Phycoerythrin
| Phycoerythrin, alpha/beta chain | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Identifiers | |||||||||
| Symbol | Phycoerythr_ab | ||||||||
| Pfam | PF02972 | ||||||||
| InterPro | IPR004228 | ||||||||
| SCOP2 | 1qgw / SCOPe / SUPFAM | ||||||||
| |||||||||
Phycoerythrin (PE) is a red protein-pigment complex from the light-harvesting phycobiliprotein family, present in cyanobacteria,[1] red algae[2] and cryptophytes,[3] accessory to the main chlorophyll pigments responsible for photosynthesis.The red pigment is due to the prosthetic group, phycoerythrobilin, which gives phycoerythrin its red color.[4]
Like all phycobiliproteins, it is composed of a protein part covalently binding chromophores called phycobilins. In the phycoerythrin family, the most known phycobilins are: phycoerythrobilin, the typical phycoerythrin acceptor chromophore. Phycoerythrobilin is a linear tetrapyrrole molecule found in cyanobacteria, red algae, and cryptomonads. Together with other bilins such as phycocyanobilin it serves as a light-harvesting pigment in the photosynthetic light-harvesting structures of cyanobacteria called phycobilisomes.[5] Phycoerythrins are composed of (αβ) monomers, usually organised in a disk-shaped trimer (αβ)3 or hexamer (αβ)6 (second one is the functional unit of the antenna rods). These typical complexes also contain a third type of subunit, the γ chain.[2]
Phycobilisomes[edit]

Phycobiliproteins are part of huge light harvesting antennae protein complexes called phycobilisomes. In red algae they are anchored to the stromal side of thylakoid membranes of chloroplasts, whereas in cryptophytes phycobilisomes are reduced and (phycobiliprotein 545 PE545 molecules here) are densely packed inside the lumen of thylakoides. [3][7]
Phycobiliproteins have many practical application to them including imperative properties like hepato-protective, anti-oxidants, anti-inflammatory and anti-aging activity of PBPs enable their use in food, cosmetics, pharmaceutical and biomedical industries. PBPs have been also noted to show beneficial effect in therapeutics of some disease like Alzheimer and cancer.[8]
Phycoerythrin is an accessory pigment to the main chlorophyll pigments responsible for photosynthesis. The light energy is captured by phycoerythrin and is then passed on to the reaction centre chlorophyll pair, most of the time via the phycobiliproteins phycocyanin and allophycocyanin.
Structural characteristics[edit]
Phycoerythrins except phycoerythrin 545 (PE545) are composed of (αβ) monomers assembled into disc-shaped (αβ)6 hexamers or (αβ)3 trimers with 32 or 3 symmetry and enclosing central channel. In phycobilisomes (PBS) each trimer or hexamer contains at least one linker protein located in central channel. B-phycoerythrin (B-PE) and R-phycoerythrin (R-PE) from red algae in addition to α and β chains have a third, γ subunit contributing both linker and light-harvesting functions, because it bears chromophores. [2]

4.
R-phycoerythrin is predominantly produced by red algae. The protein is made up of at least three different subunits and varies according to the species of algae that produces it. The subunit structure of the most common R-PE is (αβ)6γ. The α subunit has two phycoerythrobilins (PEB), the β subunit has 2 or 3 PEBs and one phycourobilin (PUB), while the different gamma subunits are reported to have 3 PEB and 2 PUB (γ1) or 1 or 2 PEB and 1 PUB (γ2). The molecular weight of R-PE is 250,000 daltons.
Crystal structures available in the Protein Data Bank [12] contain in one (αβ)2 or (αβγ)2 asymmetric unit of different phycoerythrins:


| Chromophore or other non-protein molecule |
Phycoerythrin | Chain | |||
|---|---|---|---|---|---|
| PE545 | B-PE | R-PE | other types | ||
| Bilins | 8 | 10 | 10 | 10 | α and β |
| - Phycoerythrobilin (PEB) | 6 | 10 | 0 or 8 | 8 | β (PE545) or α and β |
| - 15,16-dihydrobiliverdin (DBV) | 2 | — | — | — | α (-3 and -2) |
| - Phycocyanobilin (CYC) | — | — | 8 or 7 or 0 | — | α and β |
| - Biliverdine IX alpha (BLA) | — | — | 0 or 1 | — | α |
| - Phycourobilin (PUB) | — | — | 2 | 2 | β |
| 5-hydroxylysine (LYZ) | 1 or 2 | — | — | — | α (-3 or -3 and -2) |
| N-methyl asparagine (MEN) | 2 | 2 | 0 or 2 | 2 | β |
| Sulfate ion SO42− (SO4) | — | 5 or 1 | 0 or 2 | — | α or α and β |
| Chloride ion Cl− (CL) | 1 | — | — | — | β |
| Magnesium ion Mg2+ (MG) | 2 | — | — | — | α-3 and β |
| inspected PDB files | 1XG0 1XF6 1QGW |
3V57 3V58 |
1EYX 1LIA 1B8D |
2VJH | |

The assumed biological molecule of phycoerythrin 545 (PE545) is (αβ)2 or rather (α3β)(α2β). The numbers 2 and 3 after the α letters in second formula are part of chain names here, not their counts. The synonym cryptophytan name of α3 chain is α1 chain.
The largest assembly of B-phycoerythrin (B-PE) is (αβ)3 trimer [9][10]. However, preparations from red algae yield also (αβ)6 hexamer [2]. In case of R-phycoerythrin (R-PE) the largest assumed biological molecule here is (αβγ)6, (αβγ)3(αβ)3 or (αβ)6 dependently on publication, for other phycoerythrin types (αβ)6. These γ chains from the Protein Data Bank are very small and consist only of three or six recognizable amino acids [15][16], whereas described at the beginning of this section linker γ chain is large (for example 277 amino acid long 33 kDa in case of γ33 from red algae Aglaothamnion neglectum) [17][2]. This is because the electron density of the gamma-polypeptide is mostly averaged out by its threefold crystallographic symmetry and only a few amino acids can be modeled [15][16][18][19].
For (αβγ)6, (αβ)6 or (αβγ)3(αβ)3 the values from the table should be simply multiplied by 3, (αβ)3 contain intermediate numbers of non-protein molecules. (this non sequitur needs to be corrected)
In phycoerythrin PE545 above, one α chain (-2 or -3) binds one molecule of billin, in other examples it binds two molecules. The β chain always binds to three molecules. The small γ chain binds to none.
Two molecules of N-methyl asparagine are bound to the β chain, one 5-hydroxylysine to α (-3 or -2), one Mg2+ to α-3 and β, one Cl− to β, 1–2 molecules of SO2−
4 to α or β.
Below is sample crystal structure of R-phycoerythrin from Protein Data Bank:

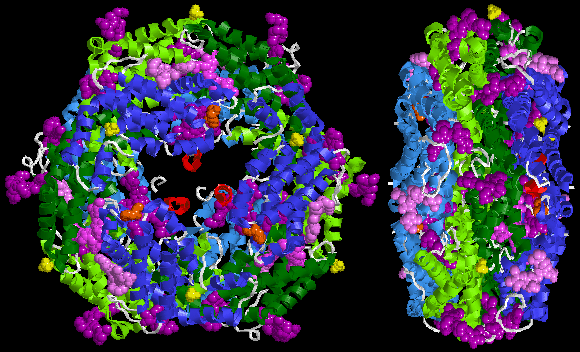
Spectral characteristics[edit]

Absorption peaks in the visible light spectrum are measured at 495 and 545/566 nm, depending on the chromophores bound and the considered organism. A strong emission peak exists at 575 ± 10 nm. (Phycoerythrin absorbs slightly blue-green/yellowish light and emits slightly orange-yellow light.)
| Property | Value |
|---|---|
| Absorption maximum | 565 nm |
| Additional Absorption peak | 498 nm |
| Emission maximum | 573 nm |
| Extinction Coefficient (ε) | 1.96×106 M−1 cm−1 |
| Quantum Yield (QY) | 0.84 |
| Brightness (ε × QY) | 1.65×106 M−1 cm−1 |
PEB and DBV bilins in PE545 absorb in the green spectral region too, with maxima at 545 and 569 nm respectively. The fluorescence emission maximum is at 580 nm. [3]
R-Phycoerythrin variations[edit]

As mentioned above, phycoerythrin can be found in a variety of algal species. As such, there can be variation in the efficiency of absorbance and emission of light required for facilitation of photosynthesis. This could be a result of the depth in the water column that a specific alga typically resides and a consequent need for greater or less efficiency of the accessory pigments.
With advances in imaging and detection technology which can avoid rapid photobleaching, protein fluorophores have become a viable and powerful tool for researchers in fields such as microscopy, microarray analysis and Western blotting. In light of this, it may be beneficial for researchers to screen these variable R-phycoerythrins to determine which one is most appropriate for their particular application. Even a small increase in fluorescent efficiency could reduce background noise and lower the rate of false-negative results.
Practical applications[edit]
R-Phycoerythrin (also known as PE or R-PE) is useful in the laboratory as a fluorescence-based indicator for the presence of cyanobacteria and for labeling antibodies, most often for flow cytometry. Its also used in microarray assays, ELISAs, and other applications that require high sensitivity but not photostability.[20] Its use is limited in immunofluorescence microscopy due to its rapid photobleaching characteristics. There are also other types of phycoerythrins, such as B-phycoerythrin, which have slightly different spectral properties. B-Phycoerythrin absorbs strongly at about 545 nm (slightly yellowish green) and emits strongly at 572 nm (yellow) instead and could be better suited for some instruments. B-Phycoerythrin may also be less "sticky" than R-phycoerythrin and contributes less to background signal due to nonspecific binding in certain applications.[citation needed] However, R-PE is much more commonly available as an antibody conjugate. Phycoerythrin (PE, λA max = 540–570 nm; λF max = 575–590 nm)[8]
R-Phycoerythrin and B-phycoerythrin are among the brightest fluorescent dyes ever identified.
References[edit]
- ^ Basheva D, Moten D, Stoyanov P, Belkinova D, Mladenov R, Teneva I (November 2018). "Content of phycoerythrin, phycocyanin, alophycocyanin and phycoerythrocyanin in some cyanobacterial strains: Applications". Engineering in Life Sciences. 18 (11): 861–866. Bibcode:2018EngLS..18..861B. doi:10.1002/elsc.201800035. PMC 6999198. PMID 32624879.
- ^ a b c d e Ficner R, Huber R (November 1993). "Refined crystal structure of phycoerythrin from Porphyridium cruentum at 0.23-nm resolution and localization of the gamma subunit". European Journal of Biochemistry. 218 (1): 103–106. doi:10.1111/j.1432-1033.1993.tb18356.x. PMID 8243457.
- ^ a b c van der Weij-De Wit CD, Doust AB, van Stokkum IH, Dekker JP, Wilk KE, Curmi PM, et al. (December 2006). "How energy funnels from the phycoerythrin antenna complex to photosystem I and photosystem II in cryptophyte Rhodomonas CS24 cells" (PDF). The Journal of Physical Chemistry B. 110 (49): 25066–25073. doi:10.1021/jp061546w. PMID 17149931. Archived from the original (PDF) on 30 October 2013.
- ^ Hine, Robert; Martin, Elizabeth, eds. (2015). "Phycoerythrin". A Dictionary of Biology. Oxford University Press. doi:10.1093/acref/9780198714378.001.0001. ISBN 978-0-19-871437-8. Retrieved 2021-11-27 – via login.aurarialibrary.idm.oclc.org.
- ^ Dammeyer T, Frankenberg-Dinkel N (September 2006). "Insights into phycoerythrobilin biosynthesis point toward metabolic channeling". The Journal of Biological Chemistry. 281 (37): 27081–27089. doi:10.1074/jbc.M605154200. PMID 16857683.
- ^ Six C, Thomas JC, Garczarek L, Ostrowski M, Dufresne A, Blot N, et al. (2007). "Diversity and evolution of phycobilisomes in marine Synechococcus spp.: a comparative genomics study". Genome Biology. 8 (12): R259. doi:10.1186/gb-2007-8-12-r259. PMC 2246261. PMID 18062815.
- ^ Glazer AN (1985). "Light harvesting by phycobilisomes". Annual Review of Biophysics and Biophysical Chemistry. 14: 47–77. doi:10.1146/annurev.bb.14.060185.000403. PMID 3924069.
- ^ a b Sonani RR, Rastogi RP, Patel R, Madamwar D (February 2016). "Recent advances in production, purification and applications of phycobiliproteins". World Journal of Biological Chemistry. 7 (1): 100–109. doi:10.4331/wjbc.v7.i1.100. PMC 4768114. PMID 26981199.
- ^ a b Camara-Artigas, A. (2011-12-16). "Crystal Structure of the B-phycoerythrin from the red algae Porphyridium cruentum at pH8". RCSB Protein Data Bank (PDB). doi:10.2210/pdb3v57/pdb. PDB ID: 3V57. Retrieved 12 October 2012.
{{cite journal}}: Cite journal requires|journal=(help) - ^ a b Camara-Artigas A, Bacarizo J, Andujar-Sanchez M, Ortiz-Salmeron E, Mesa-Valle C, Cuadri C, et al. (October 2012). "pH-dependent structural conformations of B-phycoerythrin from Porphyridium cruentum". The FEBS Journal. 279 (19): 3680–3691. doi:10.1111/j.1742-4658.2012.08730.x. PMID 22863205. S2CID 31253970. PDB ID: 3V57.
- ^ a b c d Image created with RasTop (Molecular Visualization Software).
- ^ "Protein Data Bank". RCSB Protein Data Bank (PDB). Archived from the original on 28 August 2008. Retrieved 12 October 2012.
- ^ Doust AB, Marai CN, Harrop SJ, Wilk KE, Curmi PM, Scholes GD (2004-09-16). "High resolution crystal structure of phycoerythrin 545 from the marine cryptophyte rhodomonas CS24". RCSB Protein Data Bank (PDB). doi:10.2210/pdb1xg0/pdb. PDB ID: 1XG0. Retrieved 11 October 2012.
{{cite journal}}: Cite journal requires|journal=(help) - ^ Doust AB, Marai CN, Harrop SJ, Wilk KE, Curmi PM, Scholes GD (November 2004). "Developing a structure-function model for the cryptophyte phycoerythrin 545 using ultrahigh resolution crystallography and ultrafast laser spectroscopy". Journal of Molecular Biology. 344 (1): 135–153. doi:10.1016/j.jmb.2004.09.044. PMID 15504407. PDB ID: 1XG0.
- ^ a b c d Contreras-Martel C, Martinez-Oyanedel J, Bunster M, Legrand P, Piras C, Vernede X, Fontecilla-Camps JC (2000-05-09). "Crystal structure of R-phycoerythrin at 2.2 angstroms". RCSB Protein Data Bank (PDB). doi:10.2210/pdb1eyx/pdb. PDB ID: 1EYX. Retrieved 11 October 2012.
{{cite journal}}: Cite journal requires|journal=(help) - ^ a b c d Contreras-Martel C, Martinez-Oyanedel J, Bunster M, Legrand P, Piras C, Vernede X, Fontecilla-Camps JC (January 2001). "Crystallization and 2.2 A resolution structure of R-phycoerythrin from Gracilaria chilensis: a case of perfect hemihedral twinning". Acta Crystallographica. Section D, Biological Crystallography. 57 (Pt 1): 52–60. doi:10.1107/S0907444900015274. PMID 11134927. S2CID 216930. PDB ID: 1EYX.
- ^ Apt KE, Hoffman NE, Grossman AR (August 1993). "The gamma subunit of R-phycoerythrin and its possible mode of transport into the plastid of red algae". The Journal of Biological Chemistry. 268 (22): 16208–16215. doi:10.1016/S0021-9258(19)85407-8. PMID 8344905.
- ^ Ritter S, Hiller RG, Wrench PM, Welte W, Diederichs K (1999-01-29). "Crystal structure of a phycourobilin-containing phycoerythrin". doi:10.2210/pdb1b8d/pdb. PDB ID: 1B8D.
{{cite journal}}: Cite journal requires|journal=(help) - ^ Ritter S, Hiller RG, Wrench PM, Welte W, Diederichs K (June 1999). "Crystal structure of a phycourobilin-containing phycoerythrin at 1.90-A resolution". Journal of Structural Biology. 126 (2): 86–97. doi:10.1006/jsbi.1999.4106. PMID 10388620. PDB ID: 1B8D.
- ^ "R-phycoerythrin (R-PE) - US". www.thermofisher.com. Retrieved 2021-12-11.
External links[edit]
- Phycoerythrin at the U.S. National Library of Medicine Medical Subject Headings (MeSH)
