Biological database
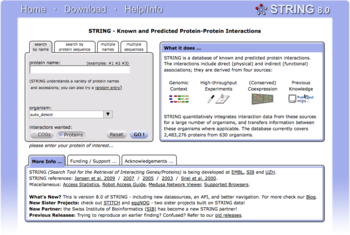
Biological databases are libraries of life sciences information, collected from scientific experiments, published literature, high-throughput experiment technology, and computational analyses.[2] They contain information from research areas including genomics, proteomics, metabolomics, microarray gene expression, and phylogenetics.[3] Information contained in biological databases includes gene function, structure, localization (both cellular and chromosomal), clinical effects of mutations as well as similarities of biological sequences and structures.
Biological databases are an important tool in assisting scientists to understand and explain a host of biological phenomena from the structure of biomolecules and their interaction, to the whole metabolism of organisms and to understanding the evolution of species. This knowledge helps facilitate the fight against diseases, assists in the development of medications and in discovering basic relationships amongst species in the history of life.
Biological knowledge is distributed amongst many different general and specialized databases. This sometimes makes it difficult to ensure the consistency of information. Integrative bioinformatics is one field attempting to tackle this problem by providing unified access. One solution is how biological databases cross-reference to other databases with accession numbers to link their related knowledge together.
Relational database concepts of computer science and Information retrieval concepts of digital libraries are important for understanding biological databases. Biological database design, development, and long-term management is a core area of the discipline of bioinformatics.[4] Data contents include gene sequences, textual descriptions, attributes and ontology classifications, citations, and tabular data. These are often described as semi-structured data, and can be represented as tables, key delimited records, and XML structures.
Nucleic Acids Research Database Issue
An important resource for finding biological databases is a special yearly issue of the journal Nucleic Acids Research (NAR). The Database Issue of NAR is freely available, and categorizes many of the publicly available on line databases related to biology and bioinformatics. A companion database to the issue called the Online Molecular Biology Database Collection lists 1,380 online databases.[5] Other collections of databases exist such as MetaBase and the Bioinformatics Links Collection.[6][7]
Access
Most biological databases are available through web sites that organise data such that users can browse through the data online. In addition the underlying data is usually available for download in a variety of formats. Biological data comes in many formats. These formats include text, sequence data, protein structure and links. Each of these can be found from certain sources, for example:
- Text formats are provided by PubMed and OMIM.
- Sequence data is provided by GenBank, in terms of DNA, and UniProt, in terms of protein.
- Protein structures are provided by PDB, SCOP, and CATH.
Species-specific databases
Species-specific databases are available for some species, mainly those that are often used in research. For example, Colibase ([1]) is an E. coli database. Other popular species specific databases include, FlyBase for Drosophila, and WormBase for the nematodes Caenorhabditis elegans and Caenorhabditis briggsae.
See also
- List of biological databases
- Biobank
- Biological data
- European Bioinformatics Institute (a provider of databases and services)
- MetaBase (a database of biological databases)
- NCBI
- PubMed (a database of biomedical literature)
- Integrative bioinformatics
References
- ^ Szklarczyk D, Franceschini A, Kuhn M; et al. (2011). "The STRING database in 2011: functional interaction networks of proteins, globally integrated and scored". Nucleic Acids Res. 39 (Database issue): D561–8. doi:10.1093/nar/gkq973. PMC 3013807. PMID 21045058.
{{cite journal}}: Explicit use of et al. in:|author=(help); Unknown parameter|month=ignored (help)CS1 maint: multiple names: authors list (link) - ^ Attwood T.K., Gisel A., Eriksson N-E. and Bongcam-Rudloff E. (2011). "Concepts, Historical Milestones and the Central Place of Bioinformatics in Modern Biology: A European Perspective". Bioinformatics - Trends and Methodologies. InTech. Retrieved 8 Jan 2012.
{{cite web}}: CS1 maint: multiple names: authors list (link) - ^ Altman RB (2004). "Building successful biological databases". Brief. Bioinformatics. 5 (1): 4–5. doi:10.1093/bib/5.1.4. PMID 15153301.
{{cite journal}}: Unknown parameter|month=ignored (help) - ^ Bourne P (2005). "Will a biological database be different from a biological journal?". PLoS Comput. Biol. 1 (3): 179–81. doi:10.1371/journal.pcbi.0010034. PMC 1193993. PMID 16158097.
{{cite journal}}: Unknown parameter|month=ignored (help)CS1 maint: unflagged free DOI (link) - ^ Galperin MY, Fernández-Suárez XM (2012). "The 2012 Nucleic Acids Research Database Issue and the online Molecular Biology Database Collection". Nucleic Acids Res. 40 (Database issue): D1–8. doi:10.1093/nar/gkr1196. PMC 3245068. PMID 22144685.
{{cite journal}}: Unknown parameter|month=ignored (help) - ^ Bolser DM, Chibon PY, Palopoli N; et al. (2012). "MetaBase--the wiki-database of biological databases". Nucleic Acids Res. 40 (Database issue): D1250–4. doi:10.1093/nar/gkr1099. PMC 3245051. PMID 22139927.
{{cite journal}}: Explicit use of et al. in:|author=(help); Unknown parameter|month=ignored (help)CS1 maint: multiple names: authors list (link) - ^ Brazas MD, Yim DS, Yamada JT, Ouellette BF (2011). "The 2011 Bioinformatics Links Directory update: more resources, tools and databases and features to empower the bioinformatics community". Nucleic Acids Res. 39 (Web Server issue): W3–7. doi:10.1093/nar/gkr514. PMC 3125814. PMID 21715385.
{{cite journal}}: Unknown parameter|month=ignored (help)CS1 maint: multiple names: authors list (link)
