OPTICS algorithm
| Part of a series on |
| Machine learning and data mining |
|---|
Ordering points to identify the clustering structure (OPTICS) is an algorithm for finding density-based[1] clusters in spatial data. It was presented by Mihael Ankerst, Markus M. Breunig, Hans-Peter Kriegel and Jörg Sander.[2] Its basic idea is similar to DBSCAN,[3] but it addresses one of DBSCAN's major weaknesses: the problem of detecting meaningful clusters in data of varying density. To do so, the points of the database are (linearly) ordered such that spatially closest points become neighbors in the ordering. Additionally, a special distance is stored for each point that represents the density that must be accepted for a cluster so that both points belong to the same cluster. This is represented as a dendrogram.
Basic idea[edit]
Like DBSCAN, OPTICS requires two parameters: ε, which describes the maximum distance (radius) to consider, and MinPts, describing the number of points required to form a cluster. A point p is a core point if at least MinPts points are found within its ε-neighborhood (including point p itself). In contrast to DBSCAN, OPTICS also considers points that are part of a more densely packed cluster, so each point is assigned a core distance that describes the distance to the MinPtsth closest point:
The reachability-distance of another point o from a point p is either the distance between o and p, or the core distance of p, whichever is bigger:
If p and o are nearest neighbors, this is the we need to assume to have p and o belong to the same cluster.
Both core-distance and reachability-distance are undefined if no sufficiently dense cluster (w.r.t. ε) is available. Given a sufficiently large ε, this never happens, but then every ε-neighborhood query returns the entire database, resulting in runtime. Hence, the ε parameter is required to cut off the density of clusters that are no longer interesting, and to speed up the algorithm.
The parameter ε is, strictly speaking, not necessary. It can simply be set to the maximum possible value. When a spatial index is available, however, it does play a practical role with regards to complexity. OPTICS abstracts from DBSCAN by removing this parameter, at least to the extent of only having to give the maximum value.
Pseudocode[edit]
The basic approach of OPTICS is similar to DBSCAN, but instead of maintaining known, but so far unprocessed cluster members in a set, they are maintained in a priority queue (e.g. using an indexed heap).
function OPTICS(DB, ε, MinPts) is
for each point p of DB do
p.reachability-distance = UNDEFINED
for each unprocessed point p of DB do
N = getNeighbors(p, ε)
mark p as processed
output p to the ordered list
if core-distance(p, ε, MinPts) != UNDEFINED then
Seeds = empty priority queue
update(N, p, Seeds, ε, MinPts)
for each next q in Seeds do
N' = getNeighbors(q, ε)
mark q as processed
output q to the ordered list
if core-distance(q, ε, MinPts) != UNDEFINED do
update(N', q, Seeds, ε, MinPts)
In update(), the priority queue Seeds is updated with the -neighborhood of and , respectively:
function update(N, p, Seeds, ε, MinPts) is
coredist = core-distance(p, ε, MinPts)
for each o in N
if o is not processed then
new-reach-dist = max(coredist, dist(p,o))
if o.reachability-distance == UNDEFINED then // o is not in Seeds
o.reachability-distance = new-reach-dist
Seeds.insert(o, new-reach-dist)
else // o in Seeds, check for improvement
if new-reach-dist < o.reachability-distance then
o.reachability-distance = new-reach-dist
Seeds.move-up(o, new-reach-dist)
OPTICS hence outputs the points in a particular ordering, annotated with their smallest reachability distance (in the original algorithm, the core distance is also exported, but this is not required for further processing).
Extracting the clusters[edit]
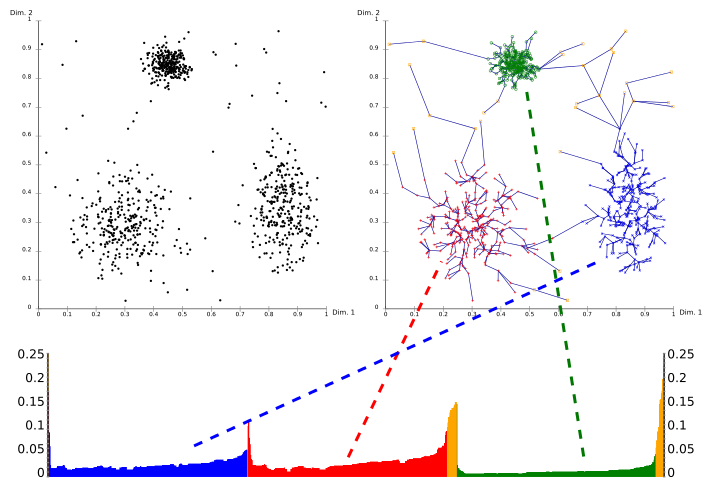
Using a reachability-plot (a special kind of dendrogram), the hierarchical structure of the clusters can be obtained easily. It is a 2D plot, with the ordering of the points as processed by OPTICS on the x-axis and the reachability distance on the y-axis. Since points belonging to a cluster have a low reachability distance to their nearest neighbor, the clusters show up as valleys in the reachability plot. The deeper the valley, the denser the cluster.
The image above illustrates this concept. In its upper left area, a synthetic example data set is shown. The upper right part visualizes the spanning tree produced by OPTICS, and the lower part shows the reachability plot as computed by OPTICS. Colors in this plot are labels, and not computed by the algorithm; but it is well visible how the valleys in the plot correspond to the clusters in above data set. The yellow points in this image are considered noise, and no valley is found in their reachability plot. They are usually not assigned to clusters, except the omnipresent "all data" cluster in a hierarchical result.
Extracting clusters from this plot can be done manually by selecting ranges on the x-axis after visual inspection, by selecting a threshold on the y-axis (the result is then similar to a DBSCAN clustering result with the same and minPts parameters; here a value of 0.1 may yield good results), or by different algorithms that try to detect the valleys by steepness, knee detection, or local maxima. A range of the plot beginning with a steep descent and ending with a steep ascent is considered a valley, and corresponds to a contiguous area of high density. Additional care must be taken to the last points in a valley to assign them to the inner or outer cluster, this can be achieved by considering the predecessor.[4] Clusterings obtained this way usually are hierarchical, and cannot be achieved by a single DBSCAN run.
Complexity[edit]
Like DBSCAN, OPTICS processes each point once, and performs one -neighborhood query during this processing. Given a spatial index that grants a neighborhood query in runtime, an overall runtime of is obtained. The worst case however is , as with DBSCAN. The authors of the original OPTICS paper report an actual constant slowdown factor of 1.6 compared to DBSCAN. Note that the value of might heavily influence the cost of the algorithm, since a value too large might raise the cost of a neighborhood query to linear complexity.
In particular, choosing (larger than the maximum distance in the data set) is possible, but leads to quadratic complexity, since every neighborhood query returns the full data set. Even when no spatial index is available, this comes at additional cost in managing the heap. Therefore, should be chosen appropriately for the data set.
Extensions[edit]
OPTICS-OF[5] is an outlier detection algorithm based on OPTICS. The main use is the extraction of outliers from an existing run of OPTICS at low cost compared to using a different outlier detection method. The better known version LOF is based on the same concepts.
DeLi-Clu,[6] Density-Link-Clustering combines ideas from single-linkage clustering and OPTICS, eliminating the parameter and offering performance improvements over OPTICS.
HiSC[7] is a hierarchical subspace clustering (axis-parallel) method based on OPTICS.
HiCO[8] is a hierarchical correlation clustering algorithm based on OPTICS.
DiSH[9] is an improvement over HiSC that can find more complex hierarchies.
FOPTICS[10] is a faster implementation using random projections.
HDBSCAN*[11] is based on a refinement of DBSCAN, excluding border-points from the clusters and thus following more strictly the basic definition of density-levels by Hartigan.[12]
Availability[edit]
Java implementations of OPTICS, OPTICS-OF, DeLi-Clu, HiSC, HiCO and DiSH are available in the ELKI data mining framework (with index acceleration for several distance functions, and with automatic cluster extraction using the ξ extraction method). Other Java implementations include the Weka extension (no support for ξ cluster extraction).
The R package "dbscan" includes a C++ implementation of OPTICS (with both traditional dbscan-like and ξ cluster extraction) using a k-d tree for index acceleration for Euclidean distance only.
Python implementations of OPTICS are available in the PyClustering library and in scikit-learn. HDBSCAN* is available in the hdbscan library.
References[edit]
- ^ Kriegel, Hans-Peter; Kröger, Peer; Sander, Jörg; Zimek, Arthur (May 2011). "Density-based clustering". Wiley Interdisciplinary Reviews: Data Mining and Knowledge Discovery. 1 (3): 231–240. doi:10.1002/widm.30. S2CID 36920706.
- ^ Mihael Ankerst; Markus M. Breunig; Hans-Peter Kriegel; Jörg Sander (1999). OPTICS: Ordering Points To Identify the Clustering Structure. ACM SIGMOD international conference on Management of data. ACM Press. pp. 49–60. CiteSeerX 10.1.1.129.6542.
- ^ Martin Ester; Hans-Peter Kriegel; Jörg Sander; Xiaowei Xu (1996). Evangelos Simoudis; Jiawei Han; Usama M. Fayyad (eds.). A density-based algorithm for discovering clusters in large spatial databases with noise. Proceedings of the Second International Conference on Knowledge Discovery and Data Mining (KDD-96). AAAI Press. pp. 226–231. CiteSeerX 10.1.1.71.1980. ISBN 1-57735-004-9.
- ^ Schubert, Erich; Gertz, Michael (2018-08-22). Improving the Cluster Structure Extracted from OPTICS Plots (PDF). Lernen, Wissen, Daten, Analysen (LWDA 2018). Vol. CEUR-WS 2191. pp. 318–329 – via CEUR-WS.
- ^ Markus M. Breunig; Hans-Peter Kriegel; Raymond T. Ng; Jörg Sander (1999). "OPTICS-OF: Identifying Local Outliers". Principles of Data Mining and Knowledge Discovery. Lecture Notes in Computer Science. Vol. 1704. Springer-Verlag. pp. 262–270. doi:10.1007/b72280. ISBN 978-3-540-66490-1. S2CID 27352458.
- ^ Achtert, Elke; Böhm, Christian; Kröger, Peer (2006). "DeLi-Clu: Boosting Robustness, Completeness, Usability, and Efficiency of Hierarchical Clustering by a Closest Pair Ranking". In Ng, Wee Keong; Kitsuregawa, Masaru; Li, Jianzhong; Chang, Kuiyu (eds.). Advances in Knowledge Discovery and Data Mining, 10th Pacific-Asia Conference, PAKDD 2006, Singapore, April 9-12, 2006, Proceedings. Lecture Notes in Computer Science. Vol. 3918. Springer. pp. 119–128. doi:10.1007/11731139_16.
- ^ Achtert, Elke; Böhm, Christian; Kriegel, Hans-Peter; Kröger, Peer; Müller-Gorman, Ina; Zimek, Arthur (2006). "Finding Hierarchies of Subspace Clusters". In Fürnkranz, Johannes; Scheffer, Tobias; Spiliopoulou, Myra (eds.). Knowledge Discovery in Databases: PKDD 2006, 10th European Conference on Principles and Practice of Knowledge Discovery in Databases, Berlin, Germany, September 18-22, 2006, Proceedings. Lecture Notes in Computer Science. Vol. 4213. Springer. pp. 446–453. doi:10.1007/11871637_42.
- ^ Achtert, E.; Böhm, C.; Kröger, P.; Zimek, A. (2006). "Mining Hierarchies of Correlation Clusters". 18th International Conference on Scientific and Statistical Database Management (SSDBM'06). pp. 119–128. CiteSeerX 10.1.1.707.7872. doi:10.1109/SSDBM.2006.35. ISBN 978-0-7695-2590-7. S2CID 2679909.
- ^ Achtert, Elke; Böhm, Christian; Kriegel, Hans-Peter; Kröger, Peer; Müller-Gorman, Ina; Zimek, Arthur (2007). "Detection and Visualization of Subspace Cluster Hierarchies". In Ramamohanarao, Kotagiri; Krishna, P. Radha; Mohania, Mukesh K.; Nantajeewarawat, Ekawit (eds.). Advances in Databases: Concepts, Systems and Applications, 12th International Conference on Database Systems for Advanced Applications, DASFAA 2007, Bangkok, Thailand, April 9-12, 2007, Proceedings. Lecture Notes in Computer Science. Vol. 4443. Springer. pp. 152–163. doi:10.1007/978-3-540-71703-4_15.
- ^ Schneider, Johannes; Vlachos, Michail (2013). "Fast parameterless density-based clustering via random projections". 22nd ACM International Conference on Information and Knowledge Management(CIKM).
- ^ Campello, Ricardo J. G. B.; Moulavi, Davoud; Zimek, Arthur; Sander, Jörg (22 July 2015). "Hierarchical Density Estimates for Data Clustering, Visualization, and Outlier Detection". ACM Transactions on Knowledge Discovery from Data. 10 (1): 1–51. doi:10.1145/2733381. S2CID 2887636.
- ^ J.A. Hartigan (1975). Clustering algorithms. John Wiley & Sons.