Slippery sequence
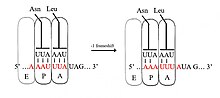
A slippery sequence is a small section of codon nucleotide sequences (usually UUUAAAC) that controls the rate and chance of ribosomal frameshifting. A slippery sequence causes a faster ribosomal transfer which in turn can cause the reading ribosome to "slip." This allows a tRNA to shift by 1 base (−1) after it has paired with its anticodon, changing the reading frame.[2][3][4][5][6] A −1 frameshift triggered by such a sequence is a Programmed −1 Ribosomal Frameshift. It is followed by a spacer region, and an RNA secondary structure. Such sequences are common in virus polyproteins.[1]
The frameshift occurs due to wobble pairing. The Gibbs free energy of secondary structures downstream give a hint at how often frameshift happens.[7] Tension on the mRNA molecule also plays a role.[8] A list of slippery sequences found in animal viruses is available from Huang et al.[9]
Slippery sequences that cause a 2-base slip (−2 frameshift) have been constructed out of the HIV UUUUUUA sequence.[8]
See also
- Nucleic acid tertiary structure
- Open reading frame
- Ribosomal frameshifting
- Translational frameshift
- Transposable element
References
- ^ a b Jacks T, Madhani HD, Masiarz FR, Varmus HE (November 1988). "Signals for ribosomal frameshifting in the Rous sarcoma virus gag-pol region". Cell. 55 (3): 447–58. doi:10.1016/0092-8674(88)90031-1. PMC 7133365. PMID 2846182. S2CID 25672863.
- ^ Green L, Kim CH, Bustamante C, Tinoco I (January 2008). "Characterization of the mechanical unfolding of RNA pseudoknots". Journal of Molecular Biology. 375 (2): 511–28. doi:10.1016/j.jmb.2007.05.058. PMC 7094456. PMID 18021801.
- ^ Yu CH, Noteborn MH, Olsthoorn RC (December 2010). "Stimulation of ribosomal frameshifting by antisense LNA". Nucleic Acids Research. 38 (22): 8277–83. doi:10.1093/nar/gkq650. PMC 3001050. PMID 20693527.
- ^ "Dr Ian Brierley Research description". Department of Pathology, University of Cambridge. Archived from the original on 2013-10-02. Retrieved 2013-07-28.
- ^ "Molecular Biology: Frameshifting occurs at slippery sequences". Molecularstudy.blogspot.com. 2012-10-16. Retrieved 2013-07-28.
- ^ Farabaugh PJ, Björk GR (March 1999). "How translational accuracy influences reading frame maintenance". The EMBO Journal. 18 (6): 1427–34. doi:10.1093/emboj/18.6.1427. PMC 1171232. PMID 10075915.
- ^ Cao S, Chen SJ (March 2008). "Predicting ribosomal frameshifting efficiency". Physical Biology. 5 (1): 016002. Bibcode:2008PhBio...5a6002C. doi:10.1088/1478-3975/5/1/016002. PMC 2442619. PMID 18367782.
- ^ a b Lin Z, Gilbert RJ, Brierley I (September 2012). "Spacer-length dependence of programmed -1 or -2 ribosomal frameshifting on a U6A heptamer supports a role for messenger RNA (mRNA) tension in frameshifting". Nucleic Acids Research. 40 (17): 8674–89. doi:10.1093/nar/gks629. PMC 3458567. PMID 22743270.
- ^ Huang X, Cheng Q, Du Z (2013). "A genome-wide analysis of RNA pseudoknots that stimulate efficient -1 ribosomal frameshifting or readthrough in animal viruses". BioMed Research International. 2013: 984028. doi:10.1155/2013/984028. PMC 3835772. PMID 24298557.
External links
- Pseudobase
- Recode
- Frameshifting,+Ribosomal at the U.S. National Library of Medicine Medical Subject Headings (MeSH)
- Wise2 - aligns a protein against a DNA sequence allowing frameshifts and introns
- FastY - compare a DNA sequence to a protein sequence database, allowing gaps and frameshifts
- Path - tool that compares two frameshift proteins (back-translation principle)
- Recode2 - Database of recoded genes, including those that require programmed Translational frameshift.
- Page for Coronavirus frameshifting stimulation element at Rfam
