Heparan sulfate

Heparan sulfate (HS) is a linear polysaccharide found in all animal tissues.[1] It occurs as a proteoglycan (HSPG, i.e. Heparan Sulfate ProteoGlycan) in which two or three HS chains are attached in close proximity to cell surface or extracellular matrix proteins.[2][3] In this form, HS binds to a variety of protein ligands, including Wnt,[4][5] and regulates a wide range of biological activities, including developmental processes, angiogenesis, blood coagulation, abolishing detachment activity by GrB (Granzyme B),[6] and tumour metastasis. HS has also been shown to serve as cellular receptor for a number of viruses, including the respiratory syncytial virus.[7] One study suggests that cellular heparan sulfate has a role in SARS-CoV-2 Infection, particularly when the virus attaches with ACE2.[8]
Proteoglycans[edit]
The major cell membrane HSPGs are the transmembrane syndecans and the glycosylphosphatidylinositol (GPI) anchored glypicans.[9][10] Other minor forms of membrane HSPG include betaglycan[11] and the V-3 isoform of CD44 present on keratinocytes and activated monocytes.[12]
In the extracellular matrix, especially basement membranes and fractones,[13] the multi-domain perlecan,[14] agrin[15] and collagen XVIII[16] core proteins are the main HS-bearing species.
Structure and differences from heparin[edit]
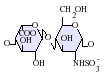
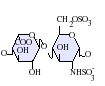
Heparan sulfate is a member of the glycosaminoglycan (GAG) family of carbohydrates and is very closely related in structure to heparin. Both consist of a variably sulfated repeating disaccharide unit. The main disaccharide units that occur in heparan sulfate and heparin are shown below.
The most common disaccharide unit within heparan sulfate is composed of a glucuronic acid (GlcA) linked to N-acetylglucosamine (GlcNAc), typically making up around 50% of the total disaccharide units. Compare this to heparin, where IdoA(2S)-GlcNS(6S) makes up 85% of heparins from beef lung and about 75% of those from porcine intestinal mucosa. Problems arise when defining hybrid GAGs that contain both 'heparin-like' and 'HS-like' structures. It has been suggested that a GAG should qualify as heparin only if its content of N-sulfate groups largely exceeds that of N-acetyl groups and the concentration of O-sulfate groups exceeds those of N-sulfate. Otherwise, it should be classified as HS.[17]
Not shown below are the rare disaccharides containing a 3-O-sulfated glucosamine (GlcNS(3S,6S) or a free amine group (GlcNH3+). Under physiological conditions the ester and amide sulfate groups are deprotonated and attract positively charged counterions to form a salt.[18] It is in this form that HS is thought to exist at the cell surface.
-
GlcA-GlcNAc
-
GlcA-GlcNS
-
IdoA-GlcNS
-
IdoA(2S)-GlcNS
-
IdoA-GlcNS(6S)
-
IdoA(2S)-GlcNS(6S)
Abbreviations[edit]
- GAG = glycosaminoglycan
- GlcA = β-D-glucuronic acid
- IdoA = α-L-iduronic acid
- IdoA(2S) = 2-O-sulfo-α-L-iduronic acid
- GlcNAc = 2-deoxy-2-acetamido-α-D-glucopyranosyl
- GlcNS = 2-deoxy-2-sulfamido-α-D-glucopyranosyl
- GlcNS(6S) = 2-deoxy-2-sulfamido-α-D-glucopyranosyl-6-O-sulfate
Biosynthesis[edit]
Many different cell types produce HS chains with many different primary structures. Therefore, there is a great deal of variability in the way HS chains are synthesised, producing structural diversity encompassed by the term "heparanome" - which defines the full range of primary structures produced by a particular cell, tissue or organism.[19] However, essential to the formation of HS regardless of primary sequence is a range of biosynthetic enzymes. These enzymes consist of multiple glycosyltransferases, sulfotransferases and an epimerase. These same enzymes also synthesize heparin.
In the 1980s, Jeffrey Esko was the first to isolate and characterize animal cell mutants altered in the assembly of heparan sulfate.[20] Many of these enzymes have now been purified, molecularly cloned and their expression patterns studied. From this and early work on the fundamental stages of HS/heparin biosynthesis using a mouse mastocytoma cell free system a lot is known about the order of enzyme reactions and specificity.[21]
Chain initiation[edit]

HS synthesis initiates with the transfer of xylose from UDP-xylose by xylosyltransferase (XT) to specific serine residues within the protein core. Attachment of two galactose (Gal) residues by galactosyltransferases I and II (GalTI and GalTII) and glucuronic acid (GlcA) by glucuronosyltransferase I (GlcATI) completes the formation of a tetrasaccharide primer O-linked to a serine of the core-protein:[22]
βGlcUA-(1→3)-βGal-(1→3)-βGal-(1→4)-βXyl-O-Ser.
The pathways for HS/heparin or chondroitin sulfate (CS) and dermatan sulfate (DS) biosynthesis diverge after the formation of this common tetrasaccharide linkage structure. The next enzyme to act, GlcNAcT-I or GalNAcT-I, directs synthesis, either to HS/heparin or CS/DS, respectively.[23]
Xylose attachment to the core protein is thought to occur in the endoplasmic reticulum (ER) with further assembly of the linkage region and the remainder of the chain occurring in the Golgi apparatus.[22][23]
Chain elongation[edit]
After attachment of the first N-acetylglucosamine (GlcNAc) residue, elongation of the tetrasacchride linker is continued by the stepwise addition of GlcA and GlcNAc residues. These are transferred from their respective UDP-sugar nucleotides. This is carried out by EXT family proteins with glycosyltransferase activities. EXT family genes are tumor suppressors. [22][24]
Mutations at the EXT1-3 gene loci in humans lead to an inability of cells to produce HS and to the development of the disease Multiple Hereditary Exostoses (MHE). MHE is characterized by cartilage-capped tumours, known as osteochondromas or exostoses, which develop primarily on the long bones of affected individuals from early childhood until puberty.[25]
Chain modification[edit]
As an HS chain polymerises, it undergoes a series of modification reactions carried out by four classes of sulfotransferases and an epimerase. The availability of the sulfate donor PAPS is crucial to the activity of the sulfotransferases.[26][27]
N-deacetylation/N-sulfation[edit]
The first polymer modification is the N-deacetylation/N-sulfation of GlcNAc residues into GlcNS. This is a prerequisite for all subsequent modification reactions, and is carried out by one or more members of a family of four GlcNAc N-deacetylase/N-sulfotransferase enzymes (NDSTs). In early studies, it was shown that modifying enzymes could recognize and act on any N-acetylated residue in the forming polymer.[28] Therefore, the modification of GlcNAc residues should occur randomly throughout the chain. However, in HS, N-sulfated residues are mainly grouped together and separated by regions of N-acetylation where GlcNAc remains unmodified.
There are four isoforms of NDST (NDST1–4). Both N-deacetylase and N-sulfotransferase activities are present in all NDST-isoforms but they differ significantly in their enzymatic activities.[29]
Generation of GlcNH2[edit]
Due to the N-deacetylase and N-sulfotransferase being carried out by the same enzyme N-sulfation is normally tightly coupled to N-acetylation. GlcNH2 residues resulting from apparent uncoupling of the two activities have been found in heparin and some species of HS.[30]
Epimerisation and 2-O-sulfation[edit]
Epimerisation is catalysed by one enzyme, the GlcA C5 epimerase or heparosan-N-sulfate-glucuronate 5-epimerase (EC 5.1.3.17). This enzyme epimerises GlcA to iduronic acid (IdoA). Substrate recognition requires that the GlcN residue linked to the non-reducing side of a potential GlcA target be N-sulfated. Uronosyl-2-O-sulfotransferase (2OST) sulfates the resulting IdoA residues.
6-O-sulfation[edit]
Three glucosaminyl 6-O-transferases (6OSTs) have been identified that result in the formation of GlcNS(6S) adjacent to sulfated or non-sulfated IdoA. GlcNAc(6S) is also found in mature HS chains.
3-O-sulfation[edit]
Currently seven glucosaminyl 3-O-sulfotransferases (3OSTs, HS3STs) are known to exist in mammals (eight in zebrafish).[31][32] The 3OST enzymes create a number of possible 3-O-sulfated disaccharides, including GlcA-GlcNS(3S±6S) (modified by HS3ST1 and HS3ST5), IdoA(2S)-GlcNH2(3S±6S)(modified by HS3ST3A1, HS3ST3B1, HS3ST5 and HS3ST6) and GlcA/IdoA(2S)-GlcNS(3S) (modified by HS3ST2 and HS3ST4).[33][34][35][36] As with all other HS sulfotransferases, the 3OSTs use 3'-phosphoadenosine-5'-phosphosulfate (PAPS) as a sulfate donor. Despite being the largest family of HS modification enzymes, the 3OSTs produce the rarest HS modification, the 3-O-sulfation of specific glucosamine residues at the C3-OH moiety.[37]
The 3OSTs are divided into two functional subcategories, those that generate an antithrombin III binding site (HS3ST1 and HS3ST5) and those that generate a herpes simplex virus 1 glycoprotein D (HSV-1 gD) binding site (HS3ST2, HS3ST3A1, HS3ST3B1, HS3ST4, HS3ST5 and HS3ST6).[33][34][35][36][38][39][40][41][42][43][44] As the 3OSTs are the largest family of HS modification enzymes and their actions are rate-limiting, substrate specific and produce rare modifications, it has been hypothesized that 3OST modified HS plays an important regulatory role in biological processes.[36][39] It has been demonstrated that 3-O-sulfation can enhance the binding of Wnt to the glypican and may play a role in regulating Wnt in cancer.[5][10]
Ligand binding[edit]
Heparan sulfate binds with a large number of extracellular proteins. These are often collectively called the “heparin interactome” or "heparin-binding proteins", because they are isolated by affinity chromatography on the related polysaccharide heparin, though the term “heparan sulfate interactome” is more correct. The functions of heparan sulfate binding proteins ranges from extracellular matrix components, to enzymes and coagulation factors, and most growth factors, cytokines, chemokines and morphogens [45] The laboratory of Mitchell Ho at the NCI isolated the HS20 human monoclonal antibody with high affinity for heparan sulfate by phage display.[46] The antibody binds heparan sulfate, not chondroitin sulfate.[5] The binding of HS20 to heparan sulfate requires sulfation at both the C2 position and C6 position. HS20 blocks the Wnt binding on heparan sulfate[5] and also inhibits infectious entry of pathogenic JC polyomavirus.[47]
Interferon-γ[edit]
The cell surface receptor binding region of Interferon-γ overlaps with the HS binding region, near the protein's C-terminal. Binding of HS blocks the receptor binding site and as a result, protein-HS complexes are inactive.[48]
Wnt[edit]
Glypican-3 (GPC3) interacts with both Wnt and Frizzled to form a complex and triggers downstream signaling.[4][10] It has been experimentally established that Wnt recognizes a heparan sulfate motif on GPC3, which contains IdoA2S and GlcNS6S, and that the 3-O-sulfation in GlcNS6S3S enhances the binding of Wnt to the glypican.[5]
The HS-binding properties of a number of other proteins are also being studied:
- Antithrombin III
- Fibroblast growth factor
- Hepatocyte growth factor
- Interleukin-8
- Vascular endothelial growth factor
- Wnt/Wingless
- Endostatin
Heparan sulfate analogue[edit]
Heparan sulfate analogues are thought to display identical properties as heparan sulfate with exception of being stable in a proteolytic environment like a wound.[49][50] Because heparan sulfate is broken down in chronic wounds by heparanase, the analogues only bind sites where natural heparan sulfate is absent and is thus resistant to enzyme degration. [51] Also the function of the heparan sulfate analogues is the same as heparan sulfate, protecting a variety of protein ligands such as growth factors and cytokines. By holding them in place, the tissue can then use the different protein ligands for proliferation.
Associated conditions[edit]
Hereditary multiple exostoses (also known as multiple hereditary exostoses or multiple osteochondromas is a hereditary disease with mutations on the EXT1 and EXT2 genes that affect biosynthesis of heparan sulfate.[52] [53]
References[edit]
- ^ Medeiros GF, Mendes A, Castro RA, Baú EC, Nader HB, Dietrich CP (July 2000). "Distribution of sulfated glycosaminoglycans in the animal kingdom: widespread occurrence of heparin-like compounds in invertebrates". Biochimica et Biophysica Acta (BBA) - General Subjects. 1475 (3): 287–94. doi:10.1016/S0304-4165(00)00079-9. PMID 10913828.
- ^ Gallagher JT, Lyon M (2000). "Molecular structure of Heparan Sulfate and interactions with growth factors and morphogens". In Iozzo MV (ed.). Proteoglycans: structure, biology and molecular interactions. New York, New York: Marcel Dekker Inc. pp. 27–59.
- ^ Iozzo RV (1998). "Matrix proteoglycans: from molecular design to cellular function". Annual Review of Biochemistry. 67: 609–52. doi:10.1146/annurev.biochem.67.1.609. PMID 9759499. S2CID 14638091.
- ^ a b Gao W, Kim H, Feng M, Phung Y, Xavier CP, Rubin JS, Ho M (August 2014). "Inactivation of Wnt signaling by a human antibody that recognizes the heparan sulfate chains of glypican-3 for liver cancer therapy". Hepatology. 60 (2): 576–87. doi:10.1002/hep.26996. PMC 4083010. PMID 24492943.
- ^ a b c d e Gao W, Xu Y, Liu J, Ho M (May 2016). "Epitope mapping by a Wnt-blocking antibody: evidence of the Wnt binding domain in heparan sulfate". Scientific Reports. 6: 26245. Bibcode:2016NatSR...626245G. doi:10.1038/srep26245. PMC 4869111. PMID 27185050.
- ^ Buzza MS, Zamurs L, Sun J, Bird CH, Smith AI, Trapani JA, et al. (June 2005). "Extracellular matrix remodeling by human granzyme B via cleavage of vitronectin, fibronectin, and laminin". The Journal of Biological Chemistry. 280 (25): 23549–58. doi:10.1074/jbc.M412001200. PMID 15843372.
- ^ Hallak LK, Spillmann D, Collins PL, Peeples ME (November 2000). "Glycosaminoglycan sulfation requirements for respiratory syncytial virus infection". Journal of Virology. 74 (22): 10508–13. doi:10.1128/JVI.74.22.10508-10513.2000. PMC 110925. PMID 11044095.
- ^ Clausen TM, Sandoval DR, Spliid CB, Pihl J, Perrett HR, Painter CD, et al. (14 September 2020). "SARS-CoV-2 Infection Depends on Cellular Heparan Sulfate and ACE2". The Journal of Cell. 183 (4): 1043–1057.e15. doi:10.1016/j.cell.2020.09.033. PMC 7489987. PMID 32970989.
- ^ Ho M, Kim H (February 2011). "Glypican-3: a new target for cancer immunotherapy". European Journal of Cancer. 47 (3): 333–8. doi:10.1016/j.ejca.2010.10.024. PMC 3031711. PMID 21112773.
- ^ a b c Li N, Gao W, Zhang YF, Ho M (November 2018). "Glypicans as Cancer Therapeutic Targets". Trends in Cancer. 4 (11): 741–754. doi:10.1016/j.trecan.2018.09.004. PMC 6209326. PMID 30352677.
- ^ Andres JL, DeFalcis D, Noda M, Massagué J (March 1992). "Binding of two growth factor families to separate domains of the proteoglycan betaglycan". The Journal of Biological Chemistry. 267 (9): 5927–30. doi:10.1016/S0021-9258(18)42643-9. PMID 1556106.
- ^ Jackson DG, Bell JI, Dickinson R, Timans J, Shields J, Whittle N (February 1995). "Proteoglycan forms of the lymphocyte homing receptor CD44 are alternatively spliced variants containing the v3 exon". The Journal of Cell Biology. 128 (4): 673–85. doi:10.1083/jcb.128.4.673. PMC 2199896. PMID 7532175.
- ^ Mercier, Frederic (2016). "Fractones: extracellular matrix niche controlling stem cell fate and growth factor activity in the brain in health and disease". Cellular and Molecular Life Sciences. 73 (24): 4661–4674. doi:10.1007/s00018-016-2314-y. PMID 27475964. S2CID 28119663.
- ^ Arikawa-Hirasawa E, Wilcox WR, Le AH, Silverman N, Govindraj P, Hassell JR, Yamada Y (April 2001). "Dyssegmental dysplasia, Silverman-Handmaker type, is caused by functional null mutations of the perlecan gene". Nature Genetics. 27 (4): 431–4. doi:10.1038/86941. PMID 11279527. S2CID 22934192.
- ^ Verbeek, Marcel M.; Otte-Höller, Irene; van den Born, Jacob; van den Heuvel, Lambert P.W.J.; David, Guido; Wesseling, Pieter; de Waal, Robert M.W. (1999). "Agrin Is a Major Heparan Sulfate Proteoglycan Accumulating in Alzheimer's Disease Brain". The American Journal of Pathology. 155 (6). Elsevier BV: 2115–2125. doi:10.1016/s0002-9440(10)65529-0. ISSN 0002-9440. PMC 1866925. PMID 10595940.
- ^ Kawashima, Hiroto; Watanabe, Norifumi; Hirose, Mayumi; Sun, Xin; Atarashi, Kazuyuki; Kimura, Tetsuya; Shikata, Kenichi; Matsuda, Mitsuhiro; Ogawa, Daisuke; Heljasvaara, Ritva; Rehn, Marko; Pihlajaniemi, Taina; Miyasaka, Masayuki (2003). "Collagen XVIII, a Basement Membrane Heparan Sulfate Proteoglycan, Interacts with L-selectin and Monocyte Chemoattractant Protein-1". Journal of Biological Chemistry. 278 (15). Elsevier BV: 13069–13076. doi:10.1074/jbc.m212244200. ISSN 0021-9258. PMID 12556525.
- ^ Gallagher JT, Walker A (September 1985). "Molecular distinctions between heparan sulphate and heparin. Analysis of sulphation patterns indicates that heparan sulphate and heparin are separate families of N-sulphated polysaccharides". The Biochemical Journal. 230 (3): 665–74. doi:10.1042/bj2300665. PMC 1152670. PMID 2933029.
- ^ LA, Fransson; I, Silverberg; I, Carlstedt (1985). "Structure of the heparan sulfate-protein linkage region. Demonstration of the sequence galactosyl-galactosyl-xylose-2-phosphate". The Journal of Biological Chemistry. 260 (27). J Biol Chem: 14722–14726. doi:10.1016/S0021-9258(17)38632-5. ISSN 0021-9258. PMID 2932448.
- ^ Turnbull J, Powell A, Guimond S (February 2001). "Heparan sulfate: decoding a dynamic multifunctional cell regulator". Trends in Cell Biology. 11 (2): 75–82. doi:10.1016/s0962-8924(00)01897-3. PMID 11166215.
- ^ Esko JD, Stewart TE, Taylor WH (May 1985). "Animal cell mutants defective in glycosaminoglycan biosynthesis". Proceedings of the National Academy of Sciences of the United States of America. 82 (10): 3197–201. Bibcode:1985PNAS...82.3197E. doi:10.1073/pnas.82.10.3197. PMC 397742. PMID 3858816.
- ^ Lindahl U, Kusche-Gullberg M, Kjellén L (September 1998). "Regulated diversity of heparan sulfate". The Journal of Biological Chemistry. 273 (39): 24979–82. doi:10.1074/jbc.273.39.24979. PMID 9737951.
- ^ a b c Kreuger, Johan; Kjellén, Lena (2012-10-04). "Heparan Sulfate Biosynthesis". Journal of Histochemistry & Cytochemistry. 60 (12). SAGE Publications: 898–907. doi:10.1369/0022155412464972. ISSN 0022-1554. PMC 3527889. PMID 23042481.
- ^ a b Jones, Courtney L.; Liu, Jian; Xu, Ding (2010). "Structure, Biosynthesis, and Function of Glycosaminoglycans". Comprehensive Natural Products II. Elsevier. pp. 407–427. doi:10.1016/b978-008045382-8.00132-5. ISBN 9780080453828.
- ^ Busse-Wicher, Marta; Wicher, Krzysztof B.; Kusche-Gullberg, Marion (2014). "The extostosin family: Proteins with many functions". Matrix Biology. 35. Elsevier BV: 25–33. doi:10.1016/j.matbio.2013.10.001. hdl:1956/10590. ISSN 0945-053X.
- ^ Beltrami G, Ristori G, Scoccianti G, Tamburini A, Capanna R (2016). "Hereditary Multiple Exostoses: a review of clinical appearance and metabolic pattern". Clinical Cases in Mineral and Bone Metabolism. 13 (2): 110–118. doi:10.11138/ccmbm/2016.13.2.110. PMC 5119707. PMID 27920806.
- ^ Silbert JE (November 1967). "Biosynthesis of heparin. 3. Formation of a sulfated glycosaminoglycan with a microsomal preparation from mast cell tumors". The Journal of Biological Chemistry. 242 (21): 5146–52. doi:10.1016/S0021-9258(18)99487-1. PMID 4228675.
- ^ Carlsson P, Presto J, Spillmann D, Lindahl U, Kjellén L (July 2008). "Heparin/heparan sulfate biosynthesis: processive formation of N-sulfated domains". The Journal of Biological Chemistry. 283 (29): 20008–14. doi:10.1074/jbc.M801652200. PMID 18487608.
- ^ Höök M, Lindahl U, Hallén A, Bäckström G (August 1975). "Biosynthesis of heparin. Studies on the microsomal sulfation process". The Journal of Biological Chemistry. 250 (15): 6065–71. doi:10.1016/S0021-9258(19)41159-9. PMID 807579.
- ^ Aikawa J, Grobe K, Tsujimoto M, Esko JD (February 2001). "Multiple isozymes of heparan sulfate/heparin GlcNAc N-deacetylase/GlcN N-sulfotransferase. Structure and activity of the fourth member, NDST4". The Journal of Biological Chemistry. 276 (8): 5876–82. doi:10.1074/jbc.M009606200. PMID 11087757.
- ^ Toida T, Yoshida H, Toyoda H, Koshiishi I, Imanari T, Hileman RE, et al. (March 1997). "Structural differences and the presence of unsubstituted amino groups in heparan sulphates from different tissues and species". The Biochemical Journal. 322 ( Pt 2) (Pt 2): 499–506. doi:10.1042/bj3220499. PMC 1218218. PMID 9065769.
- ^ Cadwallader AB, Yost HJ (February 2007). "Combinatorial expression patterns of heparan sulfate sulfotransferases in zebrafish: III. 2-O-sulfotransferase and C5-epimerases". Developmental Dynamics. 236 (2): 581–6. doi:10.1002/dvdy.21051. PMID 17195182. S2CID 38249813.
- ^ Xu D, Tiwari V, Xia G, Clement C, Shukla D, Liu J (January 2005). "Characterization of heparan sulphate 3-O-sulphotransferase isoform 6 and its role in assisting the entry of herpes simplex virus type 1". The Biochemical Journal. 385 (Pt 2): 451–9. doi:10.1042/BJ20040908. PMC 1134716. PMID 15303968.
- ^ a b Shukla D, Liu J, Blaiklock P, Shworak NW, Bai X, Esko JD, et al. (October 1999). "A novel role for 3-O-sulfated heparan sulfate in herpes simplex virus 1 entry". Cell. 99 (1): 13–22. doi:10.1016/s0092-8674(00)80058-6. PMID 10520990. S2CID 14139940.
- ^ a b Xia G, Chen J, Tiwari V, Ju W, Li JP, Malmstrom A, et al. (October 2002). "Heparan sulfate 3-O-sulfotransferase isoform 5 generates both an antithrombin-binding site and an entry receptor for herpes simplex virus, type 1". The Journal of Biological Chemistry. 277 (40): 37912–9. doi:10.1074/jbc.m204209200. PMID 12138164.
- ^ a b Xu D, Tiwari V, Xia G, Clement C, Shukla D, Liu J (January 2005). "Characterization of heparan sulphate 3-O-sulphotransferase isoform 6 and its role in assisting the entry of herpes simplex virus type 1". The Biochemical Journal. 385 (Pt 2): 451–9. doi:10.1042/bj20040908. PMC 1134716. PMID 15303968.
- ^ a b c Lawrence R, Yabe T, Hajmohammadi S, Rhodes J, McNeely M, Liu J, et al. (July 2007). "The principal neuronal gD-type 3-O-sulfotransferases and their products in central and peripheral nervous system tissues". Matrix Biology. 26 (6): 442–55. doi:10.1016/j.matbio.2007.03.002. PMC 1993827. PMID 17482450.
- ^ Shworak NW, HajMohammadi S, de Agostini AI, Rosenberg RD (2003). "Mice deficient in heparan sulfate 3-O-sulfotransferase-1: normal hemostasis with unexpected perinatal phenotypes". Glycoconjugate Journal. 19 (4–5): 355–61. doi:10.1023/a:1025377206600. PMID 12975616. S2CID 21853086.
- ^ Liu J, Shworak NW, Fritze LM, Edelberg JM, Rosenberg RD (October 1996). "Purification of heparan sulfate D-glucosaminyl 3-O-sulfotransferase". The Journal of Biological Chemistry. 271 (43): 27072–82. doi:10.1074/jbc.271.43.27072. PMID 8900198.
- ^ a b Shworak NW, Liu J, Fritze LM, Schwartz JJ, Zhang L, Logeart D, Rosenberg RD (October 1997). "Molecular cloning and expression of mouse and human cDNAs encoding heparan sulfate D-glucosaminyl 3-O-sulfotransferase". The Journal of Biological Chemistry. 272 (44): 28008–19. doi:10.1074/jbc.272.44.28008. PMID 9346953.
- ^ Shworak NW, Liu J, Petros LM, Zhang L, Kobayashi M, Copeland NG, et al. (February 1999). "Multiple isoforms of heparan sulfate D-glucosaminyl 3-O-sulfotransferase. Isolation, characterization, and expression of human cdnas and identification of distinct genomic loci". The Journal of Biological Chemistry. 274 (8): 5170–84. doi:10.1074/jbc.274.8.5170. PMID 9988767.
- ^ Chen J, Duncan MB, Carrick K, Pope RM, Liu J (November 2003). "Biosynthesis of 3-O-sulfated heparan sulfate: unique substrate specificity of heparan sulfate 3-O-sulfotransferase isoform 5". Glycobiology. 13 (11): 785–94. doi:10.1093/glycob/cwg101. PMID 12907690.
- ^ Duncan MB, Chen J, Krise JP, Liu J (March 2004). "The biosynthesis of anticoagulant heparan sulfate by the heparan sulfate 3-O-sulfotransferase isoform 5". Biochimica et Biophysica Acta (BBA) - General Subjects. 1671 (1–3): 34–43. doi:10.1016/j.bbagen.2003.12.010. PMID 15026143.
- ^ Chen J, Liu J (September 2005). "Characterization of the structure of antithrombin-binding heparan sulfate generated by heparan sulfate 3-O-sulfotransferase 5". Biochimica et Biophysica Acta (BBA) - General Subjects. 1725 (2): 190–200. doi:10.1016/j.bbagen.2005.06.012. PMID 16099108.
- ^ Girardin EP, Hajmohammadi S, Birmele B, Helisch A, Shworak NW, de Agostini AI (November 2005). "Synthesis of anticoagulantly active heparan sulfate proteoglycans by glomerular epithelial cells involves multiple 3-O-sulfotransferase isoforms and a limiting precursor pool". The Journal of Biological Chemistry. 280 (45): 38059–70. doi:10.1074/jbc.m507997200. PMID 16107334.
- ^ Ori A, Wilkinson MC, Fernig DG (May 2008). "The heparanome and regulation of cell function: structures, functions and challenges". Frontiers in Bioscience. 13 (13): 4309–38. doi:10.2741/3007. PMID 18508513.
- ^ Kim H, Ho M (November 2018). "Isolation of Antibodies to Heparan Sulfate on Glypicans by Phage Display". Current Protocols in Protein Science. 94 (1): e66. doi:10.1002/cpps.66. PMC 6205898. PMID 30091851.
- ^ Geoghegan EM, Pastrana DV, Schowalter RM, Ray U, Gao W, Ho M, et al. (October 2017). "Infectious Entry and Neutralization of Pathogenic JC Polyomaviruses". Cell Reports. 21 (5): 1169–1179. doi:10.1016/j.celrep.2017.10.027. PMC 5687836. PMID 29091757.
- ^ Sadir R, Forest E, Lortat-Jacob H (May 1998). "The heparan sulfate binding sequence of interferon-gamma increased the on rate of the interferon-gamma-interferon-gamma receptor complex formation". The Journal of Biological Chemistry. 273 (18): 10919–25. doi:10.1074/jbc.273.18.10919. PMID 9556569.
- ^ Tong M, Tuk B, Hekking IM, Vermeij M, Barritault D, van Neck JW (2009). "Stimulated neovascularization, inflammation resolution and collagen maturation in healing rat cutaneous wounds by a heparan sulfate glycosaminoglycan mimetic, OTR4120". Wound Repair and Regeneration. 17 (6): 840–52. doi:10.1111/j.1524-475X.2009.00548.x. PMID 19903305. S2CID 17262546.
- ^ Tong M, Tuk B, Hekking IM, Pleumeekers MM, Boldewijn MB, Hovius SE, van Neck JW (2011). "Heparan sulfate glycosaminoglycan mimetic improves pressure ulcer healing in a rat model of cutaneous ischemia-reperfusion injury". Wound Repair and Regeneration. 19 (4): 505–14. doi:10.1111/j.1524-475X.2011.00704.x. PMID 21649786. S2CID 7380997.
- ^ Tong, Miao; Tuk, Bastiaan; Hekking, Ineke M.; Vermeij, Marcel; Barritault, Denis; van Neck, Johan W. (2009). "Stimulated neovascularization, inflammation resolution and collagen maturation in healing rat cutaneous wounds by a heparan sulfate glycosaminoglycan mimetic, OTR4120". Wound Repair and Regeneration. 17 (6). Wiley: 840–852. doi:10.1111/j.1524-475x.2009.00548.x. ISSN 1067-1927. PMID 19903305. S2CID 17262546.
- ^ Zak, Beverly M; Crawford, Brett E; Esko, Jeffrey D (2002-12-19). "Hereditary multiple exostoses and heparan sulfate polymerization". Biochimica et Biophysica Acta (BBA) - General Subjects. Developmental Glycobiology. 1573 (3): 346–355. doi:10.1016/S0304-4165(02)00402-6. ISSN 0304-4165.
- ^ Meneghetti, Maria C. Z.; Hughes, Ashley J.; Rudd, Timothy R.; Nader, Helena B.; Powell, Andrew K.; Yates, Edwin A.; Lima, Marcelo A. (2015-09-06). "Heparan sulfate and heparin interactions with proteins". Journal of the Royal Society, Interface. 12 (110): 0589. doi:10.1098/rsif.2015.0589. ISSN 1742-5662. PMC 4614469. PMID 26289657.