Ultrasensitivity
This article may contain an excessive amount of intricate detail that may interest only a particular audience. (June 2019) |
In molecular biology, ultrasensitivity describes an output response that is more sensitive to stimulus change than the hyperbolic Michaelis-Menten response. Ultrasensitivity is one of the biochemical switches in the cell cycle and has been implicated in a number of important cellular events, including exiting G2 cell cycle arrests in Xenopus laevis oocytes, a stage to which the cell or organism would not want to return.[1]
Ultrasensitivity is a cellular system which triggers entry into a different cellular state.[2] Ultrasensitivity gives a small response to first input signal, but an increase in the input signal produces higher and higher levels of output. This acts to filter out noise, as small stimuli and threshold concentrations of the stimulus (input signal) is necessary for the trigger which allows the system to get activated quickly.[3] Ultrasensitive responses are represented by sigmoidal graphs, which resemble cooperativity. The quantification of ultrasensitivity is often performed approximately by the Hill equation:
Where Hill's coefficient (n) may represent quantitative measure of ultrasensitive response.[4]

Historical development[edit]
Zero-order ultrasensitivity was first described by Albert Goldbeter and Daniel Koshland, Jr in 1981 in a paper in the Proceedings of the National Academy of Sciences.[5] They showed using mathematical modeling that modification of enzymes operating outside of first order kinetics required only small changes in the concentration of the effector to produce larger changes in the amount of modified protein. This amplification provided added sensitivity in biological control, and implicated the importance of this in many biological systems.
Many biological processes are binary (ON-OFF), such as cell fate decisions,[6] metabolic states, and signaling pathways. Ultrasensitivity is a switch that helps decision-making in such biological processes.[7] For example, in apoptotic process, a model showed that a positive feedback of inhibition of caspase 3 (Casp3) and Casp9 by inhibitors of apoptosis can bring about ultrasensitivity (bistability). This positive feedback cooperates with Casp3-mediated feedback cleavage of Casp9 to generate irreversibility in caspase activation (switch ON), which leads to cell apoptosis.[8] Another model also showed similar but different positive feedback controls in Bcl-2 family proteins in apoptotic process.[9]
Recently, Jeyeraman et al. have proposed that the phenomenon of ultrasensitivity may be further subdivided into three sub-regimes, separated by sharp stimulus threshold values: OFF, OFF-ON-OFF, and ON. Based on their model, they proposed that this sub-regime of ultrasensitivity, OFF-ON-OFF, is like a switch-like adaption which can be accomplished by coupling N phosphorylation–dephosphorylation cycles unidirectionally, without any explicit feedback loops.[10]
Other recent work has emphasized that not only is the topology of networks important for creating ultrasensitivity responses, but that their composition (enzymes vs. transcription factors) strongly affects whether they will exhibit robust ultrasensitivity. Mathematical modeling suggests for a broad array of network topologies that a combination of enzymes and transcription factors tends to provide more robust ultrasensitivity than that seen in networks composed entirely of transcription factors or composed entirely of enzymes.[11]
Mechanisms[edit]
Ultrasensitivity can be achieved through several mechanisms:
- Multistep mechanisms (examples: cooperativity)[12] and multisite phosphorylation[13]
- Buffering mechanisms (examples: decoy phosphorylation sites)[14] or stoichiometric inhibitors[15]
- Changes in localisation (such as translocation across the nuclear envelope)
- Saturation mechanisms (also known as zero-order ultrasensitivity)[16]
- Positive feedback[17]
- Allovalency
- Non-Zero-Order Ultrasensitivity in Membrane Proteins
- Dissipative Allostery
Multistep Mechanisms[edit]
Multipstep ultrasensitivity occurs when a single effector acts on several steps in a cascade.[18] Successive cascade signals can result in higher levels of noise being introduced into the signal that can interfere with the final output. This is especially relevant for large cascades, such as the flagellar regulatory system in which the master regulator signal is transmitted through multiple intermediate regulators before activating transcription.[19] Cascade ultrasensitivity can reduce noise and therefore require less input for activation.[12] Additionally, multiple phosphorylation events are an example of ultrasensitivity. Recent modeling has shown that multiple phosphorylation sites on membrane proteins could serve to locally saturate enzyme activity. Proteins at the membrane are greatly reduced in mobility compared to those in the cytoplasm, this means that a membrane tethered enzyme acting upon a membrane protein will take longer to diffuse away. With the addition of multiple phosphorylation sites upon the membrane substrate, the enzyme can - by a combination of increased local concentration of enzyme and increased substrates - quickly reach saturation.[20]
Buffering Mechanisms[edit]
Buffering Mechanisms such as molecular titration can generate ultrasensitivity. In vitro, this can be observed for the simple mechanism:
Where the monomeric form of A is active and it can be inactivated by binding B to form the heterodimer AB. When the concentration of ( = [B] + [AB]) is much greater than the , this system exhibits a threshold determined by the concentration of .[21] At concentrations of ( = [A] +[AB]), lower than , B acts as a buffer to free A and nearly all A will be found as AB. However, at the equivalence point, when ≈ , can no longer buffer the increase in , so a small increase in causes a large increase in A.[22] The strength of the ultrasensitivity of [A] to changes in is determined by /.[22] Ultrasensitivity occurs when this ratio is greater than one and is increased as the ratio increases. Above the equivalence point, and A are again linearly related. In vivo, the synthesis of A and B as well as the degradation of all three components complicates generation of ultrasensitivity. If the synthesis rates of A and B are equal this system still exhibits ultrasensitivity at the equivalence point.[22]
One example of a buffering mechanism is protein sequestration, which is a common mechanism found in signalling and regulatory networks.[23] In 2009, Buchler and Cross constructed a synthetic genetic network that was regulated by protein sequestration of a transcriptional activator by a dominant-negative inhibitor. They showed that this system results in a flexibile ultrasensitive response in gene expression. It is flexible in that the degree of ultrasensitivity can be altered by changing expression levels of the dominant-negative inhibitor. Figure 1 in their article illustrates how an active transcription factor can be sequestered by an inhibitor into the inactive complex AB that is unable to bind DNA. This type of mechanism results in an "all-or-none" response, or ultransensitivy, when the concentration of the regulatory protein increases to the point of depleting the inhibitor. Robust buffering against a response exists below this concentration threshold, and when it is reached any small increase in input is amplified into a large change in output.[citation needed]
Changes in localization[edit]
Translocation[edit]
Signal transduction is regulated in various ways and one of the ways is translocation. Regulated translocation generates ultrasensitive response in mainly three ways:
- Regulated translocation increases the local concentration of the signaling protein. When concentration of the signaling protein is high enough to partially saturate the enzyme that inactivates it, ultrasensitive response is generated.
- Translocation of multiple components of the signaling cascade, where stimulus (input signal) causes translocation of both signaling protein and its activator in the same subcellular compartment and thereby generates ultrasensitive response which increases speed and accuracy of the signal.
- Translocation to the compartment which contains stoichiometric inhibitors.[4]
Translocation is one way of regulating signal transduction, and it can generate ultrasensitive switch-like responses or multistep-feedback loop mechanisms. A switch-like response will occur if translocation raises the local concentration of a signaling protein. For example, epidermal growth factor (EGF) receptors can be internalized through clathrin-independent endocytosis (CIE) and/or clathrin-dependent endocytosis (CDE) in ligand concentration-dependent manner. The distribution of receptors into the two pathways was shown to be EGF concentration-dependent. In the presence of low concentrations of EGF, the receptor was exclusively internalized via CDE, whereas at high concentrations, receptors were equally distributed between CDE and CIE.[4][24]
Saturation mechanisms (Zero-order ultrasensitivity)[edit]
Zero-order ultrasensitivity takes place under saturating conditions.[25] For example, consider an enzymatic step with a kinase, phosphatase, and substrate. Steady state levels of the phosphorylated substrate have an ultrasensitive response when there is enough substrate to saturate all available kinases and phosphatases.[25][26] Under these conditions, small changes in the ratio of kinase to phosphatase activity can dramatically change the number of phosphorylated substrate (For a graph illustrating this behavior, see [5]). This enhancement in sensitivity of steady state phosphorylated substrate to Km, or the ratio of kinase to phosphatase activity, is termed zero-order to distinguish it from the first order behavior described by Michaelis-Menten dynamics, wherein the steady state concentration responds in a more gradual fashion than the switch-like behavior exhibited in ultrasensitivity.[18]
Using the notation from Goldbeter & Koshland,[5] let W be a certain substrate protein and let W' be a covalently modified version of W. The conversion of W to W' is catalyzed by some enzyme and the reverse conversion of W' to W is catalyzed by a second enzyme according to following equations:
The concentrations of all necessary components (such as ATP) are assumed to be constant and represented in the kinetic constants. Using the chemical equations above, the reaction rate equations for each component are:
The total concentration of each component is given by:
The zero order mechanism assumes that the or . In other words, the system is in a Michaelis-Menten steady state, which means, to a good approximation, and are constant. From these kinetic expressions one can solve for at steady state defining and
where and
When the is plotted against the molar ratio and it can be seen that the W to W' conversion occurs over a much smaller change in the ratio than it would under first order (non-saturating) conditions, which is the telltale sign of ultrasensitivity.
Positive Feedback[edit]
Positive feedback loops can cause ultrasensitive responses. An example of this is seen in the transcription of certain eukaryotic genes in which non-cooperative transcription factor binding changes positive feedback loops of histone modification that results in an ultrasensitive activation of transcription. The binding of a transcription factor recruits histone acetyltransferases and methyltransferases. The acetylation and methylation of histones recruits more acetyltransferases and methyltransferases that results in a positive feedback loop. Ultimately, this results in activation of transcription.[17]
Additionally, positive feedback can induce bistability in Cyclin B1- by the two regulators Wee1 and Cdc25C, leading to the cell's decision to commit to mitosis. The system cannot be stable at intermediate levels of Cyclin B1, and the transition between the two stable states is abrupt when increasing levels of Cyclin B1 switches the system from low to high activity. Exhibiting hysteresis, for different levels of Cyclin B1, the switches from low to high and high to low states vary.[27] However, the emergence of a bistable system is highly influenced by the sensitivity of its feedback loops. It has been shown in Xenopus egg extracts that Cdc25C hyperphosphorylation is a highly ultrasensitive function of Cdk activity, displaying a high value of the Hill coefficient (approx. 11), and the dephosphorylation step of Ser 287 in Cdc25C (also involved in Cdc25C activation) is even more ultrasensitive, displaying a Hill coefficient of approximately 32.[28]
Allovalency[edit]
A proposed mechanism of ultrasensitivity, called allovalency, suggests that activity "derives from a high local concentration of interaction sites moving independently of each other"[29] Allovalency was first proposed when it was believed to occur in the pathway in which Sic1, is degraded in order for Cdk1-Clb (B-type cyclins) to allow entry into mitosis. Sic1 must be phosphorylated multiple times in order to be recognized and degraded by Cdc4 of the SCF Complex.[30] Since Cdc4 only has one recognition site for these phosphorylated residues it was suggested that as the amount of phosphorylation increases, it exponentially increases the likelihood that Sic1 is recognized and degraded by Cdc4. This type of interaction was thought to be relatively immune to loss of any one site and easily tuned to any given threshold by adjusting the properties of individual sites. Assumptions for the allovalency mechanism were based on a general mathematical model that describes the interaction between a polyvalent disordered ligand and a single receptor site[29] It was later found that the ultrasensitivity in Cdk1 levels by degradation of Sic1 is in fact due to a positive feedback loop.[31]
Non-Zero-Order Ultrasensitivity in Membrane Proteins[edit]
Modeling by Dushek et al.[32] proposes a possible mechanism for ultrasensitivity outside of the zero-order regime. For the case of membrane-bound enzymes acting on membrane-bound substrates with multiple enzymatic sites (such as tyrosine-phosphorylated receptors like the T-Cell receptor), ultrasensitive responses could be seen, crucially dependent on three factors: 1) limited diffusion in the membrane, 2) multiple binding sites on the substrate, and 3) brief enzymatic inactivation following catalysis.
Under these particular conditions, although the enzyme may be in excess of the substrate (first-order regime), the enzyme is effectively locally saturated with substrate due to the multiple binding sites, leading to switch-like responses. This mechanism of ultrasensitivity is independent of enzyme concentration, however the signal is significantly enhanced depending on the number of binding sites on the substrate.[32] Both conditional factors (limited diffusion and inactivation) are physiologically plausible, but have yet to be experimentally confirmed. Dushek's modeling found increasing Hill cooperativity numbers with more substrate sites (phosphorylation sites), and with greater steric/diffusional hindrance between enzyme and substrate. This mechanism of ultrasensitivity based on local enzyme saturation arises partly from passive properties of slow membrane diffusion, and therefore may be generally applicable.
Dissipative Allostery[edit]
The bacterial flagellar motor has been proposed to follow a dissipative allosteric model, where ultrasensitivity comes as a combination of protein binding affinity and energy contributions from the proton motive force (see Flagellar motors and chemotaxis below).
Impact of upstream and downstream components on module's ultrasensitivity[edit]
In a living cell, ultrasensitive modules are embedded in a bigger network with upstream and downstream components. This components may constrain the range of inputs that the module will receive as well as the range of the module's outputs that network will be able to detect. Altszyler et al. (2014)[33] studied how the effective ultrasensitivity of a modular system is affected by these restrictions. They found for some ultrasensitive motifs that dynamic range limitations imposed by downstream components can produce effective sensitivities much larger than that of the original module when considered in isolation.
Hill Coefficient[edit]
Ultrasensitive behavior is typically represented by a sigmoidal curve, as small alterations in the stimulus can trigger large changes in the response . One such relation is the Hill equation:
where is the Hill coefficient which quantifies the steepness of the sigmoidal stimulus-response curve and it is therefore a sensitivity parameter. It is often used to assess the cooperativity of a system. A Hill coefficient greater than one is indicative of positive cooperativity and thus, the system exhibits ultrasensitivity.[34] Systems with a Hill coefficient of 1 are noncooperative and follow the classical Michaelis-Menten kinetics. Enzymes exhibiting noncooperative activity are represented by hyperbolic stimulus/response curves, compared to sigmoidal curves for cooperative (ultrasensitive) enzymes.[35] In mitogen-activated protein kinase (MAPK) signaling (see example below), the ultrasensitivity of the signaling is supported by the sigmoidal stimulus/response curve that is comparable to an enzyme with a Hill coefficient of 4.0-5.0. This is even more ultrasensitive to the cooperative binding activity of hemoglobin, which has a Hill coefficient of 2.8.[35]
Calculation[edit]
From an operational point of view the Hill coefficient can be calculated as:
- .
where and are the input values needed to produce the 10% and 90% of the maximal response, respectively.
Response Coefficient[edit]
Global sensitivity measures such as the Hill coefficient do not characterise the local behaviours of the s-shaped curves. Instead, these features are well captured by the response coefficient measure [36] defined as:
In systems biology, such system responses are referred to as control coefficients. Specifically, the concentration control coefficients measure the response of concentrations to changes in a given input. In addition, within the framework of the more general biochemical control analysis, such responses can be described in terms of the individual local responses, called the elasticities.
Link between Hill Coefficient and Response coefficient[edit]
Altszyler et al. (2017) have shown that these ultrasensitivity measures can be linked by the following equation:[37]
where denoted the mean value of the variable x over the range [a,b].
Ultrasensitivity in function composition[edit]
Consider two coupled ultrasensitive modules, disregarding effects of sequestration of molecular components between layers. In this case, the expression for the system's dose-response curve, , results from the mathematical composition of the functions, , which describe the input/output relationship of isolated modules :
Brown et al. (1997) [38] have shown that the local ultrasensitivity of the different layers combines multiplicatively:
- .
In connection with this result, Ferrell et al. (1997) [39] showed, for Hill-type modules, that the overall cascade global ultrasensitivity had to be less than or equal to the product of the global ultrasensitivity estimations of each cascade's layer,
- ,
where and are the Hill coefficient of modules 1 and 2 respectively.
Altszyler et al. (2017) [37] have shown that the cascade's global ultrasensitivity can be analytically calculated:
where and delimited the Hill input's working range of the composite system, i.e. the input values for the i-layer so that the last layer (corresponding to in this case) reached the 10% and 90% of it maximal output level. It followed this equation that the system's Hill coefficient could be written as the product of two factors, and , which characterized local average sensitivities over the relevant input region for each layer: , with in this case.
For the more general case of a cascade of modules, the Hill Coefficient can be expressed as:
- ,
Supramultiplicativity[edit]
Several authors have reported the existence of supramultiplicative behavior in signaling cascades [40][33](i.e. the ultrasensitivity of the combination of layers is higher than the product of individual ultrasensitivities), but in many cases the ultimate origin of supramultiplicativity remained elusive. Altszyler et al. (2017)[37] framework naturally suggested a general scenario where supramultiplicative behavior could take place. This could occur when, for a given module, the corresponding Hill's input working range was located in an input region with local ultrasensitivities higher than the global ultrasensitivity of the respective dose-response curve.
Role in Cellular Processes[edit]
MAP Kinase Signaling Cascade[edit]
A ubiquitous signaling motif that exhibits ultrasensitivity is the MAPK (mitogen-activated protein kinase) cascade, which can take a graded input signal and produce a switch-like output, such as gene transcription or cell cycle progression. In this common motif, MAPK is activated by an earlier kinase in the cascade, called MAPK kinase, or MAPKK. Similarly, MAPKK is activated by MAPKK kinase, or MAPKKK. These kinases are sequentially phosphorylated when MAPKKK is activated, usually via a signal received by a membrane-bound receptor protein. MAPKKK activates MAPKK, and MAPKK activates MAPK.[35] Ultrasensitivity arises in this system due to several features:
- MAPK and MAPKK both require two separate phosphorylation events to be activated.
- The reversal of MAPK phosphorylation by specific phosphatases requires an increasing concentration of activation signals from each prior kinase to achieve an output of the same magnitude.
- The MAPKK is at a concentration above the KΜ for its specific phosphatase and MAPK is at a concentration above the KΜ for MAPKK.
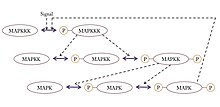
Besides the MAPK cascade, ultrasensitivity has also been reported in muscle glycolysis, in the phosphorylation of isocitrate dehydrogenase and in the activation of the calmodulin-dependent protein kinase II (CAMKII).[34]
An ultrasensitive switch has been engineered by combining a simple linear signaling protein (N-WASP) with one to five SH3 interaction modules that have autoinhibitory and cooperative properties. Addition of a single SH3 module created a switch that was activated in a linear fashion by exogenous SH3-binding peptide. Increasing number of domains increased ultrasensitivity. A construct with three SH3 modules was activated with an apparent Hill coefficient of 2.7 and a construct with five SH3 module was activated with an apparent Hill coefficient of 3.9.[41]
Translocation[edit]
During G2 phase of the cell cycle, Cdk1 and cyclin B1 makes a complex and forms maturation promoting factor (MPF). The complex accumulates in the nucleus due to phosphorylation of the cyclin B1 at multiple sites, which inhibits nuclear export of the complex. Phosphorylation of Thr19 and Tyr15 residues of Cdk1 by Wee1 and MYT1 keeps the complex inactive and inhibits entry into mitosis whereas dephosphorylation of Cdk1 by CDC25C phosphatase at Thr19 and Tyr15 residues, activates the complex which is necessary in order to enter mitosis. Cdc25C phosphatase is present in the cytoplasm and in late G2 phase it is translocated into the nucleus by signaling such as PIK1,[42] PIK3.[43] The regulated translocation and accumulation of the multiple required signaling cascade components, MPF and its activator Cdc25, in the nucleus generates efficient activation of the MPF and produces switch-like, ultrasensitive entry into mitosis.[4]
The figure[4] shows different possible mechanisms for how increased regulation of the localization of signaling components by the stimulus (input signal) shifts the output from Michaelian response to ultrasensitive response. When stimulus is regulating only inhibition of Cdk1-cyclinB1 nuclear export, the outcome is Michaelian response, Fig (a). But if the stimulus can regulate localization of multiple components of the signaling cascade, i.e. inhibition of Cdk1-cyclinB1 nuclear export and translocation of the Cdc25C to nucleus, then the outcome is ultrasensitive response, Fig (b). As more components of the signaling cascade are regulated and localized by the stimulus—i.e. inhibition of Cdk1-cyclinB1 nuclear export, translocation of the Cdc25C to the nucleus, and activation of Cdc25C—the output response becomes more and more ultrasensitive, Fig(c).[4]
Buffering (decoy)[edit]
During mitosis, mitotic spindle orientation is essential for determining the site of cleavage furrowing and position of daughter cells for subsequent cell fate determination.[44] This orientation is achieved by polarizing cortical factors and rapid alignment of the spindle with the polarity axis. In fruit flies, three cortical factors have been found to regulate the position of the spindle: heterotrimeric G protein α subunit (Gαi),[45] Partner of Inscuteable (Pins),[46] and Mushroom body defect (Mud).[47] Gαi localizes at apical cortex to recruit Pins. Upon binding to GDP-bound Gαi, Pins is activated and recruits Mud to achieve polarized distribution of cortical factors.[48] N-terminal tetratricopeptide repeats (TPRs) in Pins is the binding region for Mud, but is autoinhibited by intrinsic C-terminal GoLoco domains (GLs) in the absence of Gαi.[49][50] Activation of Pins by Gαi binding to GLs is highly ultrasensitive and is achieved through the following decoy mechanism:[14] GLs 1 and 2 act as a decoy domains, competing with the regulatory domain, GL3, for Gαi inputs. This intramolecular decoy mechanism allows Pins to establish its threshold and steepness in response to distinct Gαi concentration. At low Gαi inputs, the decoy GLs 1 and 2 are preferentially bound. At intermediate Gαi concentration, the decoys are nearly saturated, and GL3 begins to be populated. At higher Gαi concentration, the decoys are fully saturated and Gαi binds to GL3, leading to Pins activation. Ultrasensitivity of Pins in response to Gαi ensures that Pins is activated only at the apical cortex where Gαi concentration is above the threshold, allowing for maximal Mud recruitment.[citation needed]
Switching Behavior of GTPases[edit]
GTPases are enzymes capable of binding and hydrolyzing guanosine triphosphate (GTP). Small GTPases, such as Ran and Ras, can exist in either a GTP-bound form (active) or a GDP-bound form (inactive), and the conversion between these two forms grants them a switch-like behavior.[51] As such, small GTPases are involved in multiple cellular events, including nuclear translocation and signaling.[52] The transition between the active and inactive states is facilitated by guanine nucleotide exchange factors (GEFs) and GTPase activating proteins (GAPs).[53]
Computational studies on the switching behavior of GTPases have revealed that the GTPase-GAP-GEF system displays ultrasensitivity.[54] In their study, Lipshtat et al. simulated the effects of the levels of GEF and GAP activation on the Rap activation signaling network in response to signals from activated α2-adrenergic (α2R) receptors, which lead to degradation of the activated Rap GAP. They found that the switching behavior of Rap activation was ultrasensitive to changes in the concentration (i.e. amplitude) and the duration of the α2R signal, yielding Hill coefficients of nH=2.9 and nH=1.7, respectively (a Hill coefficient greater than nH=1 is characteristic of ultrasensitivity [55]). The authors confirmed this experimentally by treating neuroblasts with HU-210, which activates RAP through degradation of Rap GAP. Ultrasensitivity was observed both in a dose-dependent manner (nH=5±0.2), by treating cells with different HU-210 concentrations for a fixed time, and in a duration-dependent manner (nH=8.6±0.8), by treating cells with a fixed HU-210 concentration during varying times.[citation needed]
By further studying system, the authors determined that (the degree of responsiveness and ultrasensitivity) was heavily dependent on two parameters: the initial ratio of kGAP/kGEF, where the k's incorporate both the concentration of active GAP or GEF and their corresponding kinetic rates; and the signal impact, which is the product of the degradation rate of activated GAP and either the signal amplitude or the signal duration.[54] The parameter kGAP/kGEF affects the steepness of the transition from the two states of the GTPase switch, with higher values (~10) leading to ultrasensitivity. The signal impact affects the switching point. Therefore, by depending on the ratio of concentrations rather than on individual concentrations, the switch-like behavior of the system can also be displayed outside of the zero-order regime.[citation needed]
Ultrasensitivity and Neuronal Potentiation[edit]
Persistent stimulation at the neuronal synapse can lead to markedly different outcomes for the post-synaptic neuron. Extended weak signaling can result in long-term depression (LTD), in which activation of the post-synaptic neuron requires a stronger signal than before LTD was initiated. In contrast, long-term potentiation (LTP) occurs when the post-synaptic neuron is subjected to a strong stimulus, and this results in strengthening of the neural synapse (i.e., less neurotransmitter signal is required for activation).
In the CA1 region of the hippocampus, the decision between LTD and LTP is mediated solely by the level of intracellular at the post-synaptic dendritic spine. Low levels of (resulting from low-level stimulation) activates the protein phosphatase calcineurin, which induces LTD. Higher levels of results in activation of /calmodulin-dependent protein kinase II (CaMKII), which leads to LTP. The difference in Ca2+ concentration required for a cell to undergo LTP is only marginally higher than for LTD, and because neurons show bistability (either LTP or LTD) following persistent stimulation, this suggests that one or more components of the system respond in a switch-like, or ultrasensitive manner. Bradshaw et al. demonstrated that CaMKII (the LTP inducer) responds to intracellular calcium levels in an ultrasensitive manner, with <10% activity at 1.0 uM and ~90% activity at 1.5 uM, resulting in a Hill coefficient of ~8. Further experiments showed that this ultrasenstivity was mediated by cooperative binding of CaMKII by two molecules of calmodulin (CaM), and autophosphorylation of activated CaMKII leading to a positive feedback loop.[56]
In this way, intracellular calcium can induce a graded, non-ultrasensitive activation of calcineurin at low levels, leading to LTD, whereas the ultrasensitive activation of CaMKII results in a threshold intracellular calcium level that generates a positive feedback loop that amplifies the signal and leads to the opposite cellular outcome: LTP. Thus, binding of a single substrate to multiple enzymes with different sensitivities facilitates a bistable decision for the cell to undergo LTD or LTP.[citation needed]
Ultrasensitivity in Development[edit]
It has been suggested that zero-order ultrasensitivity may generate thresholds during development allowing for the conversion of a graded morphogen input to a binary switch-like response.[57] Melen et al. (2005) have found evidence for such a system in the patterning of the Drosophila embryonic ventral ectoderm.[58] In this system, graded mitogen activated protein kinase (MAPK) activity is converted to a binary output, the all-or-none degradation of the Yan transcriptional repressor. They found that MAPK phosphorylation of Yan is both essential and sufficient for Yan's degradation. Consistent with zero-order ultrasensitivity an increase in Yan protein lengthened the time required for degradation but had no effect on the border of Yan degradation in developing embryos. Their results are consistent with a situation where a large pool of Yan becomes either completely degraded or maintained. The particular response of each cell depends on whether or not the rate of reversible Yan phosphorylation by MAPK is greater or less than dephosphorylation. Thus, a small increase in MAPK phosphorylation can cause it to be the dominant process in the cell and lead to complete degradation of Yan.
Multistep-feedback loop mechanism also leads to ultrasensitivity[edit]
Multistep-feedback loop mechanism also leads to ultrasensitivity. There is paper introducing that engineering synthetic feedback loops using yeast mating mitogen-activated protein (MAP) kinase pathway as a model system.
In Yeast mating pathway: alpha-factor activates receptor, Ste2, and Ste4 and activated Ste4 recruits Ste5 complex to membrane, allowing PAK-like kinase Ste20 (membrane-localized) to activate MAPKKK Ste11. Ste11 and downstream kinases, Ste7 (MAPKK) and Fus3 (MAPK), are colocalized on the scaffold and activation of cascade leads to transcriptional program. They used pathway modulators outside of core cascade, Ste50 promotes activation of Ste11 by Ste20; Msg5 (negative, red) is MAPK phosphatase that deactivates Fus3 (Fig.2A).
What they built was circuit with enhanced ultrasensitive switch behavior by constitutively expressing a negative modulator, Msg5 which is one of MAPK phosphatase and inducibly expressing a positive modulator, Ste50 which is pathway modulators outside of core cascade(Fig.2B). The success of this recruitment-based engineering strategy suggests that it may be possible to reprogram cellular responses with high precision.[59]
Flagellar motors and chemotaxis[edit]
The rotational direction of E. coli is controlled by the flagellar motor switch. A ring of 34 FliM proteins around the rotor bind CheY, whose phosphorylation state determines whether the motor rotates in a clockwise or counterclockwise manner. The rapid switching mechanism is attributed to an ultrasensitive response, which has a Hill coefficient of ~10. This system has been proposed to follow a dissipative allosteric model, in which rotational switching is a result of both CheY binding and energy consumption from the proton motive force, which also powers the flagellar rotation.[60]
Development of a Synthetic Ultrasensitive Signaling Pathway[edit]
Recently it has been shown that a Michaelian signaling pathway can be converted to an ultrasensitive signaling pathway by the introduction of two positive feedback loops.[61] In this synthetic biology approach, Palani and Sarkar began with a linear, graded response pathway, a pathway that showed a proportional increase in signal output relative to the amount of signal input, over a certain range of inputs. This simple pathway was composed of a membrane receptor, a kinase and a transcription factor. Upon activation the membrane receptor phosphorylates the kinase, which moves into the nucleus and phosphorylates the transcription factor, which turns on gene expression. To transform this graded response system into an ultrasensitive, or switch-like signaling pathway, the investigators created two positive feedback loops. In the engineered system, activation of the membrane receptor resulted in increased expression of both the receptor itself and the transcription factor. This was accomplished by placing a promoter specific for this transcription factor upstream of both genes. The authors were able to demonstrate that the synthetic pathway displayed high ultrasensitivity and bistability.
Recent computational analysis of the effects of a signaling protein's concentration on the presence of an ultrasensitive response has come to complementary conclusions about the influence of a signaling protein's concentration on the conversion of a graded response to an ultrasensitive one. Rather than focus on the generation of signaling proteins through positive feedback, however, the study instead focused on how the dynamics of a signaling protein's exit from the system influences the response. Soyer, Kuwahara, and Csika´sz-Nagy[62] devised a signaling pathway composed of a protein (P) that possesses two possible states (unmodified P or modified P*) and can be modified by an incoming stimulus E. Furthermore, while the unmodified form, P, is permitted to enter or leave the system, P* is only allowed to leave (i.e. it is not generated elsewhere). After varying the parameters of this system, the researchers discovered that the modification of P to P* can shift between a graded response and an ultrasensitive response via the modification of the exit rates of P and P* relative to each other. The transition from an ultrasensitive response to E and a graded response to E was generated when the two rates went from highly similar to highly dissimilar, irrespective of the kinetics of the conversion from P to P* itself. This finding suggests at least two things: 1) the simplifying assumption that the levels of signaling molecules stay constant in a system can severely limit the understanding of ultrasensitivity's complexity; and 2) it may be possible to induce or inhibit ultrasensitivity artificially by regulating the rates of the entry and exit of signaling molecules occupying a system of interest.
Limitations in Modularity[edit]
It has been shown that the integration of a given synthetic ultrasensitive module with upstream and downstream components often alters its information-processing capabilities.[33] This effects must be taken into account in the design process.
See also[edit]
References[edit]
- ^ Ferrell Jr, JE; Machleder, EM (1998). "The biochemical basis of an all-or-none cell fate switch in Xenopus oocytes". Science. 280 (5365): 895–8. Bibcode:1998Sci...280..895F. doi:10.1126/science.280.5365.895. PMID 9572732.
- ^ Mutalik, VK; Venkatesh, KV (2005). "Quantification of the glycogen cascade system: The ultrasensitive responses of liver glycogen synthase and muscle phosphorylase are due to distinctive regulatory designs". Theoretical Biology & Medical Modelling. 2: 19. doi:10.1186/1742-4682-2-19. PMC 1180476. PMID 15907212.
- ^ Greenwald, EC; Saucerman, JJ (2011). "Bigger, Better, Faster: Principles and Models of AKAP Anchoring Protein Signaling". Journal of Cardiovascular Pharmacology. 58 (5): 462–9. doi:10.1097/FJC.0b013e31822001e3. PMC 3173587. PMID 21562426.
- ^ a b c d e f Ferrell Jr, JE (1998). "How regulated protein translocation can produce switch-like responses". Trends in Biochemical Sciences. 23 (12): 461–5. doi:10.1016/S0968-0004(98)01316-4. PMID 9868363.
- ^ a b c Goldbeter, Albert; Koshland, Daniel E. (1981). "An Amplified Sensitivity Arising from Covalent Modification in Biological Systems". Proceedings of the National Academy of Sciences of the United States of America. 78 (11): 6840–6844. Bibcode:1981PNAS...78.6840G. doi:10.1073/pnas.78.11.6840. JSTOR 11361. PMC 349147. PMID 6947258.
- ^ Joh, RI; Weitz, JS (2011). o. Wilke, Claus (ed.). "To lyse or not to lyse: Transient-mediated stochastic fate determination in cells infected by bacteriophages". PLOS Computational Biology. 7 (3): e1002006. Bibcode:2011PLSCB...7E0020J. doi:10.1371/journal.pcbi.1002006. PMC 3053317. PMID 21423715.

- ^ Chatterjee, A; Kaznessis, YN; Hu, WS (2008). "Tweaking biological switches through a better understanding of bistability behavior". Current Opinion in Biotechnology. 19 (5): 475–81. doi:10.1016/j.copbio.2008.08.010. PMC 2766094. PMID 18804166.
- ^ Legewie, S; Blüthgen, N; Herzel, H (2006). "Mathematical modeling identifies inhibitors of apoptosis as mediators of positive feedback and bistability". PLOS Computational Biology. 2 (9): e120. Bibcode:2006PLSCB...2..120L. doi:10.1371/journal.pcbi.0020120. PMC 1570177. PMID 16978046.

- ^ Cui, J; Chen, C; Lu, H; Sun, T; Shen, P (2008). Hatakeyama, Mariko (ed.). "Two independent positive feedbacks and bistability in the Bcl-2 apoptotic switch". PLOS ONE. 3 (1): e1469. Bibcode:2008PLoSO...3.1469C. doi:10.1371/journal.pone.0001469. PMC 2194625. PMID 18213378.

- ^ Srividhya, Jeyaraman; Li, Yongfeng; Pomerening, Joseph R (2011). "Open cascades as simple solutions to providing ultrasensitivity and adaptation in cellular signaling". Physical Biology. 8 (4): 046005. Bibcode:2011PhBio...8d6005S. doi:10.1088/1478-3975/8/4/046005. PMC 3151678. PMID 21566270.
- ^ Shah, Najaf A.; Sarkar, Casim A. (2011). Haugh, Jason M. (ed.). "Robust Network Topologies for Generating Switch-Like Cellular Responses". PLOS Computational Biology. 7 (6): e1002085. Bibcode:2011PLSCB...7E2085S. doi:10.1371/journal.pcbi.1002085. PMC 3121696. PMID 21731481.

- ^ a b Thattai, M; Van Oudenaarden, A (2002). "Attenuation of noise in ultrasensitive signaling cascades". Biophysical Journal. 82 (6): 2943–50. Bibcode:2002BpJ....82.2943T. doi:10.1016/S0006-3495(02)75635-X. PMC 1302082. PMID 12023217.
- ^ Markevich, NI; Hoek, JB; Kholodenko, BN (2004). "Signaling switches and bistability arising from multisite phosphorylation in protein kinase cascades". The Journal of Cell Biology. 164 (3): 353–9. doi:10.1083/jcb.200308060. PMC 2172246. PMID 14744999.
- ^ a b Smith, Nicholas R.; Prehoda, Kenneth E. (2011). "Robust Spindle Alignment in Drosophila Neuroblasts by Ultrasensitive Activation of Pins". Molecular Cell. 43 (4): 540–9. doi:10.1016/j.molcel.2011.06.030. PMC 3161515. PMID 21855794.
- ^ Kim, Sun Young; Ferrell, James E. (2007). "Substrate Competition as a Source of Ultrasensitivity in the Inactivation of Wee1". Cell. 128 (6): 1133–45. doi:10.1016/j.cell.2007.01.039. PMID 17382882.
- ^ Huang, C. Y.; Ferrell Jr, J. E. (1996). "Ultrasensitivity in the mitogen-activated protein kinase cascade". Proceedings of the National Academy of Sciences of the United States of America. 93 (19): 10078–10083. Bibcode:1996PNAS...9310078H. doi:10.1073/pnas.93.19.10078. PMC 38339. PMID 8816754.
- ^ a b Sneppen, Kim; Micheelsen, Mille A; Dodd, Ian B (2008). "Ultrasensitive gene regulation by positive feedback loops in nucleosome modification". Molecular Systems Biology. 4 (1): 182. doi:10.1038/msb.2008.21. PMC 2387233. PMID 18414483.
- ^ a b Goldbeter, A; Koshland Jr, DE (1984). "Ultrasensitivity in biochemical systems controlled by covalent modification. Interplay between zero-order and multistep effects". The Journal of Biological Chemistry. 259 (23): 14441–7. doi:10.1016/S0021-9258(17)42619-6. PMID 6501300.
- ^ Kalir, S; McClure, J; Pabbaraju, K; Southward, C; Ronen, M; Leibler, S; Surette, MG; Alon, U (2001). "Ordering genes in a flagella pathway by analysis of expression kinetics from living bacteria". Science. 292 (5524): 2080–3. doi:10.1126/science.1058758. PMID 11408658. S2CID 14396458.
- ^ Dushek, O; Van Der Merwe, PA; Shahrezaei, V (2011). "Ultrasensitivity in multisite phosphorylation of membrane-anchored proteins". Biophysical Journal. 100 (5): 1189–97. Bibcode:2011BpJ...100.1189D. doi:10.1016/j.bpj.2011.01.060. PMC 3043222. PMID 21354391.
- ^ McCarrey, JR; Riggs, AD (1986). "Determinator-inhibitor pairs as a mechanism for threshold setting in development: A possible function for pseudogenes". Proceedings of the National Academy of Sciences of the United States of America. 83 (3): 679–83. Bibcode:1986PNAS...83..679M. doi:10.1073/pnas.83.3.679. PMC 322927. PMID 2418440.
- ^ a b c Buchler, NE; Louis, M (2008). "Molecular titration and ultrasensitivity in regulatory networks". Journal of Molecular Biology. 384 (5): 1106–19. doi:10.1016/j.jmb.2008.09.079. PMID 18938177.
- ^ Buchler, NE; Cross, FR (2009). "Protein sequestration generates a flexible ultrasensitive response in a genetic network". Molecular Systems Biology. 5 (1): 272. doi:10.1038/msb.2009.30. PMC 2694680. PMID 19455136.
- ^ Schmidt-Glenewinkel, H; Vacheva, I; Hoeller, D; Dikic, I; Eils, R (2008). "An ultrasensitive sorting mechanism for EGF receptor endocytosis". BMC Systems Biology. 2: 32. doi:10.1186/1752-0509-2-32. PMC 2377235. PMID 18394191.
- ^ a b Goldbeter, Albert (2005). "Zero-order switches and developmental thresholds". Molecular Systems Biology. 1 (1): E1–E2. doi:10.1038/msb4100042. PMC 1681457. PMID 16729066.
- ^ Meinke, Marilyn H.; Jonathan S. Bishop; Ronald D. Edstrom (1986). "Zero-order ultrasensitivity in the regulation of glycogen phosphorylase". PNAS. 83 (9): 2865–2868. Bibcode:1986PNAS...83.2865M. doi:10.1073/pnas.83.9.2865. PMC 323407. PMID 3458247.
- ^ Goulev, Youlian; Charvin, Gilles (2011). "Ultrasensitivity and Positive Feedback to Promote Sharp Mitotic Entry". Molecular Cell. 41 (3): 243–4. doi:10.1016/j.molcel.2011.01.016. PMID 21292155.
- ^ Trunnell, Nicole B.; Poon, Andy C.; Kim, Sun Young; Ferrell, James E. (2011). "Ultrasensitivity in the Regulation of Cdc25C by Cdk1". Molecular Cell. 41 (3): 263–74. doi:10.1016/j.molcel.2011.01.012. PMC 3060667. PMID 21292159.
- ^ a b Klein, Peter; Pawson, Tony; Tyers, Mike (2003). "Mathematical Modeling Suggests Cooperative Interactions between a Disordered Polyvalent Ligand and a Single Receptor Site". Current Biology. 13 (19): 1669–78. doi:10.1016/j.cub.2003.09.027. PMID 14521832.
- ^ Ravid, Tommer; Hochstrasser, Mark (2008). "Diversity of degradation signals in the ubiquitin–proteasome system". Nature Reviews Molecular Cell Biology. 9 (9): 679–89. doi:10.1038/nrm2468. PMC 2606094. PMID 18698327.
- ^ Kõivomägi, Mardo; Valk, Ervin; Venta, Rainis; Iofik, Anna; Lepiku, Martin; Balog, Eva Rose M.; Rubin, Seth M.; Morgan, David O.; Loog, Mart (2011). "Cascades of multisite phosphorylation control Sic1 destruction at the onset of S phase". Nature. 480 (7375): 128–31. Bibcode:2011Natur.480..128K. doi:10.1038/nature10560. PMC 3228899. PMID 21993622.
- ^ a b Dushek, Omer; Van Der Merwe, P.Anton; Shahrezaei, Vahid (2011). "Ultrasensitivity in Multisite Phosphorylation of Membrane-Anchored Proteins". Biophysical Journal. 100 (5): 1189–97. Bibcode:2011BpJ...100.1189D. doi:10.1016/j.bpj.2011.01.060. PMC 3043222. PMID 21354391.
- ^ a b c Altszyler, E; Ventura, A. C.; Colman-Lerner, A.; Chernomoretz, A. (2014). "Impact of upstream and downstream constraints on a signaling module's ultrasensitivity". Physical Biology. 11 (6): 066003. Bibcode:2014PhBio..11f6003A. doi:10.1088/1478-3975/11/6/066003. PMC 4233326. PMID 25313165.
- ^ a b Bluethgen, Nils; Legewie, Stefan; Herzel, Hanspeter; Kholodenko, Boris (2007). "Mechanisms Generating Ultrasensitivity, Bistability, and Oscillations in Signal Transduction". Introduction to Systems Biology. Humana Press. pp. 282–99. doi:10.1007/978-1-59745-531-2_15. ISBN 978-1-58829-706-8.
- ^ a b c d Huang, CY; Ferrell Jr, JE (1996). "Ultrasensitivity in the mitogen-activated protein kinase cascade". Proceedings of the National Academy of Sciences of the United States of America. 93 (19): 10078–83. Bibcode:1996PNAS...9310078H. doi:10.1073/pnas.93.19.10078. PMC 38339. PMID 8816754.
- ^ Kholodenko, Boris N.; et al. (1997). "Quantification of information transfer via cellular signal transduction pathways". FEBS Letters. 414 (2): 430–434. doi:10.1016/S0014-5793(97)01018-1. PMID 9315734.
- ^ a b c Altszyler, E; Ventura, A. C.; Colman-Lerner, A.; Chernomoretz, A. (2017). "Ultrasensitivity in signaling cascades revisited: Linking local and global ultrasensitivity estimations". PLOS ONE. 12 (6): e0180083. arXiv:1608.08007. Bibcode:2017PLoSO..1280083A. doi:10.1371/journal.pone.0180083. PMC 5491127. PMID 28662096.
- ^ Brown, GC; Hoek, J B; Kholodenko B, N (1997). "Why do protein kinase cascades have more than one level?". Trends Biochem. Sci. 22 (8): 288. doi:10.1016/s0968-0004(97)82216-5. PMID 9270298.
- ^ Ferrell, J E (1997). "How responses get more switch-like as you move down a protein kinase cascade". Trends Biochem. Sci. 22 (8): 288–289. doi:10.1016/s0968-0004(97)82217-7. PMID 9270299.
- ^ Racz,E; Slepchenko, B M (2008). "On sensitivity amplification in intracellular signaling cascades". Phys. Biol. 5 (3): 36004. Bibcode:2008PhBio...5c6004R. doi:10.1088/1478-3975/5/3/036004. PMC 2675913. PMID 18663279.
- ^ Dueber, John E; Mirsky, Ethan A; Lim, Wendell A (2007). "Engineering synthetic signaling proteins with ultrasensitive input/output control". Nature Biotechnology. 25 (6): 660–2. doi:10.1038/nbt1308. PMID 17515908. S2CID 10465894.
- ^ Toyoshima-Morimoto, F.; Taniguchi, E; Nishida, E (2002). "Plk1 promotes nuclear translocation of human Cdc25C during prophase". EMBO Reports. 3 (4): 341–8. doi:10.1093/embo-reports/kvf069. PMC 1084057. PMID 11897663.
- ^ Bahassi, EL Mustapha; Hennigan, Robert F; Myer, David L; Stambrook, Peter J (2004). "Cdc25C phosphorylation on serine 191 by Plk3 promotes its nuclear translocation". Oncogene. 23 (15): 2658–63. doi:10.1038/sj.onc.1207425. PMID 14968113.
- ^ Doe, CQ (2008). "Neural stem cells: Balancing self-renewal with differentiation". Development. 135 (9): 1575–87. doi:10.1242/dev.014977. PMID 18356248.
- ^ Yu, F; Cai, Y; Kaushik, R; Yang, X; Chia, W (2003). "Distinct roles of Galphai and Gbeta13F subunits of the heterotrimeric G protein complex in the mediation of Drosophila neuroblast asymmetric divisions". The Journal of Cell Biology. 162 (4): 623–33. doi:10.1083/jcb.200303174. PMC 2173805. PMID 12925708.
- ^ Izumi, Yasushi; Ohta, Nao; Hisata, Kanako; Raabe, Thomas; Matsuzaki, Fumio (2006). "Drosophila Pins-binding protein Mud regulates spindle-polarity coupling and centrosome organization". Nature Cell Biology. 8 (6): 586–93. doi:10.1038/ncb1409. PMID 16648846. S2CID 23909489.
- ^ Bowman, SK; Neumüller, RA; Novatchkova, M; Du, Q; Knoblich, JA (2006). "The Drosophila NuMA Homolog Mud regulates spindle orientation in asymmetric cell division". Developmental Cell. 10 (6): 731–42. doi:10.1016/j.devcel.2006.05.005. PMID 16740476.
- ^ Siller, KH; Cabernard, C; Doe, CQ (2006). "The NuMA-related Mud protein binds Pins and regulates spindle orientation in Drosophila neuroblasts". Nature Cell Biology. 8 (6): 594–600. doi:10.1038/ncb1412. PMID 16648843. S2CID 2455200.
- ^ Du, Q; MacAra, IG (2004). "Mammalian Pins is a conformational switch that links NuMA to heterotrimeric G proteins". Cell. 119 (4): 503–16. doi:10.1016/j.cell.2004.10.028. PMID 15537540. S2CID 12900159.
- ^ Nipper, RW; Siller, KH; Smith, NR; Doe, CQ; Prehoda, KE (2007). "Galphai generates multiple Pins activation states to link cortical polarity and spindle orientation in Drosophila neuroblasts". Proceedings of the National Academy of Sciences of the United States of America. 104 (36): 14306–11. Bibcode:2007PNAS..10414306N. doi:10.1073/pnas.0701812104. PMC 1964812. PMID 17726110.
- ^ Yang, Z (2002). "Small GTPases: Versatile signaling switches in plants". The Plant Cell. 14 Suppl (Suppl): S375–88. doi:10.1105/tpc.001065. PMC 151267. PMID 12045289.
- ^ Heider, D; Hauke, S; Pyka, M; Kessler, D (2010). "Insights into the classification of small GTPases". Advances and Applications in Bioinformatics and Chemistry. 3: 15–24. doi:10.2147/aabc.s8891. PMC 3170009. PMID 21918623.
- ^ Bourne, Henry R.; Sanders, David A.; McCormick, Frank (1991). "The GTPase superfamily: Conserved structure and molecular mechanism". Nature. 349 (6305): 117–27. Bibcode:1991Natur.349..117B. doi:10.1038/349117a0. PMID 1898771. S2CID 4349901.
- ^ a b Lipshtat, A.; Jayaraman, G.; He, J. C.; Iyengar, R. (2010). "Design of versatile biochemical switches that respond to amplitude, duration, and spatial cues". Proceedings of the National Academy of Sciences. 107 (3): 1247–52. Bibcode:2010PNAS..107.1247L. doi:10.1073/pnas.0908647107. PMC 2824311. PMID 20080566.
- ^ Ferrell, James E. (1999). "Building a cellular switch: More lessons from a good egg". BioEssays. 21 (10): 866–870. CiteSeerX 10.1.1.540.1905. doi:10.1002/(SICI)1521-1878(199910)21:10<866::AID-BIES9>3.0.CO;2-1. PMID 10497337.
- ^ Bradshaw, JM; Kubota, Y; Meyer, T; Schulman, H (2003). "An ultrasensitive Ca2+/calmodulin-dependent protein kinase II-protein phosphatase 1 switch facilitates specificity in postsynaptic calcium signaling". Proceedings of the National Academy of Sciences of the United States of America. 100 (18): 10512–7. Bibcode:2003PNAS..10010512B. doi:10.1073/pnas.1932759100. PMC 193592. PMID 12928489.
- ^ Goldbeter, A; Wolpert, L (1990). "Covalent modification of proteins as a threshold mechanism in development". Journal of Theoretical Biology. 142 (2): 243–50. Bibcode:1990JThBi.142..243G. doi:10.1016/s0022-5193(05)80225-5. PMID 2161972.
- ^ Melen, GJ; Levy, S; Barkai, N; Shilo, BZ (2005). "Threshold responses to morphogen gradients by zero-order ultrasensitivity". Molecular Systems Biology. 1 (1): 2005.0028. doi:10.1038/msb4100036. PMC 1681448. PMID 16729063.
- ^ Bashor, C. J.; Helman, N. C.; Yan, S.; Lim, W. A. (2008). "Using Engineered Scaffold Interactions to Reshape MAP Kinase Pathway Signaling Dynamics". Science. 319 (5869): 1539–43. Bibcode:2008Sci...319.1539B. doi:10.1126/science.1151153. PMID 18339942. S2CID 365578.
- ^ Tu, Y. (2008). "The nonequilibrium mechanism for ultrasensitivity in a biological switch: Sensing by Maxwell's demons". Proceedings of the National Academy of Sciences. 105 (33): 11737–11741. Bibcode:2008PNAS..10511737T. doi:10.1073/pnas.0804641105. JSTOR 25463752. PMC 2575293. PMID 18687900.
- ^ Palani, S; Sarkar, CA (2011). "Synthetic conversion of a graded receptor signal into a tunable, reversible switch". Molecular Systems Biology. 7 (1): 480. doi:10.1038/msb.2011.13. PMC 3094063. PMID 21451590.
- ^ Soyer, OS; Kuwahara, H; Csikász-Nagy, A (2009). "Regulating the total level of a signaling protein can vary its dynamics in a range from switch like ultrasensitivity to adaptive responses". The FEBS Journal. 276 (12): 3290–8. doi:10.1111/j.1742-4658.2009.07054.x. PMID 19438711.








![{\displaystyle {\begin{array}{rcrcr}\\W+E_{1}&{\ce {<=>[a_1][d_1]}}&WE_{1}&{\ce {->[k_1]}}&W'+E_{1}\\{}\\W'+E_{2}&{\ce {<=>[a_2][d_2]}}&W'E_{2}&{\ce {->[k_2]}}&W+E_{2}\\{}\end{array}}}](https://wikimedia.org/api/rest_v1/media/math/render/svg/b99db825c4ff803840dbb716ddf91cf25350aee2)
![{\displaystyle {\frac {d[W]}{dt}}=-a_{1}[W][E_{1}]+d_{1}[WE_{1}]+k_{2}[W'E_{2}]}](https://wikimedia.org/api/rest_v1/media/math/render/svg/8d94c07c264d2a743d2af4229c9cb395795430cf)
![{\displaystyle {\frac {d[WE_{1}]}{dt}}=a_{1}[W][E_{1}]-(d_{1}+k_{1})[WE_{1}]}](https://wikimedia.org/api/rest_v1/media/math/render/svg/240963c6e424e624af0df0c5fd9c408b2bf0dabc)
![{\displaystyle {\frac {d[W']}{dt}}=-a_{2}[W'][E_{2}]+d_{2}[W'E_{2}]+k1[WE_{1}]}](https://wikimedia.org/api/rest_v1/media/math/render/svg/26666a5dd399063d4c38813fb9853fd95de8e985)
![{\displaystyle {\frac {d[W'E_{2}]}{dt}}=a_{2}[W'][E_{2}]-(d_{2}+k_{2})[W'E_{2}]}](https://wikimedia.org/api/rest_v1/media/math/render/svg/5f2755c33130ae77257089cc6cd8edbda76ee4be)
![{\displaystyle [W_{T}]=[W]+[W']+[WE_{1}]+[W'E_{2}]}](https://wikimedia.org/api/rest_v1/media/math/render/svg/74f4c088eb41138b9bed456fffb71fb5a51bf293)
![{\displaystyle [E_{1T}]=[E_{1}]+[WE_{1}]}](https://wikimedia.org/api/rest_v1/media/math/render/svg/00df8b30b56b0b703a1f4bb53c2d70cdb4e63ac1)
![{\displaystyle [E_{2T}]=[E_{2}]+[W'E_{2}]}](https://wikimedia.org/api/rest_v1/media/math/render/svg/aa4d15730374a3959d60d78b8506d2ef6be59377)
![{\displaystyle [W_{T}]\gg [E_{1}]}](https://wikimedia.org/api/rest_v1/media/math/render/svg/f2b067fc15f94fb3ac4fcab8cf8c72a85f2a7126)
![{\displaystyle [E_{2}]}](https://wikimedia.org/api/rest_v1/media/math/render/svg/c800f8bc3fbf23eb2b33e87f84459309ac58a636)
![{\displaystyle [WE_{1}]}](https://wikimedia.org/api/rest_v1/media/math/render/svg/b17c8921bcdc4458ca5b2d4f792d05479279d34d)
![{\displaystyle [W'E_{2}]}](https://wikimedia.org/api/rest_v1/media/math/render/svg/4756360ea10e8a76069a18341737bfa9397a91ee)

![{\displaystyle W=[W]/[W_{T}]}](https://wikimedia.org/api/rest_v1/media/math/render/svg/b6f8bd24362ea433b22305591115664c6e6c7b4c)


![{\displaystyle k_{1}[WE_{1}]=k_{2}[W'E_{2}]}](https://wikimedia.org/api/rest_v1/media/math/render/svg/4e4e37afe68c067a0ee4e3fd0b8a055212e94edd)
![{\displaystyle V_{1}=k_{1}[E_{1T}],}](https://wikimedia.org/api/rest_v1/media/math/render/svg/763a5a97e5c465ea9bf779d5b3e41035a40e1528)
![{\displaystyle V_{2}=k_{2}[E_{2T}],}](https://wikimedia.org/api/rest_v1/media/math/render/svg/3a2d41808c31348290dd5cfba0c3d1516326f1aa)
![{\displaystyle K_{1}={\frac {d_{1}+k_{1}}{a_{1}[W_{T}]}}={\frac {K_{M1}}{[W_{T}]}},}](https://wikimedia.org/api/rest_v1/media/math/render/svg/bc8f2d2a0612bbf57fd2e27774ec3fde531dc387)
![{\displaystyle K_{2}={\frac {d_{2}+k_{2}}{a_{2}[W_{T}]}}={\frac {K_{M2}}{[W_{T}]}}.}](https://wikimedia.org/api/rest_v1/media/math/render/svg/82a9c2ed5487c0e38272fb390ac0ada579811ee1)


![{\displaystyle [L]}](https://wikimedia.org/api/rest_v1/media/math/render/svg/c1eba88cad4b17de726ed2c779b29eecdf5819a2)

![{\displaystyle \theta ={\frac {[L]^{n}}{K_{d}+[L]^{n}}}={\frac {[L]^{n}}{(K_{A})^{n}+[L]^{n}}},}](https://wikimedia.org/api/rest_v1/media/math/render/svg/ff9d9d9370beac68a7957b42a48f719478d612ed)





















![{\displaystyle [X10_{i},X90_{i}]}](https://wikimedia.org/api/rest_v1/media/math/render/svg/2a6a9f701cc2d5bd5754f6206af5ce8e2b644e8f)



