DNA binding site
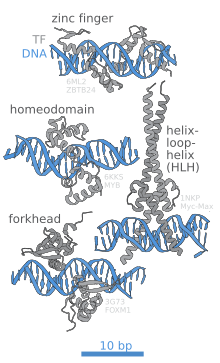
DNA binding sites are a type of binding site found in DNA where other molecules may bind. DNA binding sites are distinct from other binding sites in that (1) they are part of a DNA sequence (e.g. a genome) and (2) they are bound by DNA-binding proteins. DNA binding sites are often associated with specialized proteins known as transcription factors, and are thus linked to transcriptional regulation. The sum of DNA binding sites of a specific transcription factor is referred to as its cistrome. DNA binding sites also encompasses the targets of other proteins, like restriction enzymes, site-specific recombinases (see site-specific recombination) and methyltransferases.[1]
DNA binding sites can be thus defined as short DNA sequences (typically 4 to 30 base pairs long, but up to 200 bp for recombination sites) that are specifically bound by one or more DNA-binding proteins or protein complexes. It has been reported that some binding sites have potential to undergo fast evolutionary change.[2]
Types of DNA binding sites[edit]
DNA binding sites can be categorized according to their biological function. Thus, we can distinguish between transcription factor-binding sites, restriction sites and recombination sites. Some authors have proposed that binding sites could also be classified according to their most convenient mode of representation.[3] On the one hand, restriction sites can be generally represented by consensus sequences. This is because they target mostly identical sequences and restriction efficiency decreases abruptly for less similar sequences. On the other hand, DNA binding sites for a given transcription factor are usually all different, with varying degrees of affinity of the transcription factor for the different binding sites. This makes it difficult to accurately represent transcription factor binding sites using consensus sequences, and they are typically represented using position specific frequency matrices (PSFM), which are often graphically depicted using sequence logos. This argument, however, is partly arbitrary. Restriction enzymes, like transcription factors, yield a gradual, though sharp, range of affinities for different sites [4] and are thus also best represented by PSFM. Likewise, site-specific recombinases also show a varied range of affinities for different target sites.[5][6]
History and main experimental techniques[edit]
The existence of something akin to DNA binding sites was suspected from the experiments on the biology of the bacteriophage lambda[7] and the regulation of the Escherichia coli lac operon.[8] DNA binding sites were finally confirmed in both systems [9][10][11] with the advent of DNA sequencing techniques. From then on, DNA binding sites for many transcription factors, restriction enzymes and site-specific recombinases have been discovered using a profusion of experimental methods. Historically, the experimental techniques of choice to discover and analyze DNA binding sites have been the DNAse footprinting assay and the Electrophoretic Mobility Shift Assay (EMSA). However, the development of DNA microarrays and fast sequencing techniques has led to new, massively parallel methods for in-vivo identification of binding sites, such as ChIP-chip and ChIP-Seq.[12] To quantify the binding affinity[13] of proteins and other molecules to specific DNA binding sites the biophysical method Microscale Thermophoresis[14] is used.
Databases[edit]
Due to the diverse nature of the experimental techniques used in determining binding sites and to the patchy coverage of most organisms and transcription factors, there is no central database (akin to GenBank at the National Center for Biotechnology Information) for DNA binding sites. Even though NCBI contemplates DNA binding site annotation in its reference sequences (RefSeq), most submissions omit this information. Moreover, due to the limited success of bioinformatics in producing efficient DNA binding site prediction tools (large false positive rates are often associated with in-silico motif discovery / site search methods), there has been no systematic effort to computationally annotate these features in sequenced genomes.
There are, however, several private and public databases devoted to compilation of experimentally reported, and sometimes computationally predicted, binding sites for different transcription factors in different organisms. Below is a non-exhaustive table of available databases:
| Name | Organisms | Source | Access | URL |
|---|---|---|---|---|
| PlantRegMap | 165 plant species (e.g., Arabidopsis thaliana, Oryza sativa, Zea mays, etc.) | Expert curation and projection | Public | [1] |
| JASPAR | Vertebrates, Plants, Fungi, Flies, and Worms | Expert curation with literature support | Public | [2] |
| CIS-BP | All Eukaryotes | Experimentally derived motifs and predictions | Public | [3] |
| CollecTF | Prokaryotes | Literature curation | Public | [4] |
| RegPrecise | Prokaryotes | Expert curation | Public | [5] |
| RegTransBase | Prokaryotes | Expert/literature curation | Public | [6] |
| RegulonDB | Escherichia coli | Expert curation | Public | [7] Archived 2017-05-07 at the Wayback Machine |
| PRODORIC | Prokaryotes | Expert curation | Public | [8] Archived 2007-05-16 at the Wayback Machine |
| TRANSFAC | Mammals | Expert/literature curation | Public/Private | [9] |
| TRED | Human, Mouse, Rat | Computer predictions, manual curation | Public | [10] |
| DBSD | Drosophila species | Literature/Expert curation | Public | [11] |
| HOCOMOCO | Human, Mouse | Literature/Expert curation | Public | [12],[13] |
| MethMotif | Human, Mouse | Expert curation | Public | [14] |
Representation of DNA binding sites[edit]
A collection of DNA binding sites, typically referred to as a DNA binding motif, can be represented by a consensus sequence. This representation has the advantage of being compact, but at the expense of disregarding a substantial amount of information.[15] A more accurate way of representing binding sites is through Position Specific Frequency Matrices (PSFM). These matrices give information on the frequency of each base at each position of the DNA binding motif.[3] PSFM are usually conceived with the implicit assumption of positional independence (different positions at the DNA binding site contribute independently to the site function), although this assumption has been disputed for some DNA binding sites.[16] Frequency information in a PSFM can be formally interpreted under the framework of Information Theory,[17] leading to its graphical representation as a sequence logo.
| 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | 10 | 11 | 12 | 13 | 14 | 15 | 16 | |
| A | 1 | 0 | 1 | 5 | 32 | 5 | 35 | 23 | 34 | 14 | 43 | 13 | 34 | 4 | 52 | 3 |
| C | 50 | 1 | 0 | 1 | 5 | 6 | 0 | 4 | 4 | 13 | 3 | 8 | 17 | 51 | 2 | 0 |
| G | 0 | 0 | 54 | 15 | 5 | 5 | 12 | 2 | 7 | 1 | 1 | 3 | 1 | 0 | 1 | 52 |
| T | 5 | 55 | 1 | 35 | 14 | 40 | 9 | 27 | 11 | 28 | 9 | 32 | 4 | 1 | 1 | 1 |
| Sum | 56 | 56 | 56 | 56 | 56 | 56 | 56 | 56 | 56 | 56 | 56 | 56 | 56 | 56 | 56 | 56 |
PSFM for the transcriptional repressor LexA as derived from 56 LexA-binding sites stored in Prodoric. Relative frequencies are obtained by dividing the counts in each cell by the total count (56)
Computational search and discovery of binding sites[edit]
In bioinformatics, one can distinguish between two separate problems regarding DNA binding sites: searching for additional members of a known DNA binding motif (the site search problem) and discovering novel DNA binding motifs in collections of functionally related sequences (the sequence motif discovery problem).[18] Many different methods have been proposed to search for binding sites. Most of them rely on the principles of information theory and have available web servers (Yellaboina)(Munch), while other authors have resorted to machine learning methods, such as artificial neural networks.[3][19][20] A plethora of algorithms is also available for sequence motif discovery. These methods rely on the hypothesis that a set of sequences share a binding motif for functional reasons. Binding motif discovery methods can be divided roughly into enumerative, deterministic and stochastic.[21] MEME[22] and Consensus [23] are classical examples of deterministic optimization, while the Gibbs sampler[24] is the conventional implementation of a purely stochastic method for DNA binding motif discovery. Another instance of this class of methods is SeSiMCMC[25] that is focused of weak TFBS sites with symmetry. While enumerative methods often resort to regular expression representation of binding sites, PSFM and their formal treatment under Information Theory methods are the representation of choice for both deterministic and stochastic methods. Hybrid methods, e.g. ChIPMunk[26] that combines greedy optimization with subsampling, also use PSFM. Recent advances in sequencing have led to the introduction of comparative genomics approaches to DNA binding motif discovery, as exemplified by PhyloGibbs.[27][28]
More complex methods for binding site search and motif discovery rely on the base stacking and other interactions between DNA bases, but due to the small sample sizes typically available for binding sites in DNA, their efficiency is still not completely harnessed. An example of such tool is the ULPB[29]
See also[edit]
References[edit]
- ^ Halford E.S.; Marko J.F. (2004). "How do site-specific DNA-binding proteins find their targets?". Nucleic Acids Research. 32 (10): 3040–3052. doi:10.1093/nar/gkh624. PMC 434431. PMID 15178741.
- ^ Borneman, A.R.; Gianoulis, T.A.; Zhang, Z.D.; Yu, H.; Rozowsky, J.; Seringhaus, M.R.; Wang, L.Y.; Gerstein, M. & Snyder, M. (2007). "Divergence of transcription factor binding sites across related yeast species". Science. 317 (5839): 815–819. Bibcode:2007Sci...317..815B. doi:10.1126/science.1140748. PMID 17690298. S2CID 21535866.
- ^ a b c Stormo GD (2000). "DNA binding sites: representation and discovery". Bioinformatics. 16 (1): 16–23. doi:10.1093/bioinformatics/16.1.16. PMID 10812473.
- ^ Pingoud A, Jeltsch A (1997). "Recognition and Cleavage of DNA by Type-II Restriction Endonucleases". European Journal of Biochemistry. 246 (1): 1–22. doi:10.1111/j.1432-1033.1997.t01-6-00001.x. PMID 9210460.
- ^ Gyohda A, Komano T (2000). "Purification and characterization of the R64 shufflon-specific recombinase". Journal of Bacteriology. 182 (10): 2787–2792. doi:10.1128/JB.182.10.2787-2792.2000. PMC 101987. PMID 10781547.
- ^ Birge, E.A. (2006). "15: Site Specific Recombination". Bacterial and Bacteriophage Genetics (5th ed.). Springer. pp. 463–478. ISBN 978-0-387-23919-4.
- ^ Campbell A (1963). "Fine Structure Genetics and its Relation to Function". Annual Review of Microbiology. 17 (1): 2787–2792. doi:10.1146/annurev.mi.17.100163.000405. PMID 14145311.
- ^ Jacob F, Monod J (1961). "Genetic regulatory mechanisms in the synthesis of proteins". Journal of Molecular Biology. 3 (3): 318–356. doi:10.1016/S0022-2836(61)80072-7. PMID 13718526. S2CID 19804795.
- ^ Gilbert W, Maxam A (1973). "The nucleotide sequence of the lac operator". Proceedings of the National Academy of Sciences of the United States of America. 70 (12): 3581–3584. Bibcode:1973PNAS...70.3581G. doi:10.1073/pnas.70.12.3581. PMC 427284. PMID 4587255.
- ^ Maniatis T, Ptashne M, Barrell BG, Donelson J (1974). "Sequence of a repressor-binding site in the DNA of bacteriophage lambda". Nature. 250 (465): 394–397. Bibcode:1974Natur.250..394M. doi:10.1038/250394a0. PMID 4854243. S2CID 4204720.
- ^ Nash H. A. (1975). "Integrative recombination of bacteriophage lambda DNA in vitro". Proceedings of the National Academy of Sciences of the United States of America. 72 (3): 1072–1076. Bibcode:1975PNAS...72.1072N. doi:10.1073/pnas.72.3.1072. PMC 432468. PMID 1055366.
- ^ Elnitski L, Jin VX, Farnham PJ, Jones SJ (2006). "Locating mammalian transcription factor binding sites: a survey of computational and experimental techniques". Genome Research. 16 (12): 1455–1464. doi:10.1101/gr.4140006. PMID 17053094.
- ^ Baaske P, Wienken CJ, Reineck P, Duhr S, Braun D (Feb 2010). "Optical Thermophoresis quantifies Buffer dependence of Aptamer Binding". Angew. Chem. Int. Ed. 49 (12): 2238–41. doi:10.1002/anie.200903998. PMID 20186894. S2CID 42489892.
- "A hot road to new drugs". Phys.org. February 24, 2010.
- ^ Wienken CJ; et al. (2010). "Protein-binding assays in biological liquids using microscale thermophoresis". Nature Communications. 1 (7): 100. Bibcode:2010NatCo...1..100W. doi:10.1038/ncomms1093. PMID 20981028.
- ^ Schneider T.D. (2002). "Consensus sequence Zen". Applied Bioinformatics. 1 (3): 111–119. PMC 1852464. PMID 15130839.
- ^ Bulyk M.L.; Johnson P.L.; Church G.M. (2002). "Nucleotides of transcription factor binding sites exert interdependent effects on the binding affinities of transcription factors". Nucleic Acids Research. 30 (5): 1255–1261. doi:10.1093/nar/30.5.1255. PMC 101241. PMID 11861919.
- ^ Schneider TD, Stormo GD, Gold L, Ehrenfeucht A (1986). "Information content of binding sites on nucleotide sequences". Journal of Molecular Biology. 188 (3): 415–431X. doi:10.1016/0022-2836(86)90165-8. PMID 3525846.
- ^ Erill I; O'Neill MC (2009). "A reexamination of information theory-based methods for DNA-binding site identification". BMC Bioinformatics. 10 (1): 57. doi:10.1186/1471-2105-10-57. PMC 2680408. PMID 19210776.
- ^ Bisant D, Maizel J (1995). "Identification of ribosome binding sites in Escherichia coli using neural network models". Nucleic Acids Research. 23 (9): 1632–1639. doi:10.1093/nar/23.9.1632. PMC 306908. PMID 7784221.
- ^ O'Neill M.C. (1991). "Training back-propagation neural networks to define and detect DNA-binding sites". Nucleic Acids Research. 19 (2): 133–318. doi:10.1093/nar/19.2.313. PMC 333596. PMID 2014171.
- ^ Bailey T.L. (2008). "Discovering Sequence Motifs". Bioinformatics (PDF). Methods in Molecular Biology. Vol. 452. pp. 231–251. doi:10.1007/978-1-60327-159-2_12. ISBN 978-1-58829-707-5. PMID 18566768.
- ^ Bailey T.L. (2002). "Discovering novel sequence motifs with MEME". Current Protocols in Bioinformatics. 2 (4): 2.4.1–2.4.35. doi:10.1002/0471250953.bi0204s00. PMID 18792935. S2CID 205157795.
- ^ Stormo GD, Hartzell GW 3rd (1989). "Identifying protein-binding sites from unaligned DNA fragments". Proceedings of the National Academy of Sciences of the United States of America. 86 (4): 1183–1187. Bibcode:1989PNAS...86.1183S. doi:10.1073/pnas.86.4.1183. PMC 286650. PMID 2919167.
- ^ Lawrence CE, Altschul SF, Boguski MS, Liu JS, Neuwald AF, Wootton JC (1993). "Detecting subtle sequence signals: a Gibbs sampling strategy for multiple alignment". Science. 262 (5131): 208–214. Bibcode:1993Sci...262..208L. doi:10.1126/science.8211139. PMID 8211139. S2CID 3040614.
- ^ Favorov, A V; M S Gelfand; A V Gerasimova; D A Ravcheev; A A Mironov; V J Makeev (2005-05-15). "A Gibbs sampler for identification of symmetrically structured, spaced DNA motifs with improved estimation of the signal length". Bioinformatics. 21 (10): 2240–2245. doi:10.1093/bioinformatics/bti336. ISSN 1367-4803. PMID 15728117.
- ^ Kulakovskiy, I V; V A Boeva; A V Favorov; V J Makeev (2010-08-24). "Deep and wide digging for binding motifs in ChIP-Seq data". Bioinformatics. 26 (20): 2622–3. doi:10.1093/bioinformatics/btq488. ISSN 1367-4811. PMID 20736340.
- ^ Das MK, Dai HK (2007). "A survey of DNA motif finding algorithms". BMC Bioinformatics. 8 (Suppl 7): S21. doi:10.1186/1471-2105-8-S7-S21. PMC 2099490. PMID 18047721.
- ^ Siddharthan R, Siggia ED, van Nimwegen E (2005). "PhyloGibbs: A Gibbs sampling motif finder that incorporates phylogeny". PLOS Comput Biol. 1 (7): e67. Bibcode:2005PLSCB...1...67S. doi:10.1371/journal.pcbi.0010067. PMC 1309704. PMID 16477324.
- ^ Salama RA, Stekel DJ (2010). "Inclusion of neighboring base interdependencies substantially improves genome-wide prokaryotic transcription factor binding site prediction". Nucleic Acids Research. 38 (12): e135. doi:10.1093/nar/gkq274. PMC 2896541. PMID 20439311.
External links[edit]
- ENCODE threads Explorer Transcription factor motifs in Nature
- Manually Curated TF Binding Motifs for 157 plant species
