Biological database
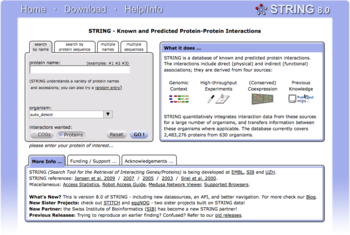
Biological databases are libraries of biological sciences, collected from scientific experiments, published literature, high-throughput experiment technology, and computational analysis.[citation needed] They contain information from research areas including genomics, proteomics, metabolomics, microarray gene expression, and phylogenetics.[2] Information contained in biological databases includes gene function, structure, localization (both cellular and chromosomal), clinical effects of mutations as well as similarities of biological sequences and structures.
Biological databases can be classified by the kind of data they collect (see below). Broadly, there are molecular databases (for sequences, molecules, etc.), functional databases (for physiology, enzyme activities, phenotypes, ecology etc), taxonomic databases (for species and other taxonomic ranks), images and other media, or specimens (for museum collections etc.)
Databases are important tools in assisting scientists to analyze and explain a host of biological phenomena from the structure of biomolecules and their interaction, to the whole metabolism of organisms and to understanding the evolution of species. This knowledge helps facilitate the fight against diseases, assists in the development of medications, predicting certain genetic diseases and in discovering basic relationships among species in the history of life.
Technical basis and theoretical concepts[edit]
Relational database concepts of computer science and Information retrieval concepts of digital libraries are important for understanding biological databases. Biological database design, development, and long-term management is a core area of the discipline of bioinformatics.[3] Data contents include gene sequences, textual descriptions, attributes and ontology classifications, citations, and tabular data. These are often described as semi-structured data, and can be represented as tables, key delimited records, and XML structures.[citation needed]
Access[edit]
Most biological databases are available through web sites that organise data such that users can browse through the data online. In addition the underlying data is usually available for download in a variety of formats. Biological data comes in many formats. These formats include text, sequence data, protein structure and links. Each of these can be found from certain sources, for example:[citation needed]
- Text formats are provided by PubMed and OMIM.
- Sequence data is provided by GenBank, in terms of DNA, and UniProt, in terms of protein.
- Protein structures are provided by PDB, SCOP, and CATH.
Problems and challenges[edit]
Biological knowledge is distributed among countless databases. This sometimes makes it difficult to ensure the consistency of information, e.g. when different names are used for the same species or different data formats. As a consequence, inter-operability is a constant challenge for information exchange. For instance, if a DNA sequence database stores the DNA sequence along the name of a species, a name change of that species may break the links to other databases which may use a different name. Integrative bioinformatics is one field attempting to tackle this problem by providing unified access. One solution is how biological databases cross-reference to other databases with accession numbers to link their related knowledge together (e.g. so that the accession number stays the same even if a species name changes). Redundancy is another problem, as many databases must store the same information, e.g. protein structure databases also contain the sequence of the proteins they cover, their sequence, and their bibliographic information.
Model-organism databases[edit]
Species-specific databases are available for some species, mainly those that are often used in research (model organisms). For example, EcoCyc is an E. coli database. Other popular model organism databases include Mouse Genome Informatics for the laboratory mouse, Mus musculus, the Rat Genome Database for Rattus, ZFIN for Danio Rerio (zebrafish), PomBase[4] for the fission yeast Schizosaccharomyces pombe, FlyBase for Drosophila, WormBase for the nematodes Caenorhabditis elegans and Caenorhabditis briggsae, and Xenbase for Xenopus tropicalis and Xenopus laevis frogs.
Biodiversity and species databases[edit]
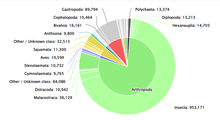
Numerous databases attempt to document the diversity of life on earth. A prominent example is the Catalogue of Life, first created in 2001 by Species 2000 and the Integrated Taxonomic Information System.[6] The Catalogue of Life [1] is a collaborative project that aims to document taxonomic categorization of all currently accepted species in the world.[7] The Catalogue of Life provides a consolidated and consistent database for researchers and policymakers to reference. The Catalogue of Life curates up-to-date datasets from other sources such as Conifer Database, ICTV MSL (for viruses), and LepIndex (for butterflies and moths). In total, the Catalogue of Life draws from 165 databases as of May 2022.[8] Operational costs of the Catalogue of Life are paid for by the Global Biodiversity Information Facility, the Illinois Natural History Survey, the Naturalis Biodiversity Center, and the Smithsonian Institution.[9]
Some biological databases also document geographical distribution of different species. Shuang Dai et al. created a new multi-source database to document spatial/geographical distribution of 1,371 bird species in China, as existing databases had been severely lacking in spatial distribution data for many species.[10] Sources for this new database included books, literature, GPS tracking, and online webpage data. The new database displayed taxonomy, distribution, species info, and data sources for each species. After completion of the bird spatial distribution database, it was discovered that 61% of known species in China were found to be distributed in regions beyond where they were previously known.[11]
Medical databases[edit]

Medical databases are a special case of biomedical data resource and can range from bibliographies, such as PubMed, to image databases for the development of AI based diagnostic software. For instance, one such image database was developed with the goal of aiding in the development of wound monitoring algorithms.[13] Over 188 multi-modal image sets were curated from 79 patient visits, consisting of photographs, thermal images, and 3D mesh depth maps. Wound outlines were manually drawn and added to the photo datasets.[14] The database was made publicly available in the form of a program called WoundsDB, downloadable from the Chronic Wound Database website. [2]
Nucleic Acids Research Database Issue[edit]
An important resource for finding biological databases is a special yearly issue of the journal Nucleic Acids Research (NAR). The Database Issue of NAR is freely available, and categorizes many of the public biological databases. A companion database to the issue called the Online Molecular Biology Database Collection lists 1,380 online databases.[15] Other collections of databases exist such as MetaBase and the Bioinformatics Links Collection.[16][17]
See also[edit]
- Biobank
- Biological data
- Chemical database
- Death Domain database
- European Bioinformatics Institute
- Gene Disease Database
- Integrative bioinformatics
- List of biological databases
- Model organism databases
- NCBI
- PubMed (a database of biomedical literature)
References[edit]
- ^ Szklarczyk D; Franceschini A; Kuhn M; et al. (January 2011). "The STRING database in 2011: functional interaction networks of proteins, globally integrated and scored". Nucleic Acids Res. 39 (Database issue): D561–8. doi:10.1093/nar/gkq973. PMC 3013807. PMID 21045058.
- ^ Altman RB (March 2004). "Building successful biological databases". Brief. Bioinformatics. 5 (1): 4–5. doi:10.1093/bib/5.1.4. PMID 15153301.
- ^ Bourne P (August 2005). "Will a biological database be different from a biological journal?". PLOS Comput. Biol. 1 (3): 179–81. Bibcode:2005PLSCB...1...34B. doi:10.1371/journal.pcbi.0010034. PMC 1193993. PMID 16158097.
- ^ Lock, A; Rutherford, K; Harris, MA; Hayles, J; Oliver, SG; Bähler, J; Wood, V (13 October 2018). "PomBase 2018: user-driven reimplementation of the fission yeast database provides rapid and intuitive access to diverse, interconnected information". Nucleic Acids Research. 47 (D1): D821–D827. doi:10.1093/nar/gky961. PMC 6324063. PMID 30321395.
- ^ Catalogue of Life (2001). "Homepage". Search. Species 2000. Retrieved 2022-05-05.
- ^ Jones, Andrew C. (2011). "Identifying and Relating Biological Concepts in the Catalogue of Life". Journal of Biomedical Semantics. 2 (1): 7. doi:10.1186/2041-1480-2-7. PMC 3245425. PMID 22004596.
- ^ Catalogue of Life (2001). "What is Catalogue of Life?". Our Mission. Species 2000. Retrieved 2022-05-05.
- ^ Catalogue of Life (2001). "Source Datasets". Species 2000. Retrieved 2022-05-05.
- ^ Catalogue of Life (2001). "Funding". Species 2000. Retrieved 2022-05-05.
- ^ Dai, Shuang (2019). "A Spatialized Digital Database for All Bird Species in China". Science China Life Sciences. 62 (5): 661–667. doi:10.1007/s11427-018-9419-2. PMID 30900164. S2CID 84845653. Retrieved 2022-05-05.
- ^ Dai, Shuang (2019). "A Spatialized Digital Database for All Bird Species in China". Science China Life Sciences. 62 (5): 661–667. doi:10.1007/s11427-018-9419-2. PMID 30900164. S2CID 84845653. Retrieved 2022-05-05.
- ^ "Chronic Wound Database". WoundsDB. Silesian University of Technology. 2020. Retrieved 2022-05-05.
- ^ Kręcichwost, Michał (2021). "Chronic Wounds Multimodal Image Database". Computerized Medical Imaging and Graphics. 88: 101844. doi:10.1016/j.compmedimag.2020.101844. PMID 33477091. S2CID 231676950. Retrieved 2022-05-05.
- ^ "Chronic Wound Database". WoundsDB. Silesian University of Technology. 2020. Retrieved 2022-05-05.
- ^ Galperin MY; Fernández-Suárez XM (January 2012). "The 2012 Nucleic Acids Research Database Issue and the online Molecular Biology Database Collection". Nucleic Acids Res. 40 (Database issue): D1–8. doi:10.1093/nar/gkr1196. PMC 3245068. PMID 22144685.
- ^ Bolser DM; Chibon PY; Palopoli N; et al. (January 2012). "MetaBase--the wiki-database of biological databases". Nucleic Acids Res. 40 (Database issue): D1250–4. doi:10.1093/nar/gkr1099. PMC 3245051. PMID 22139927.
- ^ Brazas MD; Yim DS; Yamada JT; Ouellette BF (July 2011). "The 2011 Bioinformatics Links Directory update: more resources, tools and databases and features to empower the bioinformatics community". Nucleic Acids Res. 39 (Web Server issue): W3–7. doi:10.1093/nar/gkr514. PMC 3125814. PMID 21715385.
External links[edit]
- Interactive list of biological databases, classified by categories, from Nucleic Acids Research, 2010
- DBD: Database of Biological Databases
- Biosharing (a database of biological databases)
- Chronic Wounds Database WoundsDB
- Catalogue of Life Catalogue of Life
