Hemoprotein

A hemeprotein (or haemprotein; also hemoprotein or haemoprotein), or heme protein, is a protein that contains a heme prosthetic group.[1] They are a very large class of metalloproteins. The heme group confers functionality, which can include oxygen carrying, oxygen reduction, electron transfer, and other processes. Heme is bound to the protein either covalently or noncovalently or both.[2]
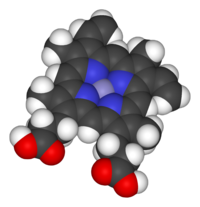
The heme consists of iron cation bound at the center of the conjugate base of the porphyrin, as well as other ligands attached to the "axial sites" of the iron. The porphyrin ring is a planar dianionic, tetradentate ligand. The iron is typically Fe2+ or Fe3+. One or two ligands are attached at the axial sites. The porphyrin ring has 4 nitrogen atoms that bind to the iron, leaving two other coordination positions of the iron available for bonding to the histidine of the protein and a divalent atom.[2]
Hemeproteins probably evolved to incorporate the iron atom contained within the protoporphyrin IX ring of heme into proteins. As it makes hemeproteins responsive to molecules that can bind divalent iron, this strategy has been maintained throughout evolution as it plays crucial physiological functions. The serum iron pool maintains iron in soluble form, making it more accessible for cells.[3] Oxygen (O2), nitric oxide (NO), carbon monoxide (CO) and hydrogen sulfide (H2S) bind to the iron atom in heme proteins. Once bound to the prosthetic heme groups, these molecules can modulate the activity/function of those hemeproteins, affording signal transduction. Therefore, when produced in biologic systems (cells), these gaseous molecules are referred to as gasotransmitters.
Because of their diverse biological functions and widespread abundance, hemeproteins are among the most studied biomolecules.[4] Data on heme protein structure and function has been aggregated into The Heme Protein Database (HPD), a secondary database to the Protein Data Bank.[5]
Roles[edit]
Hemeproteins have diverse biological functions including oxygen transport, which is completed via hemeproteins including hemoglobin, hemocyanin,[6] myoglobin, neuroglobin, cytoglobin, and leghemoglobin.[7]
Some hemeproteins—cytochrome P450s, cytochrome c oxidase, ligninases, catalase, and peroxidases—are enzymes. They often activate O2 for oxidation or hydroxylation.
Hemeproteins also enable electron transfer as they form part of the electron transport chain. Cytochrome a, cytochrome b, and cytochrome c have such electron transfer functions. It is now known that cytochrome a and cytochrome a3 make up one protein and was deemed the name cytochrome aa3.[8] The sensory system also relies on some hemeproteins including FixL, an oxygen sensor, CooA, a carbon monoxide sensor, and soluble guanylyl cyclase.
Hemoglobin and myoglobin[edit]
Hemoglobin and myoglobin are examples of hemeproteins that respectively transport and store of oxygen in mammals and in some fish.[9] Hemoglobin is a quaternary protein that occurs in the red blood cell, whereas, myoglobin is a tertiary protein found in the muscle cells of mammals. Although they might differ in location and size, their function are similar. Being hemeproteins, they both contain a heme prosthetic group.
His-F8 of the myoglobin, also known as the proximal histidine, is covalently bonded to the 5th coordination position of the iron. Oxygen interacts with the distal His by way of a hydrogen bond, not a covalent one. It binds to the 6th coordination position of the iron, His-E7 of the myoglobin binds to the oxygen that is now covalently bonded to the iron. The same is true for hemoglobin; however, being a protein with four subunits, hemoglobin contains four heme units in total, allowing four oxygen molecules in total to bind to the protein.
Myoglobin and hemoglobin are globular proteins that serve to bind and deliver oxygen using a prosthetic group. These globins dramatically improve the concentration of molecular oxygen that can be carried in the biological fluids of vertebrates and some invertebrates.
Differences occur in ligand binding and allosteric regulation.
Myoglobin[edit]
Myoglobin is found in vertebrate muscle cells and is a water-soluble globular protein.[10] Muscle cells, when put into action, can quickly require a large amount of oxygen for respiration due to their energy requirements. Therefore, muscle cells use myoglobin to accelerate oxygen diffusion and act as localized oxygen reserves for times of intense respiration. Myoglobin also stores the required amount of oxygen and makes it available for the muscle cell mitochondria.
Hemoglobin[edit]
In vertebrates, hemoglobin is found in the cytosol of red blood cells. Hemoglobin is sometimes referred to as the oxygen transport protein, in order to contrast it with myoglobin, which is stationary.
In vertebrates, oxygen is taken into the body by the tissues of the lungs, and passed to the red blood cells in the bloodstream where it's used in aerobic metabolic pathways.[10] Oxygen is then distributed to all of the tissues in the body and offloaded from the red blood cells to respiring cells. The hemoglobin then picks up carbon dioxide to be returned to the lungs. Thus, hemoglobin binds and off-loads both oxygen and carbon dioxide at the appropriate tissues, serving to deliver the oxygen needed for cellular metabolism and removing the resulting waste product, CO2.
Neuroglobin[edit]
Found in neurons, neuroglobin is responsible for driving nitric oxide to promote neuron cell survival[11] Neuroglobin is believed to increase the oxygen supply for neurons, sustaining ATP production, but they also function as storage proteins.[12]
Peroxidases and catalases[edit]
Almost all human peroxidases are hemoproteins, except glutathione peroxidase. They use hydrogen peroxide as a substrate. Metalloenzymes catalyze reactions using peroxide as an oxidant.[13] Catalases are hemoproteins responsible for the catalysis of converting hydrogen peroxide into water and oxygen.[14] They are made up of 4 subunits, each subunit having a Fe3+ heme group. They have an average molecular weight of ~240,000 g/mol.
Haloperoxidases involved in the innate immune system also contain a heme prosthetic group.
Electron transport chain and other redox catalysts[edit]
Cytochromes, cytochrome c oxidase, and coenzyme Q – cytochrome c reductase are heme-containing proteins or protein subunits embedded in the inner membrane of mitochondria which play an essential role in cellular respiration.
Sulfite oxidase, a molybdenum-dependent cytochrome, oxidizes sulfite to sulfate.
Nitric oxide synthase[edit]
Designed heme proteins[edit]

Due to the diverse functions of the heme molecule: as an electron transporter, an oxygen carrier, and as an enzyme cofactor, heme binding proteins have consistently attracted the attention of protein designers. Initial design attempts focused on α-helical heme binding proteins, in part, due to the relative simplicity of designing self-assembling helical bundles. Heme binding sites were designed inside the inter-helical hydrophobic grooves. Examples of such designs include:
- Helichrome[16][17]
- Globin-1[18]
- Cy-AA-EK[19]
- Peptides IIa/IId[20]
- α2[21]
- Transmembrane helical designs[22][23][24]
Later design attempts focused on creating functional heme binding helical bundles, such as:
- Oxidoreductases[25][26]
- Peroxidases[27][28]
- Electron transport proteins[29]
- Oxygen transport proteins[30]
- Photosensitive proteins[25]
Design techniques have matured to such an extent that it is now possible to generate entire libraries of heme binding helical proteins.[31]
Recent design attempts have focused on creating all-beta heme binding proteins, whose novel topology is very rare in nature. Such designs include:
Some methodologies attempt to incorporate cofactors into the hemoproteins who typically endure harsh conditions. In order to incorporate a synthetic cofactor, what must first occur is the denaturing of the holoprotein to remove the heme. The apoprotein is then rebuilt with the cofactor.[35]
References[edit]
- ^ "Heme Prosthetic Group Definition". earth.callutheran.edu. Retrieved 2023-04-27.
- ^ a b Nelson DL, Cox MN (2000). Lehninger, Principles of Biochemistry (3rd ed.). New Yorkm: Worth Publishing. ISBN 1-57259-153-6.
- ^ Frazer, David M.; Anderson, Gregory J. (March 2014). "The regulation of iron transport: The Regulation of Iron Transport". BioFactors. 40 (2): 206–214. doi:10.1002/biof.1148. PMID 24132807. S2CID 2998785.
- ^ Reedy CJ, Elvekrog MM, Gibney BR (January 2008). "Development of a heme protein structure-electrochemical function database". Nucleic Acids Research. 36 (Database issue): D307–D313. doi:10.1093/nar/gkm814. PMC 2238922. PMID 17933771.
- ^ Gibney BR. "Heme Protein Database". Brooklyn, NY: Brooklyn College.
- ^ "hemoproteins - Humpath.com - Human pathology". www.humpath.com. Retrieved 2023-04-27.
- ^ Lippard J, Berg JM (1994). Principles of Bioinorganic Chemistry. Mill Valley, CA: University Science Books. ISBN 0-935702-73-3.
- ^ Mahinthichaichan, Paween; Gennis, Robert B.; Tajkhorshid, Emad (2018-04-10). "Cytochrome aa 3 Oxygen Reductase Utilizes the Tunnel Observed in the Crystal Structures To Deliver O 2 for Catalysis". Biochemistry. 57 (14): 2150–2161. doi:10.1021/acs.biochem.7b01194. ISSN 0006-2960. PMC 5936630. PMID 29546752.
- ^ Gerber, Lucie; Clow, Kathy A.; Driedzic, William R.; Gamperl, Anthony K. (July 2021). "The Relationship between Myoglobin, Aerobic Capacity, Nitric Oxide Synthase Activity and Mitochondrial Function in Fish Hearts". Antioxidants. 10 (7): 1072. doi:10.3390/antiox10071072. ISSN 2076-3921. PMC 8301165. PMID 34356305.
- ^ a b "Hemoglobin and Myoglobin | Integrative Medical Biochemistry Examination and Board Review | AccessPharmacy | McGraw Hill Medical". accesspharmacy.mhmedical.com. Retrieved 2023-04-27.
- ^ DellaValle, Brian; Hempel, Casper; Kurtzhals, Jørgen A. L.; Penkowa, Milena (2010-08-01). "In vivo expression of neuroglobin in reactive astrocytes during neuropathology in murine models of traumatic brain injury, cerebral malaria, and autoimmune encephalitis: Neuroglobin in Reactive Astrogliosis". Glia. 58 (10): 1220–1227. doi:10.1002/glia.21002. PMID 20544857. S2CID 8563830.
- ^ Burmester, Thorsten; Hankeln, Thomas (June 2004). "Neuroglobin: A Respiratory Protein of the Nervous System". Physiology. 19 (3): 110–113. doi:10.1152/nips.01513.2003. ISSN 1548-9213. PMID 15143204.
- ^ Winterbourn, Christine C. (2013-01-01), "Chapter One - The Biological Chemistry of Hydrogen Peroxide", in Cadenas, Enrique; Packer, Lester (eds.), Hydrogen Peroxide and Cell Signaling, Part C, Methods in Enzymology, vol. 528, Academic Press, pp. 3–25, doi:10.1016/B978-0-12-405881-1.00001-X, ISBN 978-0-12-405881-1, PMID 23849856, retrieved 2023-04-27
- ^ Brzozowska, Ewa; Bazan, Justyna; Gamian, Andrzej (2011-03-25). "Funkcje białek bakteriofagowych". Postępy Higieny i Medycyny Doświadczalnej. 65: 167–176. doi:10.5604/17322693.936090. ISSN 1732-2693. PMID 21502693.
- ^ a b Nagarajan D, Sukumaran S, Deka G, Krishnamurthy K, Atreya HS, Chandra N (June 2018). "Design of a heme-binding peptide motif adopting a β-hairpin conformation". The Journal of Biological Chemistry. 293 (24): 9412–9422. doi:10.1074/jbc.RA118.001768. PMC 6005436. PMID 29695501.
- ^ Sasaki T, Kaiser ET (1989-01-01). "Helichrome: synthesis and enzymic activity of a designed hemeprotein". Journal of the American Chemical Society. 111 (1): 380–381. doi:10.1021/ja00183a065. ISSN 0002-7863.
- ^ Sasaki T, Kaiser ET (January 1990). "Synthesis and structural stability of helichrome as an artificial hemeproteins". Biopolymers. 29 (1): 79–88. doi:10.1002/bip.360290112. PMID 2328295. S2CID 35536899.
- ^ Isogai Y, Ota M, Fujisawa T, Izuno H, Mukai M, Nakamura H, et al. (June 1999). "Design and synthesis of a globin fold". Biochemistry. 38 (23): 7431–7443. doi:10.1021/bi983006y. PMID 10360940.
- ^ Rosenblatt MM, Wang J, Suslick KS (November 2003). "De novo designed cyclic-peptide heme complexes". Proceedings of the National Academy of Sciences of the United States of America. 100 (23): 13140–13145. Bibcode:2003PNAS..10013140R. doi:10.1073/pnas.2231273100. PMC 263730. PMID 14595023.
- ^ Robertson DE, Farid RS, Moser CC, Urbauer JL, Mulholland SE, Pidikiti R, et al. (March 1994). "Design and synthesis of multi-haem proteins". Nature. 368 (6470): 425–432. Bibcode:1994Natur.368..425R. doi:10.1038/368425a0. PMID 8133888. S2CID 4360174.
- ^ Choma CT, Lear JD, Nelson MJ, Dutton PL, Robertson DE, DeGrado WF (1994-02-01). "Design of a heme-binding four-helix bundle". Journal of the American Chemical Society. 116 (3): 856–865. doi:10.1021/ja00082a005. ISSN 0002-7863.
- ^ Discher BM, Noy D, Strzalka J, Ye S, Moser CC, Lear JD, et al. (September 2005). "Design of amphiphilic protein maquettes: controlling assembly, membrane insertion, and cofactor interactions". Biochemistry. 44 (37): 12329–12343. doi:10.1021/bi050695m. PMC 2574520. PMID 16156646.
- ^ Mahajan M, Bhattacharjya S (June 2014). "Designed di-heme binding helical transmembrane protein". ChemBioChem. 15 (9): 1257–1262. doi:10.1002/cbic.201402142. PMID 24829076. S2CID 20982919.
- ^ Korendovych IV, Senes A, Kim YH, Lear JD, Fry HC, Therien MJ, et al. (November 2010). "De novo design and molecular assembly of a transmembrane diporphyrin-binding protein complex". Journal of the American Chemical Society. 132 (44): 15516–15518. doi:10.1021/ja107487b. PMC 3016712. PMID 20945900.
- ^ a b Farid TA, Kodali G, Solomon LA, Lichtenstein BR, Sheehan MM, Fry BA, et al. (December 2013). "Elementary tetrahelical protein design for diverse oxidoreductase functions". Nature Chemical Biology. 9 (12): 826–833. doi:10.1038/nchembio.1362. PMC 4034760. PMID 24121554.
- ^ Huang SS, Koder RL, Lewis M, Wand AJ, Dutton PL (April 2004). "The HP-1 maquette: from an apoprotein structure to a structured hemoprotein designed to promote redox-coupled proton exchange". Proceedings of the National Academy of Sciences of the United States of America. 101 (15): 5536–5541. doi:10.1073/pnas.0306676101. PMC 397418. PMID 15056758.
- ^ Faiella M, Maglio O, Nastri F, Lombardi A, Lista L, Hagen WR, Pavone V (December 2012). "De novo design, synthesis and characterisation of MP3, a new catalytic four-helix bundle hemeprotein". Chemistry: A European Journal. 18 (50): 15960–15971. doi:10.1002/chem.201201404. PMID 23150230.
- ^ Cherry JR, Lamsa MH, Schneider P, Vind J, Svendsen A, Jones A, Pedersen AH (April 1999). "Directed evolution of a fungal peroxidase". Nature Biotechnology. 17 (4): 379–384. doi:10.1038/7939. PMID 10207888. S2CID 41233353.
- ^ Anderson JL, Armstrong CT, Kodali G, Lichtenstein BR, Watkins DW, Mancini JA, et al. (February 2014). "Constructing a man-made c-type cytochrome maquette in vivo: electron transfer, oxygen transport and conversion to a photoactive light harvesting maquette". Chemical Science. 5 (2): 507–514. doi:10.1039/C3SC52019F. PMC 3952003. PMID 24634717.
- ^ Koder RL, Anderson JL, Solomon LA, Reddy KS, Moser CC, Dutton PL (March 2009). "Design and engineering of an O(2) transport protein". Nature. 458 (7236): 305–309. Bibcode:2009Natur.458..305K. doi:10.1038/nature07841. PMC 3539743. PMID 19295603.
- ^ Moffet DA, Foley J, Hecht MH (September 2003). "Midpoint reduction potentials and heme binding stoichiometries of de novo proteins from designed combinatorial libraries". Biophysical Chemistry. Walter Kauzmann's 85th Birthday. 105 (2–3): 231–239. doi:10.1016/S0301-4622(03)00072-3. PMID 14499895.
- ^ Mahajan M, Bhattacharjya S (June 2013). "β-Hairpin peptides: heme binding, catalysis, and structure in detergent micelles". Angewandte Chemie. 52 (25): 6430–6434. doi:10.1002/anie.201300241. PMID 23640811.
- ^ D'Souza A, Wu X, Yeow EK, Bhattacharjya S (May 2017). "Designed Heme-Cage β-Sheet Miniproteins". Angewandte Chemie. 56 (21): 5904–5908. doi:10.1002/anie.201702472. PMID 28440962.
- ^ D'Souza A, Mahajan M, Bhattacharjya S (April 2016). "Designed multi-stranded heme binding β-sheet peptides in membrane". Chemical Science. 7 (4): 2563–2571. doi:10.1039/C5SC04108B. PMC 5477022. PMID 28660027.
- ^ Lemon, Christopher M.; Marletta, Michael A. (2021-12-21). "Designer Heme Proteins: Achieving Novel Function with Abiological Heme Analogues". Accounts of Chemical Research. 54 (24): 4565–4575. doi:10.1021/acs.accounts.1c00588. ISSN 0001-4842. PMC 8754152. PMID 34890183.
External links[edit]
- Heme Protein Database
- Hemeproteins at the U.S. National Library of Medicine Medical Subject Headings (MeSH)
