Phyllosphere: Difference between revisions
Plantsurfer (talk | contribs) Adding local short description: "The plant surface as a habitat for microorganisms", overriding Wikidata description "The phyllosphere" (Shortdesc helper) |
Epipelagic (talk | contribs) add copy from plant microbiome – see that page's history for attribution |
||
| Line 1: | Line 1: | ||
{{short description|The plant surface as a habitat for microorganisms}} |
{{short description|The plant surface as a habitat for microorganisms}} |
||
[[File:Healthy and unhealthy leaf.png|thumb|upright=1|right| A leaf from a healthy ''[[Arabidopsis]]'' plant (left) and a leaf from a dysbiosis mutant plant (right)<ref name=He2020>[[Sheng-Yang He|He, Sheng Yang]] (2020) [https://theconversation.com/when-plants-and-their-microbes-are-not-in-sync-the-results-can-be-disastrous-143802 When plants and their microbes are not in sync, the results can be disastrous] ''The Conversation'', 28 August 2020.</ref>]] |
|||
{{Refimprove|date=September 2014}} |
|||
The '''phyllosphere''' is a term used in [[microbiology]] to refer to the total above-ground portions of plants as habitat for [[microorganisms]].<ref name=Last1955/> |
The '''phyllosphere''' is a term used in [[microbiology]] to refer to the total above-ground portions of plants as habitat for [[microorganisms]].<ref name=Last1955/> |
||
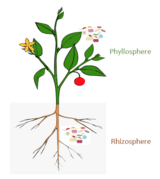
<ref name=Ruinen1956/> The phyllosphere can be further subdivided into the '''caulosphere''' (stems), '''phylloplane''' (leaves), '''anthosphere''' (flowers), and '''carposphere''' (fruits). The below-ground microbial habitats (i.e. the thin-volume of soil surrounding root or subterranean stem surfaces) are referred to as the [[rhizosphere]] and [[laimosphere]]. |
<ref name=Ruinen1956/> The phyllosphere can be further subdivided into the '''caulosphere''' (stems), '''phylloplane''' (leaves), '''anthosphere''' (flowers), and '''carposphere''' (fruits). The below-ground microbial habitats (i.e. the thin-volume of soil surrounding root or subterranean stem surfaces) are referred to as the [[rhizosphere]] and [[laimosphere]]. |
||
Most plants host diverse communities of microorganisms including [[bacteria]], [[fungi]], [[archaea]], and [[protists]] . Some are beneficial to the plant, others function as [[plant pathogen]]s and may damage the host plant or even kill it. However, the majority of microbial colonists on any given plant have no detectable effect on plant growth or function. |
Most plants host diverse communities of microorganisms including [[bacteria]], [[fungi]], [[archaea]], and [[protists]] . Some are beneficial to the plant, others function as [[plant pathogen]]s and may damage the host plant or even kill it. However, the majority of microbial colonists on any given plant have no detectable effect on plant growth or function. |
||
==The phyllosphere microbiome== |
|||
Research into the characteristics of microbial life in the phyllosphere is of great commercial importance to the agricultural industry for two reasons. First, understanding the survival of plant disease-causing bacteria and fungi is vital for developing new ways to control their spread. Second, there has been food poisoning cases associated with fruit and vegetables contaminated with bacteria, such as ''[[Salmonella]]'' and ''[[E. coli O157:H7]]''. This is particularly true of fresh fruits and salads which are not cooked prior to consumption. Preventing these outbreaks by developing better decontamination strategies is important to protect public health. For microorganisms the phyllosphere can be considered a hostile environment however, since there is a limited availability of nutrients, strong sun irradiation and variation in water availability.<ref>{{Cite journal|last=Berlec|first=Aleš|date=2012-09-01|title=Novel techniques and findings in the study of plant microbiota: Search for plant probiotics|journal=Plant Science|volume=193–194|pages=96–102|doi=10.1016/j.plantsci.2012.05.010|pmid=22794922}}</ref><ref>{{Cite journal|last=Lindow|first=Steven E|author1-link=Steven E. Lindow|last2=Leveau|first2=Johan H. J|date=2002-06-01|title=Phyllosphere microbiology|journal=Current Opinion in Biotechnology|volume=13|issue=3|pages=238–243|doi=10.1016/S0958-1669(02)00313-0}}</ref><ref>{{Cite journal|last=Bringel|first=Françoise|last2=Couée|first2=Ivan|date=2015-05-22|title=Pivotal roles of phyllosphere microorganisms at the interface between plant functioning and atmospheric trace gas dynamics|journal=Frontiers in Microbiology|volume=6|pages=486|doi=10.3389/fmicb.2015.00486|pmid=26052316|pmc=4440916}}</ref> |
|||
{{microbiomes|plant}} |
|||
The leaf surface, or phyllosphere, harbours a [[microbiome]] comprising diverse communities of [[bacteria]], [[archaea]], [[fungi]], [[algae]] and [[virus]]es.<ref name=Leveau2019>{{cite journal |doi = 10.1016/j.mib.2019.10.002|title = A brief from the leaf: Latest research to inform our understanding of the phyllosphere microbiome|year = 2019|last1 = Leveau|first1 = Johan HJ|journal = Current Opinion in Microbiology|volume = 49|pages = 41–49|pmid = 31707206}}</ref><ref>Ruinen, J. (1956) "Occurrence of Beijerinckia species in the 'phyllosphere'". ''Nature'', '''177'''(4501): 220–221.</ref> Microbial colonizers are subjected to diurnal and seasonal fluctuations of heat, moisture, and radiation. In addition, these environmental elements affect plant physiology (such as photosynthesis, respiration, water uptake etc.) and indirectly influence microbiome composition.<ref name=Dastogeer2020>Dastogeer, K.M., Tumpa, F.H., Sultana, A., Akter, M.A. and Chakraborty, A. (2020) "Plant microbiome–an account of the factors that shape community composition and diversity". ''Current Plant Biology'': 100161. {{doi|10.1016/j.cpb.2020.100161}}. [[File:CC-BY icon.svg|50px]] Material was copied from this source, which is available under a [https://creativecommons.org/licenses/by/4.0/ Creative Commons Attribution 4.0 International License].</ref> Rain and wind also cause temporal variation to the phyllosphere microbiome.<ref>{{cite book |doi = 10.1007/978-0-585-34164-4_10|chapter = Role of Immigration and Other Processes in Determining Epiphytic Bacterial Populations|title = Aerial Plant Surface Microbiology|year = 1996|last1 = Lindow|first1 = Steven E.|pages = 155–168|isbn = 978-0-306-45382-3}}</ref> |
|||
The phyllosphere includes the total [[aerial]] (above-ground) surface of a plant, and as such includes the surface of the stem, flowers and fruit, but most particularly the leaf surfaces. Compared with the [[rhizosphere]] and the [[endosphere]] the phyllosphere is nutrient poor and its environment more dynamic. |
|||
Interactions between plants and their associated microorganisms in many of these microbiomes can play pivotal roles in [[host plant]] health, function, and evolution.<ref>{{cite journal |doi = 10.1146/annurev-ecolsys-102710-145039|title = Microbially Mediated Plant Functional Traits|year = 2011|last1 = Friesen|first1 = Maren L.|last2 = Porter|first2 = Stephanie S.|last3 = Stark|first3 = Scott C.|last4 = von Wettberg|first4 = Eric J.|last5 = Sachs|first5 = Joel L.|last6 = Martinez-Romero|first6 = Esperanza|journal = Annual Review of Ecology, Evolution, and Systematics|volume = 42|pages = 23–46}}</ref> Interactions between the host plant and phyllosphere bacteria have the potential to drive various aspects of host plant physiology.<ref name=Vogel2016>{{cite journal |doi = 10.1111/nph.14036|title = The Arabidopsis leaf transcriptome reveals distinct but also overlapping responses to colonization by phyllosphere commensals and pathogen infection with impact on plant health|year = 2016|last1 = Vogel|first1 = Christine|last2 = Bodenhausen|first2 = Natacha|last3 = Gruissem|first3 = Wilhelm|last4 = Vorholt|first4 = Julia A.|journal = New Phytologist|volume = 212|pages = 192–207|url = https://www.zora.uzh.ch/id/eprint/131259/1/PMID27306148.pdf}}</ref><ref name=Ruinen1956>{{cite journal |doi = 10.1007/s00792-018-1015-x|title = Draft genome sequences of bacteria isolated from the Deschampsia antarctica phyllosphere|year = 2018|last1 = Cid|first1 = Fernanda P.|last2 = Maruyama|first2 = Fumito|last3 = Murase|first3 = Kazunori|last4 = Graether|first4 = Steffen P.|last5 = Larama|first5 = Giovanni|last6 = Bravo|first6 = Leon A.|last7 = Jorquera|first7 = Milko A.|journal = Extremophiles|volume = 22|issue = 3|pages = 537–552|pmid = 29492666|s2cid = 4320165}}</ref><ref>{{cite journal |doi = 10.1128/genomeA.00019-18|title = Draft Genome Sequence of Plant Growth-Promoting and Drought-Tolerant Bacillus altitudinis FD48, Isolated from Rice Phylloplane|year = 2018|last1 = Kumaravel|first1 = Sowmya|last2 = Thankappan|first2 = Sugitha|last3 = Raghupathi|first3 = Sridar|last4 = Uthandi|first4 = Sivakumar|journal = Genome Announcements|volume = 6|issue = 9|pmid = 29496824|pmc = 5834328}}</ref> However, as of 2020 knowledge of these bacterial associations in the phyllosphere remains relatively modest, and there is a need to advance fundamental knowledge of phyllosphere microbiome dynamics.<ref>{{cite journal |doi = 10.1002/ajb2.1229|title = Decrypting the phyllosphere microbiota: Progress and challenges|year = 2019|last1 = Laforest‐Lapointe|first1 = Isabelle|last2 = Whitaker|first2 = Briana K.|journal = American Journal of Botany|volume = 106|issue = 2|pages = 171–173|pmid = 30726571|doi-access = free}}</ref><ref name=Noble2020 /> |
|||
The assembly of the phyllosphere microbiome, which can be strictly defined as [[Epiphytic bacteria|epiphytic bacterial]] communities on the leaf surface, can be shaped by the [[microbial communities]] present in the surrounding environment (i.e., [[stochastic]] [[Colony (biology)|colonisation]]) and the host plant (i.e., [[Biotic component|biotic]] selection).<ref name=Leveau2019 /><ref>{{cite journal |doi = 10.1038/nrmicro2910|title = Microbial life in the phyllosphere|year = 2012|last1 = Vorholt|first1 = Julia A.|journal = Nature Reviews Microbiology|volume = 10|issue = 12|pages = 828–840|pmid = 23154261|s2cid = 10447146}}</ref><ref name=Noble2020 /> However, although the leaf surface is generally considered a discrete microbial habitat,<ref name=Stone2014>{{cite journal |doi = 10.1007/s00248-016-0738-4|title = Biogeographic Patterns Between Bacterial Phyllosphere Communities of the Southern Magnolia (Magnolia grandiflora) in a Small Forest|year = 2016|last1 = Stone|first1 = Bram W. G.|last2 = Jackson|first2 = Colin R.|journal = Microbial Ecology|volume = 71|issue = 4|pages = 954–961|pmid = 26883131|s2cid = 17292307}}</ref><ref name= Redford2010>{{cite journal |doi = 10.1111/j.1462-2920.2010.02258.x|title = The ecology of the phyllosphere: Geographic and phylogenetic variability in the distribution of bacteria on tree leaves|year = 2010|last1 = Redford|first1 = Amanda J.|last2 = Bowers|first2 = Robert M.|last3 = Knight|first3 = Rob|last4 = Linhart|first4 = Yan|last5 = Fierer|first5 = Noah|journal = Environmental Microbiology|volume = 12|issue = 11|pages = 2885–2893|pmid = 20545741|pmc = 3156554}}</ref> there is no consensus on the dominant driver of community assembly across phyllosphere microbiomes. For example, host-specific bacterial communities have been reported in the phyllosphere of co-occurring plant species, suggesting a dominant role of host selection.<ref name= Redford2010 /><ref>{{cite journal |doi = 10.1007/s00248-012-0053-7|title = Exploring Biodiversity in the Bacterial Community of the Mediterranean Phyllosphere and its Relationship with Airborne Bacteria|year = 2012|last1 = Vokou|first1 = Despoina|last2 = Vareli|first2 = Katerina|last3 = Zarali|first3 = Ekaterini|last4 = Karamanoli|first4 = Katerina|last5 = Constantinidou|first5 = Helen-Isis A.|last6 = Monokrousos|first6 = Nikolaos|last7 = Halley|first7 = John M.|last8 = Sainis|first8 = Ioannis|journal = Microbial Ecology|volume = 64|issue = 3|pages = 714–724|pmid = 22544345|s2cid = 17291303}}</ref><ref name=Laforest-Lapointe2016>{{cite journal |doi = 10.1186/s40168-016-0174-1|title = Host species identity, site and time drive temperate tree phyllosphere bacterial community structure|year = 2016|last1 = Laforest-Lapointe|first1 = Isabelle|last2 = Messier|first2 = Christian|last3 = Kembel|first3 = Steven W.|journal = Microbiome|volume = 4|issue = 1|page = 27|pmid = 27316353|pmc = 4912770}}</ref><ref name=Noble2020 /> |
|||
Conversely, microbiomes of the surrounding environment have also been reported to be the primary determinant of phyllosphere community composition.<ref name=Stone2014 /><ref>{{cite journal |doi = 10.1128/mBio.02527-14|title = The Soil Microbiome Influences Grapevine-Associated Microbiota|year = 2015|last1 = Zarraonaindia|first1 = Iratxe|last2 = Owens|first2 = Sarah M.|last3 = Weisenhorn|first3 = Pamela|last4 = West|first4 = Kristin|last5 = Hampton-Marcell|first5 = Jarrad|last6 = Lax|first6 = Simon|last7 = Bokulich|first7 = Nicholas A.|last8 = Mills|first8 = David A.|last9 = Martin|first9 = Gilles|last10 = Taghavi|first10 = Safiyh|last11 = Van Der Lelie|first11 = Daniel|last12 = Gilbert|first12 = Jack A.|journal = mBio|volume = 6|issue = 2|pmid = 25805735|pmc = 4453523}}</ref><ref>{{cite journal |doi = 10.1128/AEM.05565-11|title = Geographical Location Determines the Population Structure in Phyllosphere Microbial Communities of a Salt-Excreting Desert Tree|year = 2011|last1 = Finkel|first1 = Omri M.|last2 = Burch|first2 = Adrien Y.|last3 = Lindow|first3 = Steven E.|last4 = Post|first4 = Anton F.|last5 = Belkin|first5 = Shimshon|journal = Applied and Environmental Microbiology|volume = 77|issue = 21|pages = 7647–7655|pmid = 21926212|pmc = 3209174}}</ref><ref>{{cite journal |doi = 10.1128/AEM.00888-12|title = Distance-Decay Relationships Partially Determine Diversity Patterns of Phyllosphere Bacteria on Tamrix Trees across the Sonoran Desert|year = 2012|last1 = Finkel|first1 = Omri M.|last2 = Burch|first2 = Adrien Y.|last3 = Elad|first3 = Tal|last4 = Huse|first4 = Susan M.|last5 = Lindow|first5 = Steven E.|last6 = Post|first6 = Anton F.|last7 = Belkin|first7 = Shimshon|journal = Applied and Environmental Microbiology|volume = 78|issue = 17|pages = 6187–6193|pmid = 22752165|pmc = 3416633}}</ref> As a result, the processes that drive phyllosphere community assembly are not well understood but unlikely to be universal across plant species. However, the existing evidence does indicate that phyllosphere microbiomes exhibiting host-specific associations are more likely to interact with the host than those primarily recruited from the surrounding environment.<ref name=Vogel2016 /><ref name=Kembel2014>{{cite journal |doi = 10.1073/pnas.1216057111|title = Relationships between phyllosphere bacterial communities and plant functional traits in a neotropical forest|year = 2014|last1 = Kembel|first1 = S. W.|last2 = O'Connor|first2 = T. K.|last3 = Arnold|first3 = H. K.|last4 = Hubbell|first4 = S. P.|last5 = Wright|first5 = S. J.|last6 = Green|first6 = J. L.|journal = Proceedings of the National Academy of Sciences|volume = 111|issue = 38|pages = 13715–13720|bibcode = 2014PNAS..11113715K|s2cid = 852584|doi-access = free}}</ref><ref>{{cite journal |doi = 10.1128/AEM.00133-11|title = Protection of Arabidopsis thaliana against Leaf-Pathogenic Pseudomonas syringae by Sphingomonas Strains in a Controlled Model System|year = 2011|last1 = Innerebner|first1 = Gerd|last2 = Knief|first2 = Claudia|last3 = Vorholt|first3 = Julia A.|journal = Applied and Environmental Microbiology|volume = 77|issue = 10|pages = 3202–3210|pmid = 21421777|pmc = 3126462}}</ref><ref>{{cite journal |doi = 10.1186/s40168-020-00844-7|title = Adaptive matching between phyllosphere bacteria and their tree hosts in a neotropical forest|year = 2020|last1 = Lajoie|first1 = Geneviève|last2 = Maglione|first2 = Rémi|last3 = Kembel|first3 = Steven W.|journal = Microbiome|volume = 8|issue = 1|page = 70|pmid = 32438916|pmc = 7243311}}</ref><ref name=Noble2020 /> |
|||
Overall, there remains high species richness in phyllosphere communities. Fungal communities are highly variable in the phyllosphere of temperate regions and are more diverse than in tropical regions.<ref name=Vorholt2012>{{cite journal |doi = 10.1128/AEM.05565-11|title = Geographical Location Determines the Population Structure in Phyllosphere Microbial Communities of a Salt-Excreting Desert Tree|year = 2011|last1 = Finkel|first1 = Omri M.|last2 = Burch|first2 = Adrien Y.|last3 = Lindow|first3 = Steven E.|last4 = Post|first4 = Anton F.|last5 = Belkin|first5 = Shimshon|journal = Applied and Environmental Microbiology|volume = 77|issue = 21|pages = 7647–7655|pmid = 21926212|pmc = 3209174}}</ref> There can be up to 107 microbes per square centimetre present on the leaf surfaces of plants, and the bacterial population of the phyllosphere on a global scale is estimated to be 10<sup>26</sup> cells.<ref>{{cite journal |doi = 10.1038/nrmicro2910|title = Microbial life in the phyllosphere|year = 2012|last1 = Vorholt|first1 = Julia A.|journal = Nature Reviews Microbiology|volume = 10|issue = 12|pages = 828–840|pmid = 23154261|s2cid = 10447146}}</ref> The population size of the fungal phyllosphere is likely to be smaller.<ref>{{cite journal |doi = 10.1128/AEM.69.4.1875-1883.2003|title = Microbiology of the Phyllosphere|year = 2003|last1 = Lindow|first1 = Steven E.|last2 = Brandl|first2 = Maria T.|journal = Applied and Environmental Microbiology|volume = 69|issue = 4|pages = 1875–1883|pmid = 12676659|pmc = 154815|s2cid = 2304379}}</ref> |
|||
Phyllosphere microbes from different plants appear to be somewhat similar at high levels of taxa, but at the lower levels taxa there remain significant differences. This indicates microorganisms may need finely tuned metabolic adjustment to survive in phyllosphere environment.<ref name=Vorholt2012 /> [[Proteobacteria]] seems to be the dominant colonizers, with [[Bacteroidetes]] and [[Actinobacteria]] also predominant in phyllospheres.<ref>{{cite journal |doi = 10.1371/journal.pone.0056329|title = Bacterial Communities Associated with the Leaves and the Roots of Arabidopsis thaliana|year = 2013|last1 = Bodenhausen|first1 = Natacha|last2 = Horton|first2 = Matthew W.|last3 = Bergelson|first3 = Joy|journal = PLOS ONE|volume = 8|issue = 2|pages = e56329|pmid = 23457551|pmc = 3574144}}</ref> Although there are similarities between the rhizosphere and soil microbial communities, very little similarity has been found between phyllosphere communities and microorganisms floating in open air ([[aeroplankton]]).<ref>{{cite journal |doi = 10.1007/s00248-012-0053-7|title = Exploring Biodiversity in the Bacterial Community of the Mediterranean Phyllosphere and its Relationship with Airborne Bacteria|year = 2012|last1 = Vokou|first1 = Despoina|last2 = Vareli|first2 = Katerina|last3 = Zarali|first3 = Ekaterini|last4 = Karamanoli|first4 = Katerina|last5 = Constantinidou|first5 = Helen-Isis A.|last6 = Monokrousos|first6 = Nikolaos|last7 = Halley|first7 = John M.|last8 = Sainis|first8 = Ioannis|journal = Microbial Ecology|volume = 64|issue = 3|pages = 714–724|pmid = 22544345|s2cid = 17291303}}</ref><ref name=Dastogeer2020 /> |
|||
The search for a core microbiome in host-associated microbial communities is a useful first step in trying to understand the interactions that may be occurring between a host and its microbiome.<ref name= Shade2012>{{cite journal |doi = 10.1111/j.1462-2920.2011.02585.x |title = Beyond the Venn diagram: The hunt for a core microbiome |year = 2012 |last1 = Shade |first1 = Ashley |last2 = Handelsman |first2 = Jo |journal = Environmental Microbiology |volume = 14 |issue = 1 |pages = 4–12 |pmid = 22004523 |doi-access = free }}</ref><ref>{{cite journal |doi = 10.1186/s40168-020-00875-0|title = Microbiome definition re-visited: Old concepts and new challenges|year = 2020|last1 = Berg|first1 = Gabriele|last2 = Rybakova|first2 = Daria|last3 = Fischer|first3 = Doreen|last4 = Cernava|first4 = Tomislav|last5 = Vergès|first5 = Marie-Christine Champomier|last6 = Charles|first6 = Trevor|last7 = Chen|first7 = Xiaoyulong|last8 = Cocolin|first8 = Luca|last9 = Eversole|first9 = Kellye|last10 = Corral|first10 = Gema Herrero|last11 = Kazou|first11 = Maria|last12 = Kinkel|first12 = Linda|last13 = Lange|first13 = Lene|last14 = Lima|first14 = Nelson|last15 = Loy|first15 = Alexander|last16 = MacKlin|first16 = James A.|last17 = Maguin|first17 = Emmanuelle|last18 = Mauchline|first18 = Tim|last19 = McClure|first19 = Ryan|last20 = Mitter|first20 = Birgit|last21 = Ryan|first21 = Matthew|last22 = Sarand|first22 = Inga|last23 = Smidt|first23 = Hauke|last24 = Schelkle|first24 = Bettina|last25 = Roume|first25 = Hugo|last26 = Kiran|first26 = G. Seghal|last27 = Selvin|first27 = Joseph|last28 = Souza|first28 = Rafael Soares Correa de|last29 = Van Overbeek|first29 = Leo|last30 = Singh|first30 = Brajesh K.|journal = Microbiome|volume = 8|issue = 1|page = 103|pmid = 32605663|pmc = 7329523|display-authors = 29}}</ref> The prevailing core microbiome concept is built on the notion that the persistence of a taxon across the spatiotemporal boundaries of an ecological niche is directly reflective of its functional importance within the niche it occupies; it therefore provides a framework for identifying functionally critical microorganisms that consistently associate with a host species.<ref name= Shade2012 /><ref>{{cite journal |doi = 10.1038/nature07540|title = A core gut microbiome in obese and lean twins|year = 2009|last1 = Turnbaugh|first1 = Peter J.|last2 = Hamady|first2 = Micah|last3 = Yatsunenko|first3 = Tanya|last4 = Cantarel|first4 = Brandi L.|last5 = Duncan|first5 = Alexis|last6 = Ley|first6 = Ruth E.|last7 = Sogin|first7 = Mitchell L.|last8 = Jones|first8 = William J.|last9 = Roe|first9 = Bruce A.|last10 = Affourtit|first10 = Jason P.|last11 = Egholm|first11 = Michael|last12 = Henrissat|first12 = Bernard|last13 = Heath|first13 = Andrew C.|last14 = Knight|first14 = Rob|last15 = Gordon|first15 = Jeffrey I.|journal = Nature|volume = 457|issue = 7228|pages = 480–484|pmid = 19043404|pmc = 2677729|bibcode = 2009Natur.457..480T}}</ref><ref>{{cite journal |doi = 10.1038/nature11237|title = Defining the core Arabidopsis thaliana root microbiome|year = 2012|last1 = Lundberg|first1 = Derek S.|last2 = Lebeis|first2 = Sarah L.|last3 = Paredes|first3 = Sur Herrera|last4 = Yourstone|first4 = Scott|last5 = Gehring|first5 = Jase|last6 = Malfatti|first6 = Stephanie|last7 = Tremblay|first7 = Julien|last8 = Engelbrektson|first8 = Anna|last9 = Kunin|first9 = Victor|last10 = Rio|first10 = Tijana Glavina del|last11 = Edgar|first11 = Robert C.|last12 = Eickhorst|first12 = Thilo|last13 = Ley|first13 = Ruth E.|last14 = Hugenholtz|first14 = Philip|last15 = Tringe|first15 = Susannah Green|last16 = Dangl|first16 = Jeffery L.|journal = Nature|volume = 488|issue = 7409|pages = 86–90|pmid = 22859206|pmc = 4074413|bibcode = 2012Natur.488...86L}}</ref><ref name=Noble2020 /> |
|||
Divergent definitions of “core microbiome” have arisen across scientific literature with researchers variably identifying “core taxa” as those persistent across distinct host microhabitats{{hsp}}<ref>{{cite journal |doi = 10.1111/1462-2920.14031|title = Field study reveals core plant microbiota and relative importance of their drivers|year = 2018|last1 = Hamonts|first1 = Kelly|last2 = Trivedi|first2 = Pankaj|last3 = Garg|first3 = Anshu|last4 = Janitz|first4 = Caroline|last5 = Grinyer|first5 = Jasmine|last6 = Holford|first6 = Paul|last7 = Botha|first7 = Frederik C.|last8 = Anderson|first8 = Ian C.|last9 = Singh|first9 = Brajesh K.|journal = Environmental Microbiology|volume = 20|issue = 1|pages = 124–140|pmid = 29266641|s2cid = 10650949|doi-access = free}}</ref><ref>{{cite journal |doi = 10.1186/s40168-019-0624-7|title = Enterobacteriaceae dominate the core microbiome and contribute to the resistome of arugula (Eruca sativa Mill.)|year = 2019|last1 = Cernava|first1 = Tomislav|last2 = Erlacher|first2 = Armin|last3 = Soh|first3 = Jung|last4 = Sensen|first4 = Christoph W.|last5 = Grube|first5 = Martin|last6 = Berg|first6 = Gabriele|journal = Microbiome|volume = 7|issue = 1|page = 13|pmid = 30696492|pmc = 6352427}}</ref> and even different species.<ref name=Laforest-Lapointe2016 /><ref name=Kembel2014 /> Given the functional divergence of microorganisms across different host species{{hsp}}<ref name=Laforest-Lapointe2016 /> and microhabitats,<ref>{{cite journal |doi = 10.1111/1462-2920.12695|title = Spatial structuring of bacterial communities within individual ''Ginkgo'' bilobatrees|year = 2015|last1 = Leff|first1 = Jonathan W.|last2 = Del Tredici|first2 = Peter|last3 = Friedman|first3 = William E.|last4 = Fierer|first4 = Noah|journal = Environmental Microbiology|volume = 17|issue = 7|pages = 2352–2361|pmid = 25367625}}</ref> defining core taxa ''sensu stricto'' as those persistent across broad geographic distances within tissue- and species-specific host microbiomes, represents the most biologically and ecologically appropriate application of this conceptual framework.<ref>{{cite journal |doi = 10.1016/j.tim.2016.11.003|title = Defining the Core Microbiome in Corals' Microbial Soup|year = 2017|last1 = Hernandez-Agreda|first1 = Alejandra|last2 = Gates|first2 = Ruth D.|last3 = Ainsworth|first3 = Tracy D.|journal = Trends in Microbiology|volume = 25|issue = 2|pages = 125–140|pmid = 27919551}}</ref><ref name=Noble2020 /> Tissue- and species-specific core microbiomes across host populations separated by broad geographical distances have not been widely reported for the phyllosphere using the stringent definition established by Ruinen.<ref name=Ruinen1956 /><ref name=Noble2020 /> |
|||
==Example: The mānuka phyllosphere== |
|||
{{multiple image |
|||
| align = left |
|||
| direction = horizontal |
|||
| header = Relative abundance of core phyllosphere taxa in mānuka |
|||
| header_align = <!-- left/right/center --> |
|||
| header_background = |
|||
| footer = [[Mānuka]] is a flowering scrub. The chart shows an abundance-occupancy distribution identifying core phyllosphere taxa in non-rarefied (green) and rarefied (purple) datasets. Each point represents a taxon plotted by its mean logarithmic relative abundance and occupancy. Taxa (pink) with an occupancy of 1 (i.e., detected in all 89 phyllosphere samples) were considered members of the core microbiome.<ref name=Noble2020>{{cite journal |doi = 10.1371/journal.pone.0237079|title = A core phyllosphere microbiome exists across distant populations of a tree species indigenous to New Zealand|year = 2020|last1 = Noble|first1 = Anya S.|last2 = Noe|first2 = Stevie|last3 = Clearwater|first3 = Michael J.|last4 = Lee|first4 = Charles K.|journal = PLOS ONE|volume = 15|issue = 8|pages = e0237079|pmid = 32790769|pmc = 7425925|bibcode = 2020PLoSO..1537079N}} [[File:CC-BY icon.svg|50px]] Material was copied from this source, which is available under a [https://creativecommons.org/licenses/by/4.0/ Creative Commons Attribution 4.0 International License]. [[File:CC-BY icon.svg|50px]] Material was copied from this source, which is available under a [https://creativecommons.org/licenses/by/4.0/ Creative Commons Attribution 4.0 International License].</ref> |
|||
| footer_align = left |
|||
| footer_background = |
|||
| background color = |
|||
| image1 = Starr-010419-0063-Leptospermum scoparium-habit-Kula-Maui (24236713820).jpg |
|||
| width1 = 198 |
|||
| alt1 = |
|||
| caption1 = |
|||
| image2 = Relative abundance of core phyllosphere taxa in the mānuka phyllosphere (crop).png |
|||
| width2 = 167 |
|||
| alt2 = |
|||
| caption2 = |
|||
}} |
|||
The flowering tea tree commonly known as [[mānuka]] is indigenous to New Zealand.<ref>{{cite journal |doi = 10.1080/0028825X.2005.9512966|title = A review of ''Leptospermum'' scoparium(Myrtaceae) in New Zealand|year = 2005|last1 = Stephens|first1 = J. M. C.|last2 = Molan|first2 = P. C.|last3 = Clarkson|first3 = B. D.|journal = New Zealand Journal of Botany|volume = 43|issue = 2|pages = 431–449|s2cid = 53515334}}</ref> [[Mānuka honey]], produced from the [[nectar]] of mānuka flowers, is known for its non-peroxide antibacterial properties.<ref>{{cite journal |doi = 10.1046/j.1365-2672.2002.01761.x|title = The sensitivity to honey of Gram-positive cocci of clinical significance isolated from wounds|year = 2002|last1 = Cooper|first1 = R.A.|last2 = Molan|first2 = P.C.|last3 = Harding|first3 = K.G.|journal = Journal of Applied Microbiology|volume = 93|issue = 5|pages = 857–863|pmid = 12392533|s2cid = 24517001|doi-access = free}}</ref><ref>{{cite journal |doi = 10.3109/01913123.2016.1154914|title = How methylglyoxal kills bacteria: An ultrastructural study|year = 2016|last1 = Rabie|first1 = Erika|last2 = Serem|first2 = June Cheptoo|last3 = Oberholzer|first3 = Hester Magdalena|last4 = Gaspar|first4 = Anabella Regina Marques|last5 = Bester|first5 = Megan Jean|journal = Ultrastructural Pathology|volume = 40|issue = 2|pages = 107–111|pmid = 26986806|hdl = 2263/52156|s2cid = 13372064|hdl-access = free}}</ref> These non-peroxide antibacterial properties have been principally linked to the accumulation of the three-carbon sugar [[dihydroxyacetone]] (DHA) in the nectar of the mānuka flower, which undergoes a chemical conversion to [[methylglyoxal]] (MGO) in mature honey.<ref>{{cite journal |doi = 10.1016/j.carres.2009.03.020 |title = The origin of methylglyoxal in New Zealand manuka (Leptospermum scoparium) honey |year = 2009 |last1 = Adams |first1 = Christopher J. |last2 = Manley-Harris |first2 = Merilyn |last3 = Molan |first3 = Peter C. |journal = Carbohydrate Research |volume = 344 |issue = 8 |pages = 1050–1053 |pmid = 19368902 }}</ref><ref>{{cite journal |doi = 10.1016/j.carres.2012.07.025|title = Studies on the formation of methylglyoxal from dihydroxyacetone in Manuka (Leptospermum scoparium) honey|year = 2012|last1 = Atrott|first1 = Julia|last2 = Haberlau|first2 = Steffi|last3 = Henle|first3 = Thomas|journal = Carbohydrate Research|volume = 361|pages = 7–11|pmid = 22960208}}</ref><ref>{{cite journal |doi = 10.1002/mnfr.200700282|title = Identification and quantification of methylglyoxal as the dominant antibacterial constituent of Manuka (Leptospermum scoparium)honeys from New Zealand|year = 2008|last1 = Mavric|first1 = Elvira|last2 = Wittmann|first2 = Silvia|last3 = Barth|first3 = Gerold|last4 = Henle|first4 = Thomas|journal = Molecular Nutrition & Food Research|volume = 52|issue = 4|pages = 483–489}}</ref> However, the concentration of DHA in the nectar of mānuka flowers is notoriously variable, and the antimicrobial efficacy of mānuka honey consequently varies from region to region and from year to year.<ref>Hamilton, G., Millner, J., Robertson, A. and Stephens, J. (2013) "Assessment of manuka provenances for production of high 'unique manuka factor' honey". ''Agronomy New Zealand'', '''43''': 139–144.</ref><ref>{{cite journal |doi = 10.1021/jf5045958|title = Regional, Annual, and Individual Variations in the Dihydroxyacetone Content of the Nectar of Ma̅nuka (Leptospermum scoparium) in New Zealand|year = 2014|last1 = Williams|first1 = Simon|last2 = King|first2 = Jessica|last3 = Revell|first3 = Maria|last4 = Manley-Harris|first4 = Merilyn|last5 = Balks|first5 = Megan|last6 = Janusch|first6 = Franziska|last7 = Kiefer|first7 = Michael|last8 = Clearwater|first8 = Michael|last9 = Brooks|first9 = Peter|last10 = Dawson|first10 = Murray|journal = Journal of Agricultural and Food Chemistry|volume = 62|issue = 42|pages = 10332–10340|pmid = 25277074}}</ref><ref>Stephens, J.M.C. (2006) "The factors responsible for the varying levels of UMF® in mānuka (''Leptospermum scoparium'') honey", Doctoral dissertation, University of Waikato.</ref> Despite extensive research efforts, no reliable correlation has been identified between DHA production and climatic,<ref>{{cite journal |doi = 10.1080/01140671.2019.1670681|title = Floral nectar of wild mānuka (Leptospermum scoparium) varies more among plants than among sites|year = 2019|last1 = Noe|first1 = Stevie|last2 = Manley-Harris|first2 = Merilyn|last3 = Clearwater|first3 = Michael J.|journal = New Zealand Journal of Crop and Horticultural Science|volume = 47|issue = 4|pages = 282–296|s2cid = 204143940}}</ref> [[edaphic]],<ref>{{cite journal |doi = 10.1080/0028825X.2016.1247732|title = Soil influences on plant growth, floral density and nectar yield in three cultivars of mānuka (Leptospermum scoparium)|year = 2017|last1 = Nickless|first1 = Elizabeth M.|last2 = Anderson|first2 = Christopher W. N.|last3 = Hamilton|first3 = Georgie|last4 = Stephens|first4 = Jonathan M.|last5 = Wargent|first5 = Jason|journal = New Zealand Journal of Botany|volume = 55|issue = 2|pages = 100–117|s2cid = 88657399}}</ref> or host genetic factors.<ref>{{cite journal |doi = 10.1093/aob/mcx183|title = Influence of genotype, floral stage, and water stress on floral nectar yield and composition of mānuka (Leptospermum scoparium)|year = 2018|last1 = Clearwater|first1 = Michael J.|last2 = Revell|first2 = Maria|last3 = Noe|first3 = Stevie|last4 = Manley-Harris|first4 = Merilyn|journal = Annals of Botany|volume = 121|issue = 3|pages = 501–512|pmid = 29300875|pmc = 5838834}}</ref><ref name=Noble2020 /> |
|||
[[File:Differential abundance of taxa in manuka phyllosphere.png|thumb|upright=1.7|right| {A} The [[heatmap]] on the left illustrates how the composition of OTUs in the mānuka phyllosphere and associated soil communities differed significantly. No core soil microbiome was detected.<br />(B) The chart on the right shows how OTUs in phyllosphere and associated soil communities differed in relative abundances.<ref name=Noble2020 />]] |
|||
{{clear left}} |
|||
Microorganisms have been studied in the mānuka rhizosphere and endosphere.<ref>{{cite journal |doi = 10.1017/S0953756297005765|title = Leaf endophytes of manuka (Leptospermum scoparium)|year = 1998|last1 = Johnston|first1 = Peter R.|journal = Mycological Research|volume = 102|issue = 8|pages = 1009–1016}}</ref><ref>{{cite journal |doi = 10.1080/0028825X.2006.9513025|title = Checklist of fungi on teatree (Kunzeaand ''Leptospermumspecies'') in New Zealand|year = 2006|last1 = McKenzie|first1 = E. H. C.|last2 = Johnston|first2 = P. R.|last3 = Buchanan|first3 = P. K.|journal = New Zealand Journal of Botany|volume = 44|issue = 3|pages = 293–335|s2cid = 84538904|doi-access = free}}</ref><ref>{{cite journal |doi = 10.1007/s13199-017-0506-3|title = Arbuscular mycorrhizal fungi associated with Leptospermum scoparium (Mānuka): Effects on plant growth and essential oil content|year = 2018|last1 = Wicaksono|first1 = Wisnu Adi|last2 = Sansom|first2 = Catherine E.|last3 = Eirian Jones|first3 = E.|last4 = Perry|first4 = Nigel B.|last5 = Monk|first5 = Jana|last6 = Ridgway|first6 = Hayley J.|journal = Symbiosis|volume = 75|pages = 39–50|s2cid = 4819178}}</ref> Earlier studies primarily focussed on fungi, and a 2016 study provided the first investigation of endophytic bacterial communities from three geographically and environmentally distinct mānuka populations using fingerprinting techniques and revealed tissue-specific core endomicrobiomes.<ref>{{cite journal |doi = 10.1371/journal.pone.0163717|title = The Bacterial Signature of Leptospermum scoparium (Mānuka) Reveals Core and Accessory Communities with Bioactive Properties|year = 2016|last1 = Wicaksono|first1 = Wisnu Adi|last2 = Jones|first2 = E. Eirian|last3 = Monk|first3 = Jana|last4 = Ridgway|first4 = Hayley J.|journal = PLOS ONE|volume = 11|issue = 9|pages = e0163717|pmid = 27676607|pmc = 5038978|bibcode = 2016PLoSO..1163717W}}</ref><ref name=Noble2020 /> A 2020 study identified a habitat-specific and relatively abundant core microbiome in the mānuka phyllosphere, which was persistent across all samples. In contrast, non-core phyllosphere microorganisms exhibited significant variation across individual host trees and populations that was strongly driven by environmental and spatial factors. The results demonstrated the existence of a dominant and ubiquitous core microbiome in the phyllosphere of mānuka.<ref name=Noble2020 /> |
|||
{{clear}} |
|||
==See also== |
|||
* [[Epiphytes]] |
|||
* [[Microbiota]] |
|||
==References== |
==References== |
||
{{ |
{{reflist|refs= |
||
<ref name=Last1955>{{cite journal|last=Last|first=F.T.|title=Seasonal incidence of Sporobolomyces on cereal leaves|journal=Trans Br Mycol Soc|year=1955|volume=38|issue=3|pages=221–239|doi=10.1016/s0007-1536(55)80069-1}}</ref> |
<ref name=Last1955>{{cite journal|last=Last|first=F.T.|title=Seasonal incidence of Sporobolomyces on cereal leaves|journal=Trans Br Mycol Soc|year=1955|volume=38|issue=3|pages=221–239|doi=10.1016/s0007-1536(55)80069-1}}</ref> |
||
<ref name=Ruinen1956>{{cite journal|last=Ruinen|first=J.|title=Occurrence of Beijerinckia species in the phyllosphere.|journal=Nature|year=1956|volume=178|issue=4501|pages=220–221|doi=10.1038/177220a0}}</ref> |
|||
}} |
}} |
||
<ref>{{cite journal |doi = 10.3389/fpls.2018.01482|title= Influence of Light on Plant–Phyllosphere Interaction}} [[File:CC-BY icon.svg|50px]] Material was copied from this source, which is available under a [https://creativecommons.org/licenses/by/4.0/ Creative Commons Attribution 4.0 International License].</ref> |
|||
== External links == |
|||
* [http://news.bbc.co.uk/1/hi/england/humber/3708820.stm A BBC news article about a salad-associated food poisoning outbreak] |
|||
* [https://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&list_uids=12180099&dopt=Abstract Link to one of many scholarly review articles about phyllosphere microbiology] |
|||
[[Category:Microbiology terms]] |
[[Category:Microbiology terms]] |
||
Revision as of 17:22, 25 February 2021

The phyllosphere is a term used in microbiology to refer to the total above-ground portions of plants as habitat for microorganisms.[2] [3] The phyllosphere can be further subdivided into the caulosphere (stems), phylloplane (leaves), anthosphere (flowers), and carposphere (fruits). The below-ground microbial habitats (i.e. the thin-volume of soil surrounding root or subterranean stem surfaces) are referred to as the rhizosphere and laimosphere. Most plants host diverse communities of microorganisms including bacteria, fungi, archaea, and protists . Some are beneficial to the plant, others function as plant pathogens and may damage the host plant or even kill it. However, the majority of microbial colonists on any given plant have no detectable effect on plant growth or function.
The phyllosphere microbiome
| Part of a series on |
| Microbiomes |
|---|
 |
The leaf surface, or phyllosphere, harbours a microbiome comprising diverse communities of bacteria, archaea, fungi, algae and viruses.[4][5] Microbial colonizers are subjected to diurnal and seasonal fluctuations of heat, moisture, and radiation. In addition, these environmental elements affect plant physiology (such as photosynthesis, respiration, water uptake etc.) and indirectly influence microbiome composition.[6] Rain and wind also cause temporal variation to the phyllosphere microbiome.[7]
The phyllosphere includes the total aerial (above-ground) surface of a plant, and as such includes the surface of the stem, flowers and fruit, but most particularly the leaf surfaces. Compared with the rhizosphere and the endosphere the phyllosphere is nutrient poor and its environment more dynamic.
Interactions between plants and their associated microorganisms in many of these microbiomes can play pivotal roles in host plant health, function, and evolution.[8] Interactions between the host plant and phyllosphere bacteria have the potential to drive various aspects of host plant physiology.[9][3][10] However, as of 2020 knowledge of these bacterial associations in the phyllosphere remains relatively modest, and there is a need to advance fundamental knowledge of phyllosphere microbiome dynamics.[11][12]
The assembly of the phyllosphere microbiome, which can be strictly defined as epiphytic bacterial communities on the leaf surface, can be shaped by the microbial communities present in the surrounding environment (i.e., stochastic colonisation) and the host plant (i.e., biotic selection).[4][13][12] However, although the leaf surface is generally considered a discrete microbial habitat,[14][15] there is no consensus on the dominant driver of community assembly across phyllosphere microbiomes. For example, host-specific bacterial communities have been reported in the phyllosphere of co-occurring plant species, suggesting a dominant role of host selection.[15][16][17][12]
Conversely, microbiomes of the surrounding environment have also been reported to be the primary determinant of phyllosphere community composition.[14][18][19][20] As a result, the processes that drive phyllosphere community assembly are not well understood but unlikely to be universal across plant species. However, the existing evidence does indicate that phyllosphere microbiomes exhibiting host-specific associations are more likely to interact with the host than those primarily recruited from the surrounding environment.[9][21][22][23][12]
Overall, there remains high species richness in phyllosphere communities. Fungal communities are highly variable in the phyllosphere of temperate regions and are more diverse than in tropical regions.[24] There can be up to 107 microbes per square centimetre present on the leaf surfaces of plants, and the bacterial population of the phyllosphere on a global scale is estimated to be 1026 cells.[25] The population size of the fungal phyllosphere is likely to be smaller.[26]
Phyllosphere microbes from different plants appear to be somewhat similar at high levels of taxa, but at the lower levels taxa there remain significant differences. This indicates microorganisms may need finely tuned metabolic adjustment to survive in phyllosphere environment.[24] Proteobacteria seems to be the dominant colonizers, with Bacteroidetes and Actinobacteria also predominant in phyllospheres.[27] Although there are similarities between the rhizosphere and soil microbial communities, very little similarity has been found between phyllosphere communities and microorganisms floating in open air (aeroplankton).[28][6]
The search for a core microbiome in host-associated microbial communities is a useful first step in trying to understand the interactions that may be occurring between a host and its microbiome.[29][30] The prevailing core microbiome concept is built on the notion that the persistence of a taxon across the spatiotemporal boundaries of an ecological niche is directly reflective of its functional importance within the niche it occupies; it therefore provides a framework for identifying functionally critical microorganisms that consistently associate with a host species.[29][31][32][12]
Divergent definitions of “core microbiome” have arisen across scientific literature with researchers variably identifying “core taxa” as those persistent across distinct host microhabitats [33][34] and even different species.[17][21] Given the functional divergence of microorganisms across different host species [17] and microhabitats,[35] defining core taxa sensu stricto as those persistent across broad geographic distances within tissue- and species-specific host microbiomes, represents the most biologically and ecologically appropriate application of this conceptual framework.[36][12] Tissue- and species-specific core microbiomes across host populations separated by broad geographical distances have not been widely reported for the phyllosphere using the stringent definition established by Ruinen.[3][12]
Example: The mānuka phyllosphere
The flowering tea tree commonly known as mānuka is indigenous to New Zealand.[37] Mānuka honey, produced from the nectar of mānuka flowers, is known for its non-peroxide antibacterial properties.[38][39] These non-peroxide antibacterial properties have been principally linked to the accumulation of the three-carbon sugar dihydroxyacetone (DHA) in the nectar of the mānuka flower, which undergoes a chemical conversion to methylglyoxal (MGO) in mature honey.[40][41][42] However, the concentration of DHA in the nectar of mānuka flowers is notoriously variable, and the antimicrobial efficacy of mānuka honey consequently varies from region to region and from year to year.[43][44][45] Despite extensive research efforts, no reliable correlation has been identified between DHA production and climatic,[46] edaphic,[47] or host genetic factors.[48][12]

(B) The chart on the right shows how OTUs in phyllosphere and associated soil communities differed in relative abundances.[12]
Microorganisms have been studied in the mānuka rhizosphere and endosphere.[49][50][51] Earlier studies primarily focussed on fungi, and a 2016 study provided the first investigation of endophytic bacterial communities from three geographically and environmentally distinct mānuka populations using fingerprinting techniques and revealed tissue-specific core endomicrobiomes.[52][12] A 2020 study identified a habitat-specific and relatively abundant core microbiome in the mānuka phyllosphere, which was persistent across all samples. In contrast, non-core phyllosphere microorganisms exhibited significant variation across individual host trees and populations that was strongly driven by environmental and spatial factors. The results demonstrated the existence of a dominant and ubiquitous core microbiome in the phyllosphere of mānuka.[12]
See also
References
- ^ He, Sheng Yang (2020) When plants and their microbes are not in sync, the results can be disastrous The Conversation, 28 August 2020.
- ^ Last, F.T. (1955). "Seasonal incidence of Sporobolomyces on cereal leaves". Trans Br Mycol Soc. 38 (3): 221–239. doi:10.1016/s0007-1536(55)80069-1.
- ^ a b c Cid, Fernanda P.; Maruyama, Fumito; Murase, Kazunori; Graether, Steffen P.; Larama, Giovanni; Bravo, Leon A.; Jorquera, Milko A. (2018). "Draft genome sequences of bacteria isolated from the Deschampsia antarctica phyllosphere". Extremophiles. 22 (3): 537–552. doi:10.1007/s00792-018-1015-x. PMID 29492666. S2CID 4320165.
- ^ a b Leveau, Johan HJ (2019). "A brief from the leaf: Latest research to inform our understanding of the phyllosphere microbiome". Current Opinion in Microbiology. 49: 41–49. doi:10.1016/j.mib.2019.10.002. PMID 31707206.
- ^ Ruinen, J. (1956) "Occurrence of Beijerinckia species in the 'phyllosphere'". Nature, 177(4501): 220–221.
- ^ a b Dastogeer, K.M., Tumpa, F.H., Sultana, A., Akter, M.A. and Chakraborty, A. (2020) "Plant microbiome–an account of the factors that shape community composition and diversity". Current Plant Biology: 100161. doi:10.1016/j.cpb.2020.100161.
 Material was copied from this source, which is available under a Creative Commons Attribution 4.0 International License.
Material was copied from this source, which is available under a Creative Commons Attribution 4.0 International License.
- ^ Lindow, Steven E. (1996). "Role of Immigration and Other Processes in Determining Epiphytic Bacterial Populations". Aerial Plant Surface Microbiology. pp. 155–168. doi:10.1007/978-0-585-34164-4_10. ISBN 978-0-306-45382-3.
- ^ Friesen, Maren L.; Porter, Stephanie S.; Stark, Scott C.; von Wettberg, Eric J.; Sachs, Joel L.; Martinez-Romero, Esperanza (2011). "Microbially Mediated Plant Functional Traits". Annual Review of Ecology, Evolution, and Systematics. 42: 23–46. doi:10.1146/annurev-ecolsys-102710-145039.
- ^ a b Vogel, Christine; Bodenhausen, Natacha; Gruissem, Wilhelm; Vorholt, Julia A. (2016). "The Arabidopsis leaf transcriptome reveals distinct but also overlapping responses to colonization by phyllosphere commensals and pathogen infection with impact on plant health" (PDF). New Phytologist. 212: 192–207. doi:10.1111/nph.14036.
- ^ Kumaravel, Sowmya; Thankappan, Sugitha; Raghupathi, Sridar; Uthandi, Sivakumar (2018). "Draft Genome Sequence of Plant Growth-Promoting and Drought-Tolerant Bacillus altitudinis FD48, Isolated from Rice Phylloplane". Genome Announcements. 6 (9). doi:10.1128/genomeA.00019-18. PMC 5834328. PMID 29496824.
- ^ Laforest‐Lapointe, Isabelle; Whitaker, Briana K. (2019). "Decrypting the phyllosphere microbiota: Progress and challenges". American Journal of Botany. 106 (2): 171–173. doi:10.1002/ajb2.1229. PMID 30726571.
- ^ a b c d e f g h i j k l Noble, Anya S.; Noe, Stevie; Clearwater, Michael J.; Lee, Charles K. (2020). "A core phyllosphere microbiome exists across distant populations of a tree species indigenous to New Zealand". PLOS ONE. 15 (8): e0237079. Bibcode:2020PLoSO..1537079N. doi:10.1371/journal.pone.0237079. PMC 7425925. PMID 32790769.
{{cite journal}}: CS1 maint: unflagged free DOI (link) Material was copied from this source, which is available under a Creative Commons Attribution 4.0 International License.
Material was copied from this source, which is available under a Creative Commons Attribution 4.0 International License.  Material was copied from this source, which is available under a Creative Commons Attribution 4.0 International License.
Material was copied from this source, which is available under a Creative Commons Attribution 4.0 International License.
- ^ Vorholt, Julia A. (2012). "Microbial life in the phyllosphere". Nature Reviews Microbiology. 10 (12): 828–840. doi:10.1038/nrmicro2910. PMID 23154261. S2CID 10447146.
- ^ a b Stone, Bram W. G.; Jackson, Colin R. (2016). "Biogeographic Patterns Between Bacterial Phyllosphere Communities of the Southern Magnolia (Magnolia grandiflora) in a Small Forest". Microbial Ecology. 71 (4): 954–961. doi:10.1007/s00248-016-0738-4. PMID 26883131. S2CID 17292307.
- ^ a b Redford, Amanda J.; Bowers, Robert M.; Knight, Rob; Linhart, Yan; Fierer, Noah (2010). "The ecology of the phyllosphere: Geographic and phylogenetic variability in the distribution of bacteria on tree leaves". Environmental Microbiology. 12 (11): 2885–2893. doi:10.1111/j.1462-2920.2010.02258.x. PMC 3156554. PMID 20545741.
- ^ Vokou, Despoina; Vareli, Katerina; Zarali, Ekaterini; Karamanoli, Katerina; Constantinidou, Helen-Isis A.; Monokrousos, Nikolaos; Halley, John M.; Sainis, Ioannis (2012). "Exploring Biodiversity in the Bacterial Community of the Mediterranean Phyllosphere and its Relationship with Airborne Bacteria". Microbial Ecology. 64 (3): 714–724. doi:10.1007/s00248-012-0053-7. PMID 22544345. S2CID 17291303.
- ^ a b c Laforest-Lapointe, Isabelle; Messier, Christian; Kembel, Steven W. (2016). "Host species identity, site and time drive temperate tree phyllosphere bacterial community structure". Microbiome. 4 (1): 27. doi:10.1186/s40168-016-0174-1. PMC 4912770. PMID 27316353.
{{cite journal}}: CS1 maint: unflagged free DOI (link) - ^ Zarraonaindia, Iratxe; Owens, Sarah M.; Weisenhorn, Pamela; West, Kristin; Hampton-Marcell, Jarrad; Lax, Simon; Bokulich, Nicholas A.; Mills, David A.; Martin, Gilles; Taghavi, Safiyh; Van Der Lelie, Daniel; Gilbert, Jack A. (2015). "The Soil Microbiome Influences Grapevine-Associated Microbiota". mBio. 6 (2). doi:10.1128/mBio.02527-14. PMC 4453523. PMID 25805735.
- ^ Finkel, Omri M.; Burch, Adrien Y.; Lindow, Steven E.; Post, Anton F.; Belkin, Shimshon (2011). "Geographical Location Determines the Population Structure in Phyllosphere Microbial Communities of a Salt-Excreting Desert Tree". Applied and Environmental Microbiology. 77 (21): 7647–7655. doi:10.1128/AEM.05565-11. PMC 3209174. PMID 21926212.
- ^ Finkel, Omri M.; Burch, Adrien Y.; Elad, Tal; Huse, Susan M.; Lindow, Steven E.; Post, Anton F.; Belkin, Shimshon (2012). "Distance-Decay Relationships Partially Determine Diversity Patterns of Phyllosphere Bacteria on Tamrix Trees across the Sonoran Desert". Applied and Environmental Microbiology. 78 (17): 6187–6193. doi:10.1128/AEM.00888-12. PMC 3416633. PMID 22752165.
- ^ a b Kembel, S. W.; O'Connor, T. K.; Arnold, H. K.; Hubbell, S. P.; Wright, S. J.; Green, J. L. (2014). "Relationships between phyllosphere bacterial communities and plant functional traits in a neotropical forest". Proceedings of the National Academy of Sciences. 111 (38): 13715–13720. Bibcode:2014PNAS..11113715K. doi:10.1073/pnas.1216057111. S2CID 852584.
- ^ Innerebner, Gerd; Knief, Claudia; Vorholt, Julia A. (2011). "Protection of Arabidopsis thaliana against Leaf-Pathogenic Pseudomonas syringae by Sphingomonas Strains in a Controlled Model System". Applied and Environmental Microbiology. 77 (10): 3202–3210. doi:10.1128/AEM.00133-11. PMC 3126462. PMID 21421777.
- ^ Lajoie, Geneviève; Maglione, Rémi; Kembel, Steven W. (2020). "Adaptive matching between phyllosphere bacteria and their tree hosts in a neotropical forest". Microbiome. 8 (1): 70. doi:10.1186/s40168-020-00844-7. PMC 7243311. PMID 32438916.
{{cite journal}}: CS1 maint: unflagged free DOI (link) - ^ a b Finkel, Omri M.; Burch, Adrien Y.; Lindow, Steven E.; Post, Anton F.; Belkin, Shimshon (2011). "Geographical Location Determines the Population Structure in Phyllosphere Microbial Communities of a Salt-Excreting Desert Tree". Applied and Environmental Microbiology. 77 (21): 7647–7655. doi:10.1128/AEM.05565-11. PMC 3209174. PMID 21926212.
- ^ Vorholt, Julia A. (2012). "Microbial life in the phyllosphere". Nature Reviews Microbiology. 10 (12): 828–840. doi:10.1038/nrmicro2910. PMID 23154261. S2CID 10447146.
- ^ Lindow, Steven E.; Brandl, Maria T. (2003). "Microbiology of the Phyllosphere". Applied and Environmental Microbiology. 69 (4): 1875–1883. doi:10.1128/AEM.69.4.1875-1883.2003. PMC 154815. PMID 12676659. S2CID 2304379.
- ^ Bodenhausen, Natacha; Horton, Matthew W.; Bergelson, Joy (2013). "Bacterial Communities Associated with the Leaves and the Roots of Arabidopsis thaliana". PLOS ONE. 8 (2): e56329. doi:10.1371/journal.pone.0056329. PMC 3574144. PMID 23457551.
{{cite journal}}: CS1 maint: unflagged free DOI (link) - ^ Vokou, Despoina; Vareli, Katerina; Zarali, Ekaterini; Karamanoli, Katerina; Constantinidou, Helen-Isis A.; Monokrousos, Nikolaos; Halley, John M.; Sainis, Ioannis (2012). "Exploring Biodiversity in the Bacterial Community of the Mediterranean Phyllosphere and its Relationship with Airborne Bacteria". Microbial Ecology. 64 (3): 714–724. doi:10.1007/s00248-012-0053-7. PMID 22544345. S2CID 17291303.
- ^ a b Shade, Ashley; Handelsman, Jo (2012). "Beyond the Venn diagram: The hunt for a core microbiome". Environmental Microbiology. 14 (1): 4–12. doi:10.1111/j.1462-2920.2011.02585.x. PMID 22004523.
- ^ Berg, Gabriele; Rybakova, Daria; Fischer, Doreen; Cernava, Tomislav; Vergès, Marie-Christine Champomier; Charles, Trevor; Chen, Xiaoyulong; Cocolin, Luca; Eversole, Kellye; Corral, Gema Herrero; Kazou, Maria; Kinkel, Linda; Lange, Lene; Lima, Nelson; Loy, Alexander; MacKlin, James A.; Maguin, Emmanuelle; Mauchline, Tim; McClure, Ryan; Mitter, Birgit; Ryan, Matthew; Sarand, Inga; Smidt, Hauke; Schelkle, Bettina; Roume, Hugo; Kiran, G. Seghal; Selvin, Joseph; Souza, Rafael Soares Correa de; Van Overbeek, Leo; et al. (2020). "Microbiome definition re-visited: Old concepts and new challenges". Microbiome. 8 (1): 103. doi:10.1186/s40168-020-00875-0. PMC 7329523. PMID 32605663.
{{cite journal}}: CS1 maint: unflagged free DOI (link) - ^ Turnbaugh, Peter J.; Hamady, Micah; Yatsunenko, Tanya; Cantarel, Brandi L.; Duncan, Alexis; Ley, Ruth E.; Sogin, Mitchell L.; Jones, William J.; Roe, Bruce A.; Affourtit, Jason P.; Egholm, Michael; Henrissat, Bernard; Heath, Andrew C.; Knight, Rob; Gordon, Jeffrey I. (2009). "A core gut microbiome in obese and lean twins". Nature. 457 (7228): 480–484. Bibcode:2009Natur.457..480T. doi:10.1038/nature07540. PMC 2677729. PMID 19043404.
- ^ Lundberg, Derek S.; Lebeis, Sarah L.; Paredes, Sur Herrera; Yourstone, Scott; Gehring, Jase; Malfatti, Stephanie; Tremblay, Julien; Engelbrektson, Anna; Kunin, Victor; Rio, Tijana Glavina del; Edgar, Robert C.; Eickhorst, Thilo; Ley, Ruth E.; Hugenholtz, Philip; Tringe, Susannah Green; Dangl, Jeffery L. (2012). "Defining the core Arabidopsis thaliana root microbiome". Nature. 488 (7409): 86–90. Bibcode:2012Natur.488...86L. doi:10.1038/nature11237. PMC 4074413. PMID 22859206.
- ^ Hamonts, Kelly; Trivedi, Pankaj; Garg, Anshu; Janitz, Caroline; Grinyer, Jasmine; Holford, Paul; Botha, Frederik C.; Anderson, Ian C.; Singh, Brajesh K. (2018). "Field study reveals core plant microbiota and relative importance of their drivers". Environmental Microbiology. 20 (1): 124–140. doi:10.1111/1462-2920.14031. PMID 29266641. S2CID 10650949.
- ^ Cernava, Tomislav; Erlacher, Armin; Soh, Jung; Sensen, Christoph W.; Grube, Martin; Berg, Gabriele (2019). "Enterobacteriaceae dominate the core microbiome and contribute to the resistome of arugula (Eruca sativa Mill.)". Microbiome. 7 (1): 13. doi:10.1186/s40168-019-0624-7. PMC 6352427. PMID 30696492.
{{cite journal}}: CS1 maint: unflagged free DOI (link) - ^ Leff, Jonathan W.; Del Tredici, Peter; Friedman, William E.; Fierer, Noah (2015). "Spatial structuring of bacterial communities within individual Ginkgo bilobatrees". Environmental Microbiology. 17 (7): 2352–2361. doi:10.1111/1462-2920.12695. PMID 25367625.
- ^ Hernandez-Agreda, Alejandra; Gates, Ruth D.; Ainsworth, Tracy D. (2017). "Defining the Core Microbiome in Corals' Microbial Soup". Trends in Microbiology. 25 (2): 125–140. doi:10.1016/j.tim.2016.11.003. PMID 27919551.
- ^ Stephens, J. M. C.; Molan, P. C.; Clarkson, B. D. (2005). "A review of Leptospermum scoparium(Myrtaceae) in New Zealand". New Zealand Journal of Botany. 43 (2): 431–449. doi:10.1080/0028825X.2005.9512966. S2CID 53515334.
- ^ Cooper, R.A.; Molan, P.C.; Harding, K.G. (2002). "The sensitivity to honey of Gram-positive cocci of clinical significance isolated from wounds". Journal of Applied Microbiology. 93 (5): 857–863. doi:10.1046/j.1365-2672.2002.01761.x. PMID 12392533. S2CID 24517001.
- ^ Rabie, Erika; Serem, June Cheptoo; Oberholzer, Hester Magdalena; Gaspar, Anabella Regina Marques; Bester, Megan Jean (2016). "How methylglyoxal kills bacteria: An ultrastructural study". Ultrastructural Pathology. 40 (2): 107–111. doi:10.3109/01913123.2016.1154914. hdl:2263/52156. PMID 26986806. S2CID 13372064.
- ^ Adams, Christopher J.; Manley-Harris, Merilyn; Molan, Peter C. (2009). "The origin of methylglyoxal in New Zealand manuka (Leptospermum scoparium) honey". Carbohydrate Research. 344 (8): 1050–1053. doi:10.1016/j.carres.2009.03.020. PMID 19368902.
- ^ Atrott, Julia; Haberlau, Steffi; Henle, Thomas (2012). "Studies on the formation of methylglyoxal from dihydroxyacetone in Manuka (Leptospermum scoparium) honey". Carbohydrate Research. 361: 7–11. doi:10.1016/j.carres.2012.07.025. PMID 22960208.
- ^ Mavric, Elvira; Wittmann, Silvia; Barth, Gerold; Henle, Thomas (2008). "Identification and quantification of methylglyoxal as the dominant antibacterial constituent of Manuka (Leptospermum scoparium)honeys from New Zealand". Molecular Nutrition & Food Research. 52 (4): 483–489. doi:10.1002/mnfr.200700282.
- ^ Hamilton, G., Millner, J., Robertson, A. and Stephens, J. (2013) "Assessment of manuka provenances for production of high 'unique manuka factor' honey". Agronomy New Zealand, 43: 139–144.
- ^ Williams, Simon; King, Jessica; Revell, Maria; Manley-Harris, Merilyn; Balks, Megan; Janusch, Franziska; Kiefer, Michael; Clearwater, Michael; Brooks, Peter; Dawson, Murray (2014). "Regional, Annual, and Individual Variations in the Dihydroxyacetone Content of the Nectar of Ma̅nuka (Leptospermum scoparium) in New Zealand". Journal of Agricultural and Food Chemistry. 62 (42): 10332–10340. doi:10.1021/jf5045958. PMID 25277074.
- ^ Stephens, J.M.C. (2006) "The factors responsible for the varying levels of UMF® in mānuka (Leptospermum scoparium) honey", Doctoral dissertation, University of Waikato.
- ^ Noe, Stevie; Manley-Harris, Merilyn; Clearwater, Michael J. (2019). "Floral nectar of wild mānuka (Leptospermum scoparium) varies more among plants than among sites". New Zealand Journal of Crop and Horticultural Science. 47 (4): 282–296. doi:10.1080/01140671.2019.1670681. S2CID 204143940.
- ^ Nickless, Elizabeth M.; Anderson, Christopher W. N.; Hamilton, Georgie; Stephens, Jonathan M.; Wargent, Jason (2017). "Soil influences on plant growth, floral density and nectar yield in three cultivars of mānuka (Leptospermum scoparium)". New Zealand Journal of Botany. 55 (2): 100–117. doi:10.1080/0028825X.2016.1247732. S2CID 88657399.
- ^ Clearwater, Michael J.; Revell, Maria; Noe, Stevie; Manley-Harris, Merilyn (2018). "Influence of genotype, floral stage, and water stress on floral nectar yield and composition of mānuka (Leptospermum scoparium)". Annals of Botany. 121 (3): 501–512. doi:10.1093/aob/mcx183. PMC 5838834. PMID 29300875.
- ^ Johnston, Peter R. (1998). "Leaf endophytes of manuka (Leptospermum scoparium)". Mycological Research. 102 (8): 1009–1016. doi:10.1017/S0953756297005765.
- ^ McKenzie, E. H. C.; Johnston, P. R.; Buchanan, P. K. (2006). "Checklist of fungi on teatree (Kunzeaand Leptospermumspecies) in New Zealand". New Zealand Journal of Botany. 44 (3): 293–335. doi:10.1080/0028825X.2006.9513025. S2CID 84538904.
- ^ Wicaksono, Wisnu Adi; Sansom, Catherine E.; Eirian Jones, E.; Perry, Nigel B.; Monk, Jana; Ridgway, Hayley J. (2018). "Arbuscular mycorrhizal fungi associated with Leptospermum scoparium (Mānuka): Effects on plant growth and essential oil content". Symbiosis. 75: 39–50. doi:10.1007/s13199-017-0506-3. S2CID 4819178.
- ^ Wicaksono, Wisnu Adi; Jones, E. Eirian; Monk, Jana; Ridgway, Hayley J. (2016). "The Bacterial Signature of Leptospermum scoparium (Mānuka) Reveals Core and Accessory Communities with Bioactive Properties". PLOS ONE. 11 (9): e0163717. Bibcode:2016PLoSO..1163717W. doi:10.1371/journal.pone.0163717. PMC 5038978. PMID 27676607.
{{cite journal}}: CS1 maint: unflagged free DOI (link)
- ^ "Influence of Light on Plant–Phyllosphere Interaction". doi:10.3389/fpls.2018.01482.
{{cite journal}}: Cite journal requires|journal=(help)CS1 maint: unflagged free DOI (link) Material was copied from this source, which is available under a Creative Commons Attribution 4.0 International License.
Material was copied from this source, which is available under a Creative Commons Attribution 4.0 International License.


