1045|edition=4th ed.}}</ref> . Buildup of homogentisic acid can result in heart valve damage, kidney stones and damage to cartilage in the body<ref>{{cite journal|last=Phornphutkul|first=Chanika|coauthors=Introne, Wendy J.; Perry, Monique B.; Bernardini, Isa; Murphey, Mark D.; Fitzpatrick, Diana L.; Anderson, Paul D.; Huizing, Marjan; Anikster, Yair; Gerber, Lynn H.; Gahl, William A.|title=Natural History of Alkaptonuria|journal=New England Journal of Medicine|date=26 December 2002|volume=347|issue=26|pages=2111–2121|doi=10.1056/NEJMoa021736}}</ref> . |
|||
- [[Pantothenate kinase-associated neurodegeneration]] (PKAN) is an autosomal recessive disorder that can lead to the development of iron granules and Lewy bodies in neurons. A study has shown that patients diagnosed with PKAN were found to have increased cysteine levels in the [[globus pallidus]] as a consequence of a [[cysteine dioxygenase]] deficiency<ref>{{cite journal|last=Perry|first=TL|coauthors=Kish SJ, Norman MG, Yong VW, Whiting S, Crichton JU, Hansen S|title=Hallervorden-Spatz disease: cysteine accumulation and cysteine dioxygenase deficiency in the globus pallidus|journal=Ann Neurol.|date=1985|volume=18|issue=4|pages=482-489|doi=doi: 10.1002/ana.410180411}}</ref> . Patients with PKAN often develop symptoms of dementia and often die at an early age in adulthood. |
|||
- In DNA repair, the Fe (II)/2-oxoglutarate-dependent dioxygenase [[AlkB]], functions in the oxidative removal of alkylation damage to DNA. Failure to remove DNA alkylation damage can result in cytotoxicity or mutagenesis during DNA replication. |
|||
- [[Cyclooxygenase|Cyclooxygenases]] (COX), which are responsible for forming [[prostanoid|prostanoids]] in the human body, are the target of many [[Nonsteroidal anti-inflammatory drugs|NSAID]] pain relievers<ref name=sono/>. Inhibition of COX leads to reduced inflammation and has an analgesic effect, due to the lowered level of prostaglandin and thromboxane synthesis. |
|||
| ⚫ | |||
<references /> |
|||
Revision as of 22:05, 22 February 2014
| Dioxygenase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
 crystal structure of acinetobacter sp. adp1 protocatechuate 3,4-dioxygenase in complex with 3,4-dihydroxybenzoate | |||||||||
| Identifiers | |||||||||
| Symbol | Dioxygenase_C | ||||||||
| Pfam | PF00775 | ||||||||
| Pfam clan | CL0287 | ||||||||
| InterPro | IPR000627 | ||||||||
| PROSITE | PDOC00079 | ||||||||
| SCOP2 | 2pcd / SCOPe / SUPFAM | ||||||||
| |||||||||
Aerobic life, from simple single-celled bacteria species to complex eukaryotic organisms, has evolved to depend on the oxidizing power of dioxygen in various metabolic pathways. From energetic ATP generation to xenobiotic degradation, the use of dioxygen as a biological oxidant is widespread and varied in the exact mechanism of its use. Enzymes employ many different schemes to use dioxygen, and this largely depends on the substrate and reaction at hand. In the monooxygenases, only a single atom of dioxygen is incorporated into a substrate with the other being reduced to a water molecule. The dioxygenases (EC 1.13.11) catalyze the oxidation a substrate without the reduction of one oxygen atom from dioxygen into a water molecule. However, this definition is ambiguous because it does not take into account how many substrates are involved in the reaction. The majority of dioxygenases fully incorporate dioxygen into a single substrate, and a variety of cofactor schemes are utilized to achieve this. For example, in the α-ketoglutarate-dependent enzymes, one atom of dioxygen is incorporated into two substrates, with one always being α-ketoglutarate, and this reaction is brought about by a mononuclear iron center.
Iron-Containing Enzymes
The most widely observed cofactor involved in dioxygenation reactions is iron, but the catalytic scheme employed by these iron-containing enzymes is highly diverse. Iron-containing dioxygenases can be subdivided into three classes on the basis of how iron is incorporated into the active site: those utilizing a heme prosthetic group, those containing a Rieske [2Fe-2S] cluster, and those employing a mononuclear iron center.
Mononuclear Iron Dioxygenases
The mononuclear iron dioxygenases, or non-heme iron-dependent dioxygenases as they are also termed, all utilize a single catalytic iron to incorporate either one or both atoms of dioxygen into a substrate. Despite this common oxygenation event, the mononuclear iron dioxygenases are diverse in how dioxygen activation is used to promote certain chemical reactions[1] . For instance, carbon-carbon bond cleavage, fatty acid hydroperoxidation, carbon-sulfur bond cleavage, and thiol oxidation are all reactions catalyzed by mononuclear iron dioxygenases[1][2][3].
Most mononculear iron dioxygenases are members of the cupin superfamily in which the overall domain structure is described as a six-stranded β-barrel fold (or jelly roll motif). At the center this barrel structure is a metal ion, most commonly ferrous iron, whose coordination environment is frequently provided by residues in two partially conserved structural motifs: G(X)5HXH(X)3-4E(X)6G and G(X)5-7PXG(X)2H(X)3N [4] [5].
The catechol dioxygenases, some of the most well-studied dioxygenase enzymes, use dioxygen to cleave a carbon-carbon bond of an aromatic catechol ring system[4] . Catechol dioxygenases are further classified as being “extradiol” or “intradiol,” and this distinction is based on mechanistic differences in the reactions (figures 1 & 2). Intradiol enzymes cleave the carbon-carbon bond between the two hydroxyl groups. The active ferric center is coordinated by four protein ligands—two histidine and two tyrosinate residues—in a trigonal bipyramidal manner with a water molecule occupying the fifth coordination site[3] . Once a catecholate substrate binds to the metal center in a bidentate fashion through the deprotonated hydroxyl groups, the ferric iron “activates” the substrate by means of abstracting an electron to produce a radical on the substrate. This then allows for reaction with dioxygen and subsequent intradiol cleavage to occur through a cyclic anhydride intermediate[2] [4] . Extradiol members utilize ferrous iron as the active redox state, and this center is commonly coordinated octahedrally through a 2-His-1-Glu motif with labile water ligands occupying empty positions. Once a substrate binds to the ferrous center, this promotes dioxygen binding and subsequent activation[2] [4] [6] . This activated oxygen species then proceeds to react with the substrate ultimately cleaving the carbon-carbon bond adjacent to the hydroxyl groups through the formation of an α-keto lactone intermediate[3].
Rieske Dioxygenases
The Rieske dioxygenases catalyze the cis-dihydroxylation of arenes to cis-dihydro-diol products. These enzymes are prominently found in soil bacteria such as ''Pseudomonas''[3] , and their reactions constitute the initial step in aromatic hydrocarbon biodegradation[2] . Rieske dioxygenases are structurally more complex than other dioxygenases due to the need for an efficient electron transfer pathway (figure 2) to mediate the additional, simultaneous two-electron reduction of the aromatic substrate.

A catalytically-competent Rieske dioxygenase is comprised of three components: an NADH-dependent FAD reductase, a ferredoxin with two [2Fe-2S] Rieske clusters, and an α3β3 oxygenase with each α-subunit containing a mononuclear iron center and a [2Fe-2S] Rieske cluster[2] . Within each α-subunit, the iron-sulfur cluster and mononuclear iron center are separated by a distance of some ~43 Å, much too far for efficient electron transfer to occur. Instead, it is proposed electron transfer is mediated through these two centers in adjacent subunits, that the iron-sulfur cluster of one subunit transfers electrons to the mononuclear iron center of the adjacent subunit which is conveniently separated by ~12 Å. While this distance would appear optimal for efficient electron transfer, replacement of the bridging aspartate residue causes a loss of enzyme function, suggesting that electron transfer instead proceeds through the hydrogen-bonding network held in place by this aspartate residue[3].

The mechanistic picture for this class of dioxygenases is not yet clear, but there is evidence supporting an iron(III) hydroperoxy intermediate in the reaction pathway[6] . This species could represent the active oxidant, or it could undergo hemolytic O-O bond cleavage to yield an iron(V)-oxo intermediate as the working oxidizing agent[3][6]. The Rieske dioxygenase are a powerful class of redox-active enzymes, and reactions such as sulfoxidation, desaturation, and benzylic oxidation have been reported in addition to dioxygenation[2] .
Heme-Containing Dioxygenases
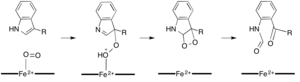
While most iron-dependent dioxygenases utilize a non-heme iron cofactor, the oxidation of L-(and D-)tryptophan to N-formylkynurenine is catalyzed by either tryptophan 2,3-dioxygenase (TDO) or indoleamine 2,3-dioxygenase (IDO), which are heme dioxygenases that utilize iron coordinated by a heme B prosthetic group[7][8] . While these dioxygenases are of interest in part because they uniquely use heme for catalysis, they are also of interest due to their importance in tryptophan regulation in the cell, which has numerous physiological implications [9] . The initial association of the substrate with the dioxygen-iron in the enzyme active site is thought to either proceed via radical or electrophilic addition, requiring either ferrous iron or ferric iron, respectively [7]. While the exact reaction mechanism for the heme-dependent dioxygenases is still under debate, it is postulated that the reaction proceeds through either a dioxetane or Criegee mechanism (figures 4, 5) [7] [9].
Cambialistic Dioxygenases
While iron is by far the most prevalent cofactor used for enzymatic dioxygenation, it is not required by all dioxygenases for catalysis. Quercetin 2,3-dioxygenase (quercetinase, QueD) catalyzes the dioxygenolytic cleavage of quercetin to 2-protocatechuoylphloroglucinolcarboxylic acid and carbon monoxide[10]. The most characterized enzyme, from Aspergillus japonicus, requires the presence of copper[4], and bacterial quercetinases have been discovered that are quite promiscuous in their requirements of a metal center, with varying degrees of activity reported with substitution of divalent Mn, Co, Fe, Ni, and Cu[10]. (Quercetin, role in metabolism). Acireductone (1,2-dihydroxy-5-(methylthio)pent-1-en-3-one) dioxygenase (ARD) is found in both prokaryotes and eukaryotes[4][10][11]. ARD enzymes from most species bind ferrous iron and catalyze the oxidation of acireductone to 4-(methylthio)-2-oxobutanoate, the α-keto acid of methionine, and formic acid. However, ARD from Klebsiella oxytoca catalyzes an additional reaction when nickel(II) is bound: it instead produces 3-(methylthio)propionate, formate, and carbon monoxide from the reaction of acireductone with dioxygen. The activity of Fe-ARD is closely interwoven with the methionine salvage pathway, in which the methylthioadenosine product of cellular SAM reactions is eventually converted to acireductone.
While the exact role of Ni-ARD is not known, it is suspected to help regulate methionine levels by acting as a shunt in the salvage pathway. This K. oxytoca enzyme represents a unique example whereby the metal ion present dictates which reaction is catalyzed. Interestingly, the quercetinases and ARD enzymes all are members of the cupin superfamily, to which the mononuclear iron enzymes also belong[12] . The metal coordination scheme for the QueD enzymes is either a 3-His or 3-His-1-Glu with the exact arrangement being organism-specific[4]. The ARD enzymes all chelate the catalytic metal (either Ni or Fe) through the 3-His-1-Glu motif [12]. In these dioxygenases, the coordinating ligands are provided by both of the typical cupin motifs. In the ARD enzymes, the metal exists in an octahedral arrangement with the three histidine residues comprising a facial triad[11]. The bacterial quercetinase metal centers typically have a trigonal bipyramidal or octahedral coordination environment when there are four protein ligands; the metal centers of the copper-dependent QueD enzymes possesses a distorted tetrahedral geometry in which only the three conserved histidine residues provide coordination ligands[4][10]. Empty coordination sites in all metal centers are occupied by aqua ligands until these are displaced by the incoming substrate.
The ability of these dioxygenases to retain activity in the presence of other metal cofactors with wide ranges of redox potentials suggests the metal center does not play an active role in the activation of dioxygen. Rather, it is thought the metal center functions to hold the substrate in the proper geometry for it to react with dioxygen. In this respect, these enzymes are reminiscent of the intradiol catechol dioxygenases whereby the metal centers activate the substrate for subsequent reaction with dioxygen.
Cofactor-Independent Dioxygenases
Dioxygenases that catalyze reactions without the need for a cofactor are much more rare in nature than those that do require them. Two dioxygenases, 1H-3-hydroxy-4-oxo-quinoline 2,4-dioxygenase (QDO) and 1H-3-hydroxy-4-oxoquinaldine 2,4-dioxygenase (HDO), have been shown to require neither an organic or metal cofactor[13]. These enzymes catalyze the degradation of quinolone heterocycles in a manner similar to quercetin dioxygenase, but are thought to mediate a radical reaction of a dioxygen molecule with a carbanion on the substrate (figure 5)[14]. Both HDO and QDO belong to the α/β hydrolase superfamily of enzymes, although the cataclytic residues in HDO and QDO do not seem to serve the same function as they do in the rest of the enzymes in the α/β hydrolase superfamily[13].

Clinical Significance
Due to the degree of diversity in the dioxygenase family, dioxygenases have a wide range of influences in biology:
- TDO is important for regulating the levels of tryptophan in the body and is expressed in a high number of human tumors[15] . The other heme iron-dependent dioxygenase, IDO, also has relevance to human health, as it functions in inflammatory responses in the context of certain diseases[16] . Since it affects levels of both tryptophan and kynurenine, IDO has also been implicated in influencing systems related to depression in humans[17] .
- Alkaptonuria is a genetic disease that results in a deficiency of homogentisate 1,2-dioxygenase, which is responsible for catalyzing the formation of 4-maleylacetoacetate from homogentisate[18] . Buildup of homogentisic acid can result in heart valve damage, kidney stones and damage to cartilage in the body[19] .
- Pantothenate kinase-associated neurodegeneration (PKAN) is an autosomal recessive disorder that can lead to the development of iron granules and Lewy bodies in neurons. A study has shown that patients diagnosed with PKAN were found to have increased cysteine levels in the globus pallidus as a consequence of a cysteine dioxygenase deficiency[20] . Patients with PKAN often develop symptoms of dementia and often die at an early age in adulthood.
- In DNA repair, the Fe (II)/2-oxoglutarate-dependent dioxygenase AlkB, functions in the oxidative removal of alkylation damage to DNA. Failure to remove DNA alkylation damage can result in cytotoxicity or mutagenesis during DNA replication.
- Cyclooxygenases (COX), which are responsible for forming prostanoids in the human body, are the target of many NSAID pain relievers[8]. Inhibition of COX leads to reduced inflammation and has an analgesic effect, due to the lowered level of prostaglandin and thromboxane synthesis.
References
- ^ a b Leitgeb, Stefan (1 December 2008). "Structural and functional comparison of 2-His- 1-carboxylate and 3-His metallocentres in non-haem iron(II)-dependent enzymes". Biochemical Society Transactions. 36 (6): 1180. doi:10.1042/BST0361180.
{{cite journal}}: Unknown parameter|coauthors=ignored (|author=suggested) (help) - ^ a b c d e f Abu-Omar, Mahdi M. (June 2005). "Reaction Mechanisms of Mononuclear Non-Heme Iron Oxygenases". Chemical Reviews. 105 (6): 2227–2252. doi:10.1021/cr040653o.
{{cite journal}}: Unknown parameter|coauthors=ignored (|author=suggested) (help) - ^ a b c d e f Iron-containing enzymes versatile catalysts of hydroxylation reactions in nature. Royal Society of Chemistry. 2011. ISBN 978-1-84973-298-7.
{{cite book}}:|first=missing|last=(help)CS1 maint: extra punctuation (link) CS1 maint: multiple names: authors list (link) - ^ a b c d e f g h Fetzner, S. (27 January 2012). "Ring-Cleaving Dioxygenases with a Cupin Fold". Applied and Environmental Microbiology. 78 (8): 2505–2514. doi:10.1128/AEM.07651-11.
- ^ Stipanuk, Martha H. (1 March 2010). "Thiol dioxygenases: unique families of cupin proteins". Amino Acids. 41 (1): 91–102. doi:10.1007/s00726-010-0518-2.
{{cite journal}}: Unknown parameter|coauthors=ignored (|author=suggested) (help) - ^ a b c Bugg, Timothy DH (April 2008). "Non-heme iron-dependent dioxygenases: unravelling catalytic mechanisms for complex enzymatic oxidations". Current Opinion in Chemical Biology. 12 (2): 134–140. doi:10.1016/j.cbpa.2007.12.007.
{{cite journal}}: Unknown parameter|coauthors=ignored (|author=suggested) (help) - ^ a b c Efimov, Igor (12 April 2011). "Structure and Reaction Mechanism in the Heme Dioxygenases". Biochemistry. 50 (14): 2717–2724. doi:10.1021/bi101732n.
{{cite journal}}: Unknown parameter|coauthors=ignored (|author=suggested) (help) - ^ a b Sono, M (1996 Nov 7). "Heme-Containing Oxygenases". Chemical reviews. 96 (7): 2841–2888. PMID 11848843.
{{cite journal}}: Check date values in:|date=(help); Unknown parameter|coauthors=ignored (|author=suggested) (help) - ^ a b Thackray, Sarah J. (1 December 2008). "Exploring the mechanism of tryptophan 2,3-dioxygenase". 36 (6): 1120. doi:10.1042/BST0361120.
{{cite journal}}: Cite journal requires|journal=(help); Unknown parameter|*=ignored (help); Unknown parameter|coauthors=ignored (|author=suggested) (help); no-break space character in|coauthors=at position 19 (help); no-break space character in|first=at position 6 (help) - ^ a b c d Schaab, MR (2006 Jan 24). "Kinetic and spectroscopic studies on the quercetin 2,3-dioxygenase from Bacillus subtilis". Biochemistry. 45 (3): 1009–16. PMID 16411777.
{{cite journal}}: Check date values in:|date=(help); Unknown parameter|coauthors=ignored (|author=suggested) (help) - ^ a b Maroney, Michael J. (26 December 2013). "Nonredox Nickel Enzymes". Chemical Reviews. doi:10.1021/cr4004488.
{{cite journal}}: Unknown parameter|coauthors=ignored (|author=suggested) (help) - ^ a b Boer, Jodi L. (February 2014). "Nickel-dependent metalloenzymes". Archives of Biochemistry and Biophysics. 544: 142–152. doi:10.1016/j.abb.2013.09.002.
{{cite journal}}: Unknown parameter|coauthors=ignored (|author=suggested) (help) - ^ a b S., Fetzner (1 November 2002). "Oxygenases without requirement for cofactors or metal ions". Applied Microbiology and Biotechnology. 60 (3): 243–257. doi:10.1007/s00253-002-1123-4.
- ^ Bugg, Timothy D.H. (September 2003). "Dioxygenase enzymes: catalytic mechanisms and chemical models". Tetrahedron. 59 (36): 7075–7101. doi:10.1016/S0040-4020(03)00944-X.
- ^ Pilotte, L. (30 January 2012). "Reversal of tumoral immune resistance by inhibition of tryptophan 2,3-dioxygenase". Proceedings of the National Academy of Sciences. 109 (7): 2497–2502. doi:10.1073/pnas.1113873109.
{{cite journal}}: Unknown parameter|coauthors=ignored (|author=suggested) (help) - ^ Murakami, Yuki (2013). "Remarkable Role of Indoleamine 2,3-Dioxygenase and Tryptophan Metabolites in Infectious Diseases: Potential Role in Macrophage-Mediated Inflammatory Diseases". Mediators of Inflammation. 2013: 1–9. doi:doi:10.1155/2013/391984.
{{cite journal}}: Check|doi=value (help); Unknown parameter|coauthors=ignored (|author=suggested) (help) - ^ Sublette, M. E. (24 August 2012). "Neuroinflammation and Depression: The Role of Indoleamine 2,3-dioxygenase (IDO) as a Molecular Pathway". Psychosomatic Medicine. 74 (7): 668–672. doi:doi:10.1097/PSY.0b013e318268de9f.
{{cite journal}}: Check|doi=value (help); Unknown parameter|coauthors=ignored (|author=suggested) (help) - ^ Voet, Donald Voet, Judith G. (2011). Biochemistry (4th ed. ed.). Hoboken, NJ: John Wiley & Sons. p. 1045. ISBN 0470917458.
{{cite book}}:|edition=has extra text (help)CS1 maint: multiple names: authors list (link) - ^ Phornphutkul, Chanika (26 December 2002). "Natural History of Alkaptonuria". New England Journal of Medicine. 347 (26): 2111–2121. doi:10.1056/NEJMoa021736.
{{cite journal}}: Unknown parameter|coauthors=ignored (|author=suggested) (help) - ^ Perry, TL (1985). "Hallervorden-Spatz disease: cysteine accumulation and cysteine dioxygenase deficiency in the globus pallidus". Ann Neurol. 18 (4): 482–489. doi:doi: 10.1002/ana.410180411.
{{cite journal}}: Check|doi=value (help); Unknown parameter|coauthors=ignored (|author=suggested) (help)