DREAM complex
The dimerization partner, RB-like, E2F and multi-vulval class B (DREAM) complex is a protein complex responsible for the regulation of cell cycle-dependent gene expression.[1][2] The complex is evolutionarily conserved, although some of its components vary from species to species. In humans, the key proteins in the complex are RBL1 (p107) and RBL2 (p130), both of which are homologs of RB (p105) and bind repressive E2F transcription factors E2F4 and E2F5; DP1, DP2 and DP3, dimerization partners of E2F; and MuvB, which is a complex of LIN9/37/52/54 and RBBP4.[1]
Discovery
[edit]Genes encoding the MuvB complex were originally identified from loss-of-function mutation studies in C. elegans. When mutated, these genes produced worms with multiple vulva-like organs, hence the name ‘Muv’. Three classes of Muv genes were classified, with class B genes encoding homologues of mammalian RB, E2F, and DP1, and others such as LIN-54, LIN-37, LIN-7 and LIN-52, whose functions were not yet understood.[3][4]
Studies in Drosophila melanogaster ovarian follicle cells identified a protein complex that bound to repeatedly amplifying chorion genes. The complex included genes that had close homology with the MuvB genes such as Mip130, Mip120 and Mip40. These Mip genes were identified as homologues of the MuvB genes LIN9, LIN54, and LIN37 respectively.[5] Further studies in the fly embryo nuclear extracts confirmed the coexistence of these proteins with others such as the RB homologues Rbf1 and Rbf2, and others like E2f and Dp. The protein complex was thus termed as the Drosophila RBF, E2f2 and Mip (dREAM) complex. Disruption of the dREAM complex through RNAi knockdown of the components of dREAM complex led to higher expression of E2f regulated genes that are typically silenced, implicating dREAM’s role in gene down-regulation.[6] Later in Drosophila melanogaster, there was also found a testis-specific paralog of the Myb-MuvB/DREAM complex known as tMAC (testis-specific meiotic arrest complex), which is involved in meiotic arrest.[7]
A protein complex similar to dREAM was subsequently identified in C. elegans extract containing DP, RB, and MuvB, and was named as DRM. This complex included mammalian homologues of RB and DP, and other members of the MuvB complex.[8]
The mammalian DREAM complex was identified following immunoprecipitation of p130 with mass-spectrometry analysis. The results showed that p130 was associated with E2F4, E2F5, the dimerization partner DP, and LIN9, LIN54, LIN37, LIN52, and RBBP4 that make up the MuvB complex. Immunoprecipitation of MuvB factors also revealed association of BMYB. Subsequent immunoprecipitation with BMYB yielded all the MuvB core proteins, but not other members of the DREAM complex – p130, p107, E2F4/5 and DP. This indicated that MuvB associated with BMYB to form the BMYB-MuvB complex or with p130/p107, E2F4/5 and DP to form the DREAM complex. The DREAM complex was found prevalent in quiescent or starved cells, and the BMYB-MuvB complex was found in actively dividing cells, hinting at separate functionalities of these two complexes.[9]
MuvB-like complexes were also recently discovered in Arabidoposis that include E2F and MYB orthologs combined with LIN9 and LIN54 orthologs.[10][11]
Function
[edit]The main function of the DREAM complex is to repress G1/S and G2/M gene expression during quiescence (G0). Entry into the cell cycle dissociates p130 from the complex and leads to subsequent recruitment of activating E2F proteins. This allows for the expression of E2F regulated late G1 and S phase genes. BMYB (MYBL2), which is repressed by the DREAM complex during G0 is also able to be expressed at this time, and binds to MuvB during S phase to promote the expression of key G2/M phase genes such as CDK1 and CCNB1. FOXM1 is then recruited in G2 to further promote gene expression (e.g. AURKA). During late S phase BMYB is degraded via CUL1 (SCF complex), while FOXM1 is degraded during mitosis by the APC/C.[1][12] Near the end of the cell cycle, the DREAM complex is re-assembled by DYRK1A to repress G1/S and G2/M genes.
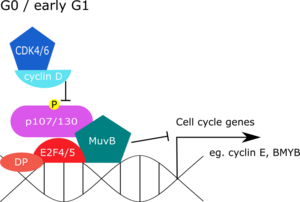
G0
[edit]During quiescence, the DREAM complex represses G1/S and G2/M gene expression. In mammalian systems, chromatin-immunoprecipitation (ChIP) studies have revealed that DREAM components are found together at promoters of genes that peak in G1/S or G2/M phase.[9] Abrogation of the DREAM complex on the other hand, led to increased expression of E2F regulated genes normally repressed in the G0 phase.[9][13] Contrary to mammalian cells, the fly dREAM complex was found at almost one-third of all promoters, which may reflect a broader role for dREAM in gene regulation, such as programmed cell death of neural precursor cells.[14][15]
Docking of the DREAM complex to promoters is achieved by binding of LIN-54 to regions known as cell cycle genes homology region (CHR). These are specific sequence of nucleotides that are commonly found in the promoters of genes expressed during late S phase or G2/M phase. Docking can also be achieved via E2F proteins binding to sequences known as cell cycle-dependent element sites (CDEs). Some cell cycle dependent genes have been found where both CHRs and CDEs are in proximity to one another. Because p130-E2F4 can form stable associations with the MuvB complex, the proximity of CHRs to CDEs suggests that affinity of binding of the DREAM complex to target genes is cooperatively improved by association with both the binding sites.[16]
When DREAM is docked onto the promoter, p130 is bound to LIN52, and this association inhibits LIN52 binding to chromatin modifier proteins.[17][18] Therefore, unlike RB-E2F, the DREAM complex is unlikely to directly recruit chromatin modifiers to repress gene expression, although some associations have been suggested.[19][20] DREAM complex may instead down-regulate gene expression by affecting nucleosome positioning. Compacted DNA at transcription start sites inhibit gene expression by blocking the docking of RNA polymerase.[21] In worms for example, loss of a MuvB complex protein, LIN35, leads to loss of repressive histone associations and high expression of cell cycle dependent genes. However, direct evidence for the link between repressive histones and the DREAM complex remains to be elucidated.[22]

G1/S
[edit]Like its counterpart, RB-E2F, the DREAM complex is also affected by similar growth stimuli and subsequent cyclin-CDK activity. Increasing cyclin D-CDK4 and cyclin E-CDK2 activity dissociates the DREAM complex from the promoter by phosphorylation of p130.[18] Hyper-phosphorylated p130 is subsequently degraded [23][24] and E2F4 exported from the nucleus.[25] Once the repressive E2Fs are vacated, activating E2Fs bind to the promoter to up-regulate G1/S genes that promote DNA synthesis and transition of the cell cycle.[26] BMYB is also up-regulated during this time, which then binds to genes that peak at G2/M phase.[9][27][28] Binding of BMYB to late cell cycle genes is dependent on its association with the MuvB core to form the BMYB-MuvB complex, which is then able to up-regulate genes in the G2/M phase.[12]
Late mitosis
[edit]Near the end of mitosis, p130 and p107 are dephosphorylated from their hyperphosphorylated state by the phosphatase PP2a.[29][30] Inhibition of PP2a activity reduced promoter binding of some of the proteins of the DREAM complex in the subsequent G1 phase and de-repression of gene expression.[31]
Other components have been shown to be phosphorylated for DREAM complex assembly to occur. Of these, LIN52 phosphorylation on its S28 residue is the most well-understood. Substitution of this serine to alanine led to reduced binding of the MuvB core to p130 and impaired the ability of cells to enter quiescence. This indicates that LIN52 S28 phosphorylation is required for proper association and function of the DREAM complex via binding with p130. One known regulator of phosphorylation of the S28 residue is the DYRK1A. The loss of this kinase leads to decreased phosphorylation of the S28 residue and association of p130 with MuvB.[13] DYRK1A was also found to degrade cyclin D1, which would increase p21 levels – both of which contribute to cell cycle exit.[32]
The DREAM complex was also shown to regulate cytokinesis through GAS2L3.[33]
Cancer therapy
[edit]Due to its regulatory role in the cell cycle, targeting the DREAM complex might enhance anticancer treatments such as imatinib.[34][35]
See also
[edit]References
[edit]- ^ a b c Sadasivam, Subhashini; DeCaprio, James A. (11 July 2013). "The DREAM complex: master coordinator of cell cycle-dependent gene expression". Nature Reviews Cancer. 13 (8): 585–595. doi:10.1038/nrc3556. PMC 3986830. PMID 23842645.
- ^ Fischer, M; Müller, GA (December 2017). "Cell cycle transcription control: DREAM/MuvB and RB-E2F complexes". Critical Reviews in Biochemistry and Molecular Biology. 52 (6): 638–662. doi:10.1080/10409238.2017.1360836. PMID 28799433. S2CID 205695213.
- ^ Beitel, GJ; Lambie, EJ; Horvitz, HR (2000-08-22). "The C. elegans gene lin-9, which acts in an Rb-related pathway, is required for gonadal sheath cell development and encodes a novel protein". Gene. 254 (1–2): 253–63. doi:10.1016/s0378-1119(00)00296-1. PMID 10974557.
- ^ Thomas, JH; Ceol, CJ; Schwartz, HT; Horvitz, HR (May 2003). "New genes that interact with lin-35 Rb to negatively regulate the let-60 ras pathway in Caenorhabditis elegans". Genetics. 164 (1): 135–51. doi:10.1093/genetics/164.1.135. PMC 1462563. PMID 12750327.
- ^ Beall, EL; Manak, JR; Zhou, S; Bell, M; Lipsick, JS; Botchan, MR (2002-12-19). "Role for a Drosophila Myb-containing protein complex in site-specific DNA replication". Nature. 420 (6917): 833–7. Bibcode:2002Natur.420..833B. doi:10.1038/nature01228. PMID 12490953. S2CID 4425307.
- ^ Korenjak, M; Taylor-Harding, B; Binné, UK; Satterlee, JS; Stevaux, O; Aasland, R; White-Cooper, H; Dyson, N; Brehm, A (2004-10-15). "Native E2F/RBF complexes contain Myb-interacting proteins and repress transcription of developmentally controlled E2F target genes". Cell. 119 (2): 181–93. doi:10.1016/j.cell.2004.09.034. PMID 15479636. S2CID 17989678.
- ^ Beall, E. L.; Lewis, P. W.; Bell, M.; Rocha, M.; Jones, D. L.; Botchan, M. R. (15 April 2007). "Discovery of tMAC: a Drosophila testis-specific meiotic arrest complex paralogous to Myb-Muv B". Genes & Development. 21 (8): 904–919. doi:10.1101/gad.1516607. PMC 1847709. PMID 17403774.
- ^ Harrison, MM; Ceol, CJ; Lu, X; Horvitz, HR (2006-11-07). "Some C. elegans class B synthetic multivulva proteins encode a conserved LIN-35 Rb-containing complex distinct from a NuRD-like complex". Proceedings of the National Academy of Sciences of the United States of America. 103 (45): 16782–7. Bibcode:2006PNAS..10316782H. doi:10.1073/pnas.0608461103. PMC 1636532. PMID 17075059.
- ^ a b c d Litovchick, L; Sadasivam, S; Florens, L; Zhu, X; Swanson, SK; Velmurugan, S; Chen, R; Washburn, MP; Liu, XS; DeCaprio, JA (2007-05-25). "Evolutionarily conserved multisubunit RBL2/p130 and E2F4 protein complex represses human cell cycle-dependent genes in quiescence". Molecular Cell. 26 (4): 539–51. doi:10.1016/j.molcel.2007.04.015. PMID 17531812.
- ^ Kobayashi, K; Suzuki, T; Iwata, E; Nakamichi, N; Suzuki, T; Chen, P; Ohtani, M; Ishida, T; Hosoya, H; Müller, S; Leviczky, T; Pettkó-Szandtner, A; Darula, Z; Iwamoto, A; Nomoto, M; Tada, Y; Higashiyama, T; Demura, T; Doonan, JH; Hauser, MT; Sugimoto, K; Umeda, M; Magyar, Z; Bögre, L; Ito, M (2015-08-04). "Transcriptional repression by MYB3R proteins regulates plant organ growth". The EMBO Journal. 34 (15): 1992–2007. doi:10.15252/embj.201490899. PMC 4551348. PMID 26069325.
- ^ Lang, Lucas; Pettkó-Szandtner, Aladár; Tunçay Elbaşı, Hasibe; Takatsuka, Hirotomo; Nomoto, Yuji; Zaki, Ahmad; Dorokhov, Stefan; De Jaeger, Geert; Eeckhout, Dominique; Ito, Masaki; Magyar, Zoltán; Bögre, László; Heese, Maren; Schnittger, Arp (December 2021). "The DREAM complex represses growth in response to DNA damage in Arabidopsis". Life Science Alliance. 4 (12): e202101141. doi:10.26508/lsa.202101141. PMC 8500230. PMID 34583930.
- ^ a b Sadasivam, S.; Duan, S.; DeCaprio, J. A. (5 March 2012). "The MuvB complex sequentially recruits B-Myb and FoxM1 to promote mitotic gene expression". Genes & Development. 26 (5): 474–489. doi:10.1101/gad.181933.111. PMC 3305985. PMID 22391450.
- ^ a b Litovchick, L; Florens, LA; Swanson, SK; Washburn, MP; DeCaprio, JA (2011-04-15). "DYRK1A protein kinase promotes quiescence and senescence through DREAM complex assembly". Genes & Development. 25 (8): 801–13. doi:10.1101/gad.2034211. PMC 3078706. PMID 21498570.
- ^ Georlette, D; Ahn, S; MacAlpine, DM; Cheung, E; Lewis, PW; Beall, EL; Bell, SP; Speed, T; Manak, JR; Botchan, MR (2007-11-15). "Genomic profiling and expression studies reveal both positive and negative activities for the Drosophila Myb MuvB/dREAM complex in proliferating cells". Genes & Development. 21 (22): 2880–96. doi:10.1101/gad.1600107. PMC 2049191. PMID 17978103.
- ^ Rovani, Margritte K.; Brachmann, Carrie Baker; Ramsay, Gary; Katzen, Alisa L. (December 2012). "The dREAM/Myb–MuvB complex and Grim are key regulators of the programmed death of neural precursor cells at the Drosophila posterior wing margin". Developmental Biology. 372 (1): 88–102. doi:10.1016/j.ydbio.2012.08.022. PMC 3621911. PMID 22960039.
- ^ Müller, GA; Engeland, K (February 2010). "The central role of CDE/CHR promoter elements in the regulation of cell cycle-dependent gene transcription". The FEBS Journal. 277 (4): 877–93. doi:10.1111/j.1742-4658.2009.07508.x. PMID 20015071. S2CID 8955433.
- ^ Forristal, C; Henley, SA; MacDonald, JI; Bush, JR; Ort, C; Passos, DT; Talluri, S; Ishak, CA; Thwaites, MJ; Norley, CJ; Litovchick, L; DeCaprio, JA; DiMattia, G; Holdsworth, DW; Beier, F; Dick, FA (June 2014). "Loss of the mammalian DREAM complex deregulates chondrocyte proliferation". Molecular and Cellular Biology. 34 (12): 2221–34. doi:10.1128/MCB.01523-13. PMC 4054284. PMID 24710275.
- ^ a b Guiley, KZ; Liban, TJ; Felthousen, JG; Ramanan, P; Litovchick, L; Rubin, SM (2015-05-01). "Structural mechanisms of DREAM complex assembly and regulation". Genes & Development. 29 (9): 961–74. doi:10.1101/gad.257568.114. PMC 4421984. PMID 25917549.
- ^ Sandoval, R; Pilkinton, M; Colamonici, OR (2009-10-15). "Deletion of the p107/p130-binding domain of Mip130/LIN-9 bypasses the requirement for CDK4 activity for the dissociation of Mip130/LIN-9 from p107/p130-E2F4 complex". Experimental Cell Research. 315 (17): 2914–20. doi:10.1016/j.yexcr.2009.07.014. PMC 2757496. PMID 19619530.
- ^ Stiegler, P; De Luca, A; Bagella, L; Giordano, A (1998-11-15). "The COOH-terminal region of pRb2/p130 binds to histone deacetylase 1 (HDAC1), enhancing transcriptional repression of the E2F-dependent cyclin A promoter". Cancer Research. 58 (22): 5049–52. PMID 9823308.
- ^ Bai, L; Morozov, AV (November 2010). "Gene regulation by nucleosome positioning". Trends in Genetics. 26 (11): 476–83. doi:10.1016/j.tig.2010.08.003. PMID 20832136.
- ^ Latorre, I; Chesney, MA; Garrigues, JM; Stempor, P; Appert, A; Francesconi, M; Strome, S; Ahringer, J (2015-03-01). "The DREAM complex promotes gene body H2A.Z for target repression". Genes & Development. 29 (5): 495–500. doi:10.1101/gad.255810.114. PMC 4358402. PMID 25737279.
- ^ Tedesco, D; Lukas, J; Reed, SI (2002-11-15). "The pRb-related protein p130 is regulated by phosphorylation-dependent proteolysis via the protein-ubiquitin ligase SCF(Skp2)". Genes & Development. 16 (22): 2946–57. doi:10.1101/gad.1011202. PMC 187481. PMID 12435635.
- ^ Bhattacharya, S; Garriga, J; Calbó, J; Yong, T; Haines, DS; Graña, X (2003-04-24). "SKP2 associates with p130 and accelerates p130 ubiquitylation and degradation in human cells". Oncogene. 22 (16): 2443–51. doi:10.1038/sj.onc.1206339. PMID 12717421. S2CID 26125392.
- ^ Gaubatz, S; Lees, JA; Lindeman, GJ; Livingston, DM (February 2001). "E2F4 is exported from the nucleus in a CRM1-dependent manner". Molecular and Cellular Biology. 21 (4): 1384–92. doi:10.1128/MCB.21.4.1384-1392.2001. PMC 99590. PMID 11158323.
- ^ Takahashi, Y; Rayman, JB; Dynlacht, BD (2000-04-01). "Analysis of promoter binding by the E2F and pRB families in vivo: distinct E2F proteins mediate activation and repression". Genes & Development. 14 (7): 804–16. doi:10.1101/gad.14.7.804. PMC 316494. PMID 10766737.
- ^ Lam, EW; Robinson, C; Watson, RJ (September 1992). "Characterization and cell cycle-regulated expression of mouse B-myb". Oncogene. 7 (9): 1885–90. PMID 1501895.
- ^ Pilkinton, M; Sandoval, R; Song, J; Ness, SA; Colamonici, OR (2007-01-05). "Mip/LIN-9 regulates the expression of B-Myb and the induction of cyclin A, cyclin B, and CDK1". The Journal of Biological Chemistry. 282 (1): 168–75. doi:10.1074/jbc.M609924200. PMID 17098733. S2CID 21963932.
- ^ Kolupaeva, V; Janssens, V (January 2013). "PP1 and PP2A phosphatases--cooperating partners in modulating retinoblastoma protein activation". The FEBS Journal. 280 (2): 627–43. doi:10.1111/j.1742-4658.2012.08511.x. PMID 22299668. S2CID 46705471.
- ^ Kurimchak, A; Graña, X (2015). "PP2A: more than a reset switch to activate pRB proteins during the cell cycle and in response to signaling cues". Cell Cycle. 14 (1): 18–30. doi:10.4161/15384101.2014.985069. PMC 4612414. PMID 25483052.
- ^ Naetar, N; Soundarapandian, V; Litovchick, L; Goguen, KL; Sablina, AA; Bowman-Colin, C; Sicinski, P; Hahn, WC; DeCaprio, JA; Livingston, DM (2014-06-19). "PP2A-mediated regulation of Ras signaling in G2 is essential for stable quiescence and normal G1 length". Molecular Cell. 54 (6): 932–45. doi:10.1016/j.molcel.2014.04.023. PMC 4118046. PMID 24857551.
- ^ Chen, JY; Lin, JR; Tsai, FC; Meyer, T (2013-10-10). "Dosage of Dyrk1a shifts cells within a p21-cyclin D1 signaling map to control the decision to enter the cell cycle". Molecular Cell. 52 (1): 87–100. doi:10.1016/j.molcel.2013.09.009. PMC 4039290. PMID 24119401.
- ^ Wolter, P; Schmitt, K; Fackler, M; Kremling, H; Probst, L; Hauser, S; Gruss, OJ; Gaubatz, S (15 May 2012). "GAS2L3, a target gene of the DREAM complex, is required for proper cytokinesis and genomic stability". Journal of Cell Science. 125 (Pt 10): 2393–406. doi:10.1242/jcs.097253. PMID 22344256.
- ^ DeCaprio, James A.; Duensing, Anette (July 2014). "The DREAM complex in antitumor activity of imatinib mesylate in gastrointestinal stromal tumors". Current Opinion in Oncology. 26 (4): 415–421. doi:10.1097/CCO.0000000000000090. PMC 4236229. PMID 24840522.
- ^ Boichuk, S.; Parry, J. A.; Makielski, K. R.; Litovchick, L.; Baron, J. L.; Zewe, J. P.; Wozniak, A.; Mehalek, K. R.; Korzeniewski, N.; Seneviratne, D. S.; Schoffski, P.; Debiec-Rychter, M.; DeCaprio, J. A.; Duensing, A. (20 June 2013). "The DREAM Complex Mediates GIST Cell Quiescence and Is a Novel Therapeutic Target to Enhance Imatinib-Induced Apoptosis". Cancer Research. 73 (16): 5120–5129. doi:10.1158/0008-5472.CAN-13-0579. PMID 23786773.
