Bio-layer interferometry

Bio-layer interferometry (BLI) is an optical biosensing technology that analyzes biomolecular interactions in real-time without the need for fluorescent labeling.[1] Alongside Surface Plasmon Resonance, BLI is one of few widely available label-free biosensing technologies, a detection style that yields more information in less time than traditional processes.[2] The technology relies on the phase shift-wavelength correlation created between interference patterns off of two unique surfaces on the tip of a biosensor.[3] BLI has significant applications in quantifying binding strength, measuring protein interactions, and identifying properties of reaction kinetics, such as rate constants and reaction rates.[4]
Method[edit]
Mechanism overview[edit]

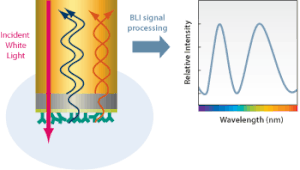
Bio-layer interferometry measures kinetics and biomolecular interactions on a basis of wave interference. To prepare for BLI analysis between two unique biomolecules, the ligand is first immobilized onto a bio compatible biosensor while the analyte is in solution.[5] Shortly after this, the biosensor tip is dipped into the solution and the target molecule will begin to associate with the analyte, producing a layer on top of the biosensor tip. This creates two separate surfaces: the substrate itself, and the substrate interacting with the molecule immobilized on the biosensor tip.[1] This essentially creates a thin-film interference, in which the created layer acts as a thin film bound by these two surfaces. White light from a tungsten lamp is shone onto the biosensor tip and reflected off both surfaces, creating two unique reflection patterns with different intensities.[5] Figure 2 expresses this phenomenon in a more general form. The wavelength shift (Δλ) between these two reflection patterns creates an interference pattern (Figure 3) from which all desired results can be obtained.[1] Since the wavelength shift is direct measure of the change in thickness of the biological layer and the biological layer thickness will change in response to molecules associating to and dissociating from the biosensor, the interference pattern will allow for real-time monitoring of molecular interactions on the biosensor surface.[6] In short, a positive wavelength shift implies an increase in biolayer thickness and thus more association, while a negative wavelength shift implies a decrease in biolayer thickness and thus more dissociation.[6]
"Dip and read" format[edit]
Bio-layer interferometry platforms achieve high throughput by utilizing a "Dip and Read" format.[1] The biosensor tips themselves are transported directly to the desired sample and "dipped" into their respective compartment, eliminating the needs for micro-fluidics and the complications (clogging, purification) that come with it.[1][7] This structure is often supported by a robot, and both 96-well and 384-well plate formats are combined to achieve this.[8] This distinct detection method ensures that sample concentration and viscosity and varying refractive indexes rarely affect the results of BLI.[1] Thus, BLI finds significant use in viscous media such as glycerol, where other techniques may struggle.[9]
Biosensor type and selection[edit]
Bio-layer interferometry relies on biosensors with a fiber optic tip upon which the ligand is immobilized.[1] The tip is additionally coated with a matrix biocompatible with the target molecule to limit any non-specific binding. For BLI calculations to work, it is necessary to assume that both the fiber optic tip and the bound ligand and analyte act as thin, reflective surfaces.[10] The biosensors are disposable, resulting in low costs and high commercial availability.[11] Biosensor selection is determined by the desired test results: kinetic analysis, quantitative analysis, or both.[12] Most commercially available biosensor types will be grouped into one of these three categories by the BLI manufacturer.[1]
Applications[edit]
Analyzing biomolecular interactions[edit]
A key use of Bio-layer interferometry is to analyze and quantify interactions between sets of biomolecules.[1] This is extremely useful in pharmaceutical research, in which biomolecule-membrane interaction determines characteristics of a given drug. Due to its ability to achieve high-resolution data and high throughput, BLI has been used to identify biophysical properties of lipid bilayers, allowing for an alternative method of study than the traditional in vitro methods currently used (microscopy, electrophoresis).[6] In addition, BLI can be used to study effector complex-target interactions. Where the traditional Electrophoretic Mobility Shift Assay (EMSA) method can be used, BLI can act as a suitable substitute if the provided benefits (label-free, real-time measurements) are desired.[3]

Measuring biomolecular kinetics[edit]
Bio-layer interferometry can be used to analyze kinetics in biomolecular systems. The benefits that BLI brings provide additional insight into kinetics on top of commonly used endpoint methods like enzyme-linked immunosorbent assay (ELISA).[1] Interference patterns found in BLI experiments can be used to calculate rate constants and other kinetic data in biomolecular interactions.[13] The (relatively) lower sensitivity of the BLI sensor results in less response to changes in sample composition. As a result, BLI can also be used to investigate allosteric effects on enzyme conformational changes.[14]
Distinguishing characteristics[edit]
BLI and SPR are both dominant technologies in the label-free instruments market.[1] Despite sharing some similarities in concept, there are significant differences between the two techniques. Micro-fluidic SPR relies on a closed architecture to transport samples to a stationary sensor chip (Figure 4). BLI instead utilizes an open system, shaking multiple wells on a plate to transport the sensors to the samples without need for micro-fluidics.[6] Being a closed system, SPR's association and dissociation phases are limited by the technology's design. BLI's open plate design results in association and dissociation length limits determined by sample evaporation instead.[15] SPR is easily reproducible due to its continuous flow microfluidics. BLI's multi well plate design allows for extremely high throughput in one batch. Assay configuration in BLI can, in stable conditions, allow for recovery of samples. Assay configuration in SPR allows for higher sensitivity. As a result, BLI results are often compared to SPR results for validation.[16]
See also[edit]
References[edit]
- ^ a b c d e f g h i j k Apiyo D, Schasfoort R, Schuck P, Marquart A, Gedig ET, Karlsson R, Abdiche YN, Eckman Y, Blum SR, Schasfoort RB (2017). Handbook of Surface Plasmon Resonance. Royal Society of Chemistry. ISBN 978-1-78801-139-6. OCLC 988866146.
- ^ Syahir A, Usui K, Tomizaki KY, Kajikawa K, Mihara H (April 2015). "Label and Label-Free Detection Techniques for Protein Microarrays". Microarrays. 4 (2): 228–244. doi:10.3390/microarrays4020228. PMC 4996399. PMID 27600222.
- ^ a b Müller-Esparza H, Osorio-Valeriano M, Steube N, Thanbichler M, Randau L (2020-05-27). "Bio-Layer Interferometry Analysis of the Target Binding Activity of CRISPR-Cas Effector Complexes". Frontiers in Molecular Biosciences. 7: 98. doi:10.3389/fmolb.2020.00098. PMC 7266957. PMID 32528975.
- ^ Rich RL, Myszka DG (February 2007). "Higher-throughput, label-free, real-time molecular interaction analysis". Analytical Biochemistry. 361 (1): 1–6. doi:10.1016/j.ab.2006.10.040. PMID 17145039.
- ^ a b Müller-Esparza H, Osorio-Valeriano M, Steube N, Thanbichler M, Randau L (2020-05-27). "Bio-Layer Interferometry Analysis of the Target Binding Activity of CRISPR-Cas Effector Complexes". Frontiers in Molecular Biosciences. 7: 98. doi:10.3389/fmolb.2020.00098. PMC 7266957. PMID 32528975.
- ^ a b c d Wallner J, Lhota G, Jeschek D, Mader A, Vorauer-Uhl K (January 2013). "Application of Bio-Layer Interferometry for the analysis of protein/liposome interactions". Journal of Pharmaceutical and Biomedical Analysis. 72: 150–154. doi:10.1016/j.jpba.2012.10.008. PMID 23146240.
- ^ Kamat V, Rafique A (November 2017). "Designing binding kinetic assay on the bio-layer interferometry (BLI) biosensor to characterize antibody-antigen interactions". Analytical Biochemistry. 536: 16–31. doi:10.1016/j.ab.2017.08.002. PMID 28802648.
- ^ Petersen RL (October 2017). "Strategies Using Bio-Layer Interferometry Biosensor Technology for Vaccine Research and Development". Biosensors. 7 (4): 49. doi:10.3390/bios7040049. PMC 5746772. PMID 29088096.
- ^ Lea WA, O'Neil PT, Machen AJ, Naik S, Chaudhri T, McGinn-Straub W, Tischer A, Auton MT, Burns JR, Baldwin MR, Khar KR, Karanicolas J, Fisher MT (September 2016). "Chaperonin-Based Biolayer Interferometry To Assess the Kinetic Stability of Metastable, Aggregation-Prone Proteins". Biochemistry. 55 (35): 4885–908. doi:10.1021/acs.biochem.6b00293. PMC 5524994. PMID 27505032.
- ^ Gao S, Zheng X, Wu J (2017). "A biolayer interferometry-based competitive biosensor for rapid and sensitive detection of saxitoxin". Sensors and Actuators B: Chemical. 246: 169–174. Bibcode:2017SeAcB.246..169G. doi:10.1016/j.snb.2017.02.078. ISSN 0925-4005.
- ^ Abdiche Y, Malashock D, Pinkerton A, Pons J (June 2008). "Determining kinetics and affinities of protein interactions using a parallel real-time label-free biosensor, the Octet". Analytical Biochemistry. 377 (2): 209–217. doi:10.1016/j.ab.2008.03.035. PMID 18405656.
- ^ Yu Y, Mitchell S, Lynaugh H, Brown M, Nobrega RP, Zhi X, et al. (January 2016). "Understanding ForteBio's Sensors for High-Throughput Kinetic and Epitope Screening for Purified Antibodies and Yeast Culture Supernatant". Journal of Biomolecular Screening. 21 (1): 88–95. doi:10.1177/1087057115609564. PMC 4708621. PMID 26442912.
- ^ Wilson JL, Scott IM, McMurry JL (November 2010). "Optical biosensing: Kinetics of protein A-IGG binding using biolayer interferometry". Biochemistry and Molecular Biology Education. 38 (6): 400–407. doi:10.1002/bmb.20442. PMID 21567869. S2CID 29689214.
- ^ Shah NB, Duncan TM (February 2014). "Bio-layer interferometry for measuring kinetics of protein-protein interactions and allosteric ligand effects". Journal of Visualized Experiments (84): e51383. doi:10.3791/51383. PMC 4089413. PMID 24638157.
- ^ Abdiche Y, Malashock D, Pinkerton A, Pons J (June 2008). "Determining kinetics and affinities of protein interactions using a parallel real-time label-free biosensor, the Octet". Analytical Biochemistry. 377 (2): 209–217. doi:10.1016/j.ab.2008.03.035. PMID 18405656.
- ^ Yang D, Singh A, Wu H, Kroe-Barrett R (September 2016). "Comparison of biosensor platforms in the evaluation of high affinity antibody-antigen binding kinetics". Analytical Biochemistry. 508: 78–96. doi:10.1016/j.ab.2016.06.024. PMID 27365220.
