Coronavirus frameshifting stimulation element
| Coronavirus frameshifting stimulation element | |
|---|---|
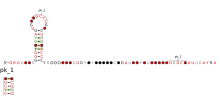 Predicted secondary structure and sequence conservation of Corona_FSE | |
| Identifiers | |
| Symbol | Corona_FSE |
| Rfam | RF00507 |
| Other data | |
| RNA type | Cis-reg; frameshift element |
| Domain(s) | Viruses |
| SO | SO:0001427 |
| PDB structures | PDBe |
In molecular biology, the coronavirus frameshifting stimulation element is a conserved stem-loop of RNA found in coronaviruses that can promote ribosomal frameshifting. Such RNA molecules interact with a downstream region to form a pseudoknot structure; the region varies according to the virus but pseudoknot formation is known to stimulate frameshifting. In the classical situation, a sequence 32 nucleotides downstream of the stem is complementary to part of the loop. In other coronaviruses, however, another stem-loop structure around 150 nucleotides downstream can interact with members of this family to form kissing stem-loops and stimulate frameshifting.[1]
Other RNA families identified in the coronavirus include the coronavirus 3′ stem-loop II-like motif (s2m), the coronavirus packaging signal and the coronavirus 3′ UTR pseudoknot.
During protein synthesis, rapidly changing conditions in the cell can cause ribosomal pausing. In coronaviruses, this can affect growth rate and trigger translational abandonment. This releases the ribosome from the mRNA and the incomplete polypeptide is targeted for destruction.[2]
See also
[edit]- Translational frameshift
- Slippery sequence
- Coronavirus 5′ UTR
- Coronavirus 3′ UTR
- Coronavirus 3′ UTR pseudoknot
- Coronavirus 3′ stem-loop II-like motif (s2m)
- Coronavirus packaging signal
References
[edit]- ^ Baranov PV, Henderson CM, Anderson CB, Gesteland RF, Atkins JF, Howard MT (February 2005). "Programmed ribosomal frameshifting in decoding the SARS-CoV genome". Virology. 332 (2): 498–510. doi:10.1016/j.virol.2004.11.038. PMC 7111862. PMID 15680415.
- ^ Buchan, J. R.; Stansfield, I. (2007). "Halting a cellular production line: responses to ribosomal pausing during translation". Biol Cell. 99 (9): 475–487. doi:10.1042/BC20070037. PMID 17696878.

