Cladistics
| Part of a series on |
| Evolutionary biology |
|---|
 |
Cladistics (from Greek κλάδος, klados, i.e. "branch"[1]) is an approach to biological classification in which items are grouped together based on whether or not they have one or more shared unique characteristics that come from the group's last common ancestor and are not present in more distant ancestors. Therefore, members of the same group are thought to share a common history and are considered to be more closely related.[2][3][4][5]
The original method for cladistic analysis and the school of taxonomy derived from it originated in the work of the German entomologist Willi Hennig, who referred to it as "phylogenetic systematics" (also the title of his 1966 book); the use of the terms "cladistics" and "clade" was popularized by other researchers. The techniques and sometimes the names have been successfully applied in other disciplines: for example, to determine the relationships between the surviving manuscripts of the Canterbury Tales,[6] or also between 53 manuscripts of the Sanskrit Charaka Samhita.[7]
History of cladistics
This section needs expansion. You can help by adding to it. (September 2012) |
What is now called the cladistic method appeared as early as 1901 with a work by P.C. Mitchell (for birds)[8][9] and subsequently by Tillyard (for insects) in 1921,[10] and W. Zimmermann (for plants) in 1943.[11] The term "clade" was introduced in 1958 by Julian Huxley after having been coined by Lucien Cuénot in 1940,[12], cladogenesis" in 1958,[13] "cladistic" by Cain and Harrison in 1960,[14], "cladist" (for an adherent of Hennig's school) by Mayr in 1965[15], and "cladistics" in 1966.[16] Hennig referred to his own approach as "phylogenetic systematics". From the time of his original formulation until the end of the 1970s, cladistics competed as an analytical and philosophical approach to phylogenetic inference with phenetics and so-called evolutionary taxonomy. In the 1990s, the development of effective polymerase chain reaction techniques allowed the application of cladistic methods to biochemical and molecular genetic traits of organisms, as well as to anatomical ones, vastly expanding the amount of data available for phylogenetics. At the same time, cladistics rapidly became the dominant set of methods of phylogenetics in evolutionary biology, because computers made it possible to process large quantities of data about organisms and their characteristics.
The way for computational phylogenetics was paved by phenetics,[17] a set of methods commonly used from the 1950s to 1980s and to some degree later. Phenetics did not try to reconstruct phylogenetic trees; rather, it tried to build dendrograms from similarity data; its algorithms required less computer power than phylogenetic ones.
Methodology
The cladistic method interprets each character state transformation implied by the distribution of shared character states among taxa (or other terminals) as a potential piece of evidence for grouping. The outcome of a cladistic analysis is a cladogram – a tree-shaped diagram (dendrogram[18]) that is interpreted to represent the best hypothesis of phylogenetic relationships. Although traditionally such cladograms were generated largely on the basis of morphological characters and originally calculated by hand, genetic sequencing data and computational phylogenetics are now very commonly used in phylogenetic analyses, and the parsimony criterion has been abandoned by many phylogeneticists in favor of more "sophisticated" but less parsimonious evolutionary models of character state transformation. Cladists contend that these models are unjustified.[clarification needed]
Every cladogram is based on a particular dataset analyzed with a particular method. Datasets are tables consisting of molecular, morphological, ethological[19] and/or other characters and a list of operational taxonomic units (OTUs) which may be genes, individuals, populations, species, or larger taxa that are presumed to be monophyletic and that are presumed to form, all together, one large clade; phylogenetic analysis infers the branching pattern within that clade. Different datasets and different methods, not to mention violations of the mentioned assumptions, often result in different cladograms. Only scientific investigation can show which is more likely to be correct.
Until recently, for example, cladograms like the following have generally been accepted as accurate representations of the ancestral relations among turtles, lizards, crocodilians, and birds:[20]
If this phylogenetic hypothesis is correct, then the last common ancestor of turtles and birds, at the ⊣ connection near the ▼ (a ⊤ in some browsers) lived earlier than the last common ancestor of lizards and birds, near the ♦. Most molecular evidence, however, produces cladograms more like this:[21]
Diapsida ♦
|
|
If this is accurate, then the last common ancestor of turtles and birds lived later than the last common ancestor of lizards and birds. Since the cladograms provide competing accounts of real events, at most one of them is correct.
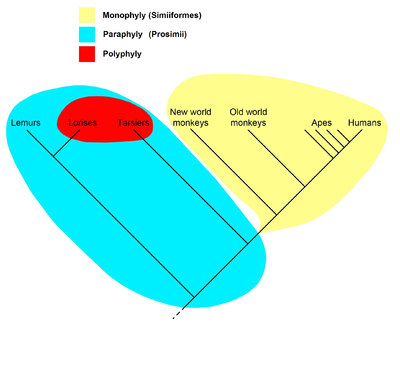
The cladogram to the right represents the current universally accepted hypothesis that all primates, including strepsirrhines like the lemurs and lorises, had a common ancestor all of whose descendants were primates, and so form a clade; the name Primates is therefore recognized for this clade. Within the primates, all anthropoids (monkeys, apes and humans) are hypothesized to have had a common ancestor all of whose descendants were anthropoids, so they form the clade called Anthropoidea. The "prosimians", on the other hand, form a paraphyletic taxon. The name Prosimii is not used in phylogenetic nomenclature, which names only clades; the "prosimians" are instead divided between the clades Strepsirhini and Haplorhini, where the latter contains Tarsiiformes and Anthropoidea.
Cladistics has developed several innovations in 7 major areas[22][23]:
- parsimony (optimization) criteria
- Camin-Sokal, 1965
- Wagner ( Kluge and Farris, 1969; Farris, 1970)
- Fitsch, 1971
- Dollo (LeQuesne, 1969; Farris, 1977)
- search strategies
- NNI (nearest-neighbour interchange), Robinson, 1971
- SPR (subtree pruning and regrafting), Swofford and Olsen, 1990
- TBR (tree bisection and reconnection), Swofford and Olsen, 1990
- branch and bound, Hendy and Penny, 1982
- consensus methods
- Adams, 1972
- Nelson, 1979
- majority, Margush and McMorris, 1980
- strict, Sokal and Rohlf, 1980
- Bremer (=combinable components=semi-strict), Bremer, 1990
- reduced, Wilkinson, 1994, 1995
- group support measures
- bootstrap, Felsenstein, 1985
- Bremer support, 1994
- symmetrical resampling, Goloboff et al, 2007
- homology measures
- CI (consistency index), Kluge and Farris, 1969
- RI (retention index), Farris, 1989
- RC (rescaled consistency index), Farris, 1989
- HER (homoplasy excess ratio), Archie, 1989
- DDI (data decisiveness index), Goloboff, 1991
- a posteriori character weighting
- dynamic, Farris, 1969
- successive, Farris, 1969
- implied, Goloboff, 1993
- algorithm, software packages
- PHYLIP, Felsenstein, 1980
- PHYSIS, Mikevich and Farris, 1982
- PAUP, Swofford
- Mesquite, Maddison and Maddison
- Hennig86, Farris, 1988
- MacClade, Maddison and Maddison, 1986
- TNT, Goloboff, Farris, Nixon, 1998
- Winclada, Nixon, 1999
Terminology for character states
The following terms, coined by Hennig, are used to identify shared or distinct character states among groups:[24][25][26]
- A plesiomorphy ("close form") or ancestral state is a character state that a taxon is inferred to have been retained from its ancestors. When two or more taxa that are not nested within each other share a plesiomorphy, it is a symplesiomorphy (from syn-, "together"). Symplesiomorphies do not mean that the taxa that exhibit that character state are necessarily closely related. For example, Reptilia is traditionally characterized by (among other things) being cold-blooded (i.e. not maintaining a constant high body temperature), whereas birds are warm-blooded. Since cold-bloodedness is a plesiomorphy, inherited from the common ancestor of traditional reptiles and birds, and thus a symplesiomorphy of turtles, snakes and crocodiles (among others), it does not mean that turtles, snakes and crocodiles form a clade that excludes the birds.
- An apomorphy ("separate form") or derived state is an innovation. It can thus be used to diagnose a clade – or even to define a clade name in phylogenetic nomenclature. Features that are derived in individual taxa (a single species or a group that is represented by a single terminal in a given phylogenetic analysis) are called autapomorphies (from auto-, "self"). Autapomorphies express nothing about relationships among groups; clades are identified (or defined) by synapomorphies (from syn-, "together"). For example, the possession of digits that are homologous with those of Homo sapiens is a synapomorphy within the vertebrates. The tetrapods can be singled out as consisting of the first vertebrate with such digits together with all descendants of this vertebrate (an apomorphy-based phylogenetic definition).[27] Importantly, snakes and other tetrapods that do not have digits are nonetheless tetrapods: other characters, such as amniotic eggs and diapsid skulls, indicate that they descended from ancestors that possessed digits which are homologous with ours.
- A character state is homoplastic or "an instance of homoplasy" if it is shared by two or more organisms, but is inferred, based on the distribution of other features, to have been absent in their common ancestor. It is therefore inferred to have evolved by convergence or reversal. Both mammals and birds are able to maintain a high constant body temperature (i.e. they are 'warm-blooded'). However, the most parsimonious cladogram explaining all of their features implies that the ancestors of each group did not share this character state, so it must have evolved independently. Warm-bloodedness is separately a synapomorphy of mammals (or a larger clade) and of birds (or a larger clade), but it is not a synapomorphy of these two clades. Hennig's Auxiliary Principle [28] states that shared character states should be considered evidence of grouping unless they are contradicted by the weight of other evidence; thus, homoplasy of some feature among members of a group may only be inferred after a phylogenetic hypothesis for that group has been established.
The terms plesiomorphy and apomorphy are relative; their application depends on the position of a group within a tree. For example, when trying to decide whether the tetrapods form a clade, an important question is whether having four limbs is a synapomorphy of all the taxa to be included within Tetrapoda: did all the possible members of the Tetrapoda inherit four limbs from a common ancestor, whereas all other vertebrates did not? By contrast, for a group within the tetrapods, such as birds, having four limbs is a plesiomorphy. Using these two terms allows a greater precision in the discussion of homology, in particular allowing clear expression of the hierarchical relationships among different homologous features.
It can be difficult to decide whether a character state is in fact the same and thus can be classified as a synapomorphy which may identify a monophyletic group or whether it only appears to be the same and is thus a homoplasy which cannot identify such a group. There is a danger of circular reasoning: assumptions about the shape of a phylogenetic tree are used to justify decisions about character states, which are then used as evidence for the shape of the tree.[29] Phylogenetics uses various forms of parsimony to decide such questions; the conclusions reached often depend on the dataset and the methods. Such is the nature of empirical science, and for this reason, most cladists refer to their cladograms as hypotheses of relationship. Cladograms that are supported by a large number and variety of different kinds of characters are viewed as more robust than those based on more limited evidence.
Terminology for taxa
Mono-, para- and polyphyletic taxa can be understood based on the shape of the tree (as done above), as well as based on their character states.[25][26][30] These are compared in the table below.
| Term | Node-based definition | Character-based definition |
|---|---|---|
| Monophyly | A monophyletic assemblage consists of an ancestral species and all of its descendants. Also known as a clade. | A clade is characterized by one or more apomorphies: derived character states inferred to have been present in the first member of the taxon, inherited by its descendants (unless secondarily lost), and not inherited by any other taxa. |
| Paraphyly | A paraphyletic assemblage consists of an ancestral species and some, but not all, of its descendants. It is like a clade from which some smaller clades have been removed.[31] (Removing one clade produces a singly paraphyletic assemblage, removing two a doubly paraphylectic assemblage, and so on.[32]) | A paraphyletic assemblage is characterized by one or more plesiomorphies: character states inferred to have been inherited from ancestors but not present in all of their descendants. As a consequence, a paraphyletic assemblage is truncated, in that it excludes one or more clades from an otherwise monophyletic taxon. An alternative name is evolutionary grade, referring to an ancestral character state within the group. While paraphyletic assemblages are popular among paleontologists and evolutionary taxonomists, cladists do not recognize paraphyletic assemblages as having any formal information content - they are merely parts of clades. |
| Polyphyly | A polyphyletic assemblage is like a paraphyletic assemblage that only includes some descendants of an ancestral species, and excludes the ancestral species itself. | A polyphyletic assemblage is characterized by one or more homoplasies: character states which are inferred to have converged or reverted so as to appear to be the same but which the cladogram implies have not been inherited from a common ancestor. No systematist recognizes polyphyletic assemblages as taxonomically meaningful entities, although ecologists sometimes consider them meaningful labels for functional participants in ecological communities (e. g., primary producers, detritivores, etc.). |
Criticism
Cladistics, either generally or in specific applications, has been criticized from its beginnings. Decisions as to whether particular character states are homologous, a precondition of their being synapomorphies, have been challenged as involving circular reasoning and subjective judgements.[33]
However, homology is usually determined from the analysis through the results which are evaluated with homology measures, mainly the CI (consistency index) and RI (retention index), which makes the process objective. Also, homology can be equated to synapomorphy, which is what Patterson has done [34].
Application to disciplines other than biology
The comparisons used to acquire data on which cladograms can be based are not limited to the field of biology.[35] Any group of individuals or classes that are hypothesized to have a common ancestor, and to which a set of common characteristics may or may not apply, can be compared pairwise. Cladograms can be used to depict the hypothetical descent relationships within groups of items in many different academic realms. The only requirement is that the items have characteristics that can be identified and measured.
Recent attempts to use cladistic methods outside of biology address the reconstruction of lineages in:
- Anthropology and archeology:[36] Cladistic methods have been used to reconstruct the development of cultures or artifacts using groups of cultural traits or artifact features.
- Historical linguistics:[37] Cladistic methods have been used to reconstruct the phylogeny of languages using linguistic features. This is similar to the traditional comparative method of historical linguistics, but is more explicit in its use of parsimony and allows much faster analysis of large datasets (computational phylogenetics).
- Textual criticism or stemmatics:[7][38] Cladistic methods have been used to reconstruct the phylogeny of manuscripts of the same work (and reconstruct the lost original) using distinctive copying errors as apomorphies. This differs from traditional historical-comparative linguistics in enabling the editor to evaluate and place in genetic relationship large groups of manuscripts with large numbers of variants that would be impossible to handle manually. It also enables parsimony analysis of contaminated traditions of transmission that would be impossible to evaluate manually in a reasonable period of time.
- Astrophysics[39] infers the history of relationships between galaxies to create branching diagram hypotheses of galaxy diversification.
See also
Notes and references
- ^ "clade". Online Etymology Dictionary.
- ^ Columbia Encyclopedia
- ^ http://www.ucmp.berkeley.edu/clad/clad1.html
- ^ Oxford Dictionary of English
- ^ Oxford English Dictionary
- ^ "Canterbury Tales Project". Retrieved 4 July 2009.
- ^ a b Maas 2010
- ^ Schuh, Randall. 2000. Biological Systematics: Principles and Applications, p.7 (citing Nelson and Platnick, 1981). Cornell U. Press (books.google)
- ^ Folinsbee, Kaila et al. 2007. 5 Quantitative Approaches to Phylogenetics, p. 172. Rev. Mex. Div. 225-52 (kfolinsb.public.iastate.edu)
- ^ Folinsbee, Kaila et al. 2007. 5 Quantitative Approaches to Phylogenetics, p.172. Rev. Mex. Div. 225-52 (kfolinsb.public.iastate.edu)
- ^ Schuh, Randall. 2000. Biological Systematics: Principles and Applications, p.7. Cornell U. Press
- ^ Cuénot 1940
- ^ Webster's 9th New Collegiate Dictionary
- ^ Cain & Harrison 1960
- ^ Dupuis 1984
- ^ Webster's 9th New Collegiate Dictionary
- ^ Mayr 1982, p. 221
- ^ Weygoldt 1998
- ^ Jerison 2003, p. 254
- ^ Benton, Michael J. (2005), Vertebrate Palaeontology, Blackwell, pp. 214, 233, ISBN 978-0-632-05637-8
- ^ Lyson, Tyler; Gilbert, Scott F. (2009), "Turtles all the way down: loggerheads at the root of the chelonian tree" (PDF), Evolution & Development, 11 (2): 133–135, doi:10.1111/j.1525-142X.2009.00325.x
- ^ Forey, Peter et al. 1992. Cladistics, Oxford U. Press
- ^ Kitching, Ian et al. 1998. Cladistics. Oxford U. Press.
- ^ Patterson 1982, pp. 21–74
- ^ a b Patterson 1988
- ^ a b de Pinna 1991
- ^ Laurin & Anderson 2004
- ^ Hennig 1966
- ^ James & Pourtless IV 2009, p. 25: "Synapomorphies are invoked to defend the hypothesis; the hypothesis is invoked to defend the synapomorphies."
- ^ Patterson 1982
- ^ Many sources give a verbal definition of 'paraphyletic' which does not require the missing groups to be monophyletic. However, when diagrams are presented representing paraphyletic groups, these invariably show the missing groups as monophyletic. See e.g. Wiley et al. 1991, p. 4.
- ^ Taylor 2003
- ^ Adrain, Edgecombe & Lieberman 2002, pp. 56–57
- ^ Forey, Peter et al. 1992. Cladistics,1st ed., p. 9, Oxford U. Press.
- ^ Mace, Clare & Shennan 2005, p. 1
- ^ Lipo et al. 2006
- ^ Oppenheimer 2006, pp. 290–300, 340–56
- ^ Robinson & O’Hara 1996
- ^ Fraix-Burnet et al. 2006
Bibliography
- Adrain, Jonathan M.; Edgecombe, Gregory D.; Lieberman, Bruce S. (2002), Fossils, Phylogeny, and Form: An Analytical Approach, New York: Kluwer Academic, ISBN 978-0-306-46721-9, retrieved 15 August 2012
{{citation}}: Unknown parameter|lastauthoramp=ignored (|name-list-style=suggested) (help) - Baron, C.; Høeg, J.T. (2005), "Gould, Scharm and the paleontologocal perspective in evolutionary biology", in Koenemann, S.; Jenner, R.A. (eds.), Crustacea and Arthropod Relationships, CRC Press, pp. 3–14, ISBN 978-0-8493-3498-6, retrieved 15 October 2008
{{citation}}: Unknown parameter|lastauthoramp=ignored (|name-list-style=suggested) (help) - Benton, M. J. (2000), "Stems, nodes, crown clades, and rank-free lists: is Linnaeus dead?" (PDF), Biological Reviews, 75 (4): 633–648, doi:10.1111/j.1469-185X.2000.tb00055.x, PMID 11117201
- Benton, M. J. (2004), Vertebrate Palaeontology (3rd ed.), Oxford: Blackwell Science, ISBN 978-0-632-05637-8
- Cain, A. J.; Harrison, G. A. (1960), "Phyletic weighting", Proceedings of the Zoological Society of London, 35: 1–31
- Cuénot, Lucien (1940), "Remarques sur un essai d'arbre généalogique du règne animal", Comptes Rendus de l'Académie des Sciences de Paris, 210: 23–27.
{{citation}}: Invalid|ref=harv(help) Available free online at http://gallica.bnf.fr (No direct URL). This is the paper credited by Hennig 1979 for the first use of the term 'clade'. - Dupuis, Claude (1984), "Willi Hennig's impact on taxonomic thought", Annual Review of Ecology and Systematics, 15: 1–24, ISSN 0066-4162.
{{citation}}: CS1 maint: ref duplicates default (link) - Farris, James S. (1977), "On the phenetic approach to vertebrate classification", in Hecht, M. K.; Goody, P. C.; Hecht, B. M. (eds.), Major Patterns in Vertebrate Evolution, Plenum, New York, pp. 823–850
- Farris, James S. (1979a), "On the naturalness of phylogenetic classification", Systematic Zoology, 28: 200–214
- Farris, James S. (1979b), "The information content of the phylogenetic system", Systematic Zoology, 28: 483–519
- Farris, James S. (1980), "The efficient diagnoses of the phylogenetic system", Systematic Zoology, 29: 386–401
- Farris, James S. (1983), "The logical basis of phylogenetic analysis", in Platnick, Norman I.; Funk, Vicki A. (eds.), Advances in Cladistics, vol. 2, Columbia University Press, New York, pp. 7–36
- Fraix-Burnet, D.; Choler, P.; Douzery, E.J.P.; Verhamme, A. (2006), "Astrocladistics: A Phylogenetic Analysis of Galaxy Evolution II. Formation and Diversification of Galaxies", Journal of Classification, 23 (1): 57–78, arXiv:astro-ph/0602580, Bibcode:2006JClas..23...57F, doi:10.1007/s00357-006-0004-4
- Hennig, Willi (1966), Phylogenetic systematics (tr. D. Dwight Davis and Rainer Zangerl), Urbana, IL: Univ. of Illinois Press (reprinted 1979 and 1999), ISBN 0-252-06814-9
- Hennig, Willi (1975), "'Cladistic analysis or cladistic classification?': a reply to Ernst Mayr" (PDF), Systematic Zoology, 24 (2): 244–256, doi:10.2307/2412765, JSTOR 2412765,
{{citation}}: CS1 maint: ref duplicates default (link) responding to Mayr 1974. - Hennig, Willi (1999), Phylogenetic systematics (3rd edition of 1966 book), Urbana: University of Illinois Press, ISBN 0-252-06814-9 Translated from manuscript in German eventually published in 1982 (Phylogenetische Systematik, Verlag Paul Parey, Berlin).
- Hull, David (1988), Science as a Process, University of Chicago Press, ISBN 978-0-226-36051-5
- James, Frances C.; Pourtless IV, John A. (2009), Cladistics and the Origin of Birds: A Review and Two New Analyses (PDF), Ornithological Monographs, No. 66, American Ornithologists' Union, ISBN 978-0-943610-85-6, retrieved 14 December 2010
{{citation}}: Unknown parameter|lastauthoramp=ignored (|name-list-style=suggested) (help) - Jerison, Harry J. (2003), "On Theory in Comparative Psychology", in Sternberg, Robert J.; Kaufman, James C. (eds.), The Evolution of Intelligence, Mahwah, NJ: Lawrence Erlbaum Associates, Inc., ISBN 0-12-385250-1
- Laurin, M.; Anderson, J. (2004), "Meaning of the Name Tetrapoda in the Scientific Literature: An Exchange" (PDF), Systematic Biology, 53 (1): 68–80, doi:10.1080/10635150490264716, PMID 14965901
{{citation}}: Unknown parameter|lastauthoramp=ignored (|name-list-style=suggested) (help) - Lipo, Carl; O'Brien, Michael J.; Collard, Mark; Shennan, Stephen J., eds. (2006), Mapping Our Ancestors: Phylogenetic Approaches in Anthropology and Prehistory, Piscataway: Transaction Publishers, ISBN 978-0-202-30751-0
- Maas, Philipp (2010), Jürgen, Hanneder; Maas, Philipp (eds.), Wiener Zeitschrift für die Kunde Südasiens, 52–53, Vienna: Austrian Academy of Sciences: 63–120, doi:10.1553/wzks2009-2010s63
{{citation}}:|contribution=ignored (help); Missing or empty|title=(help); Unknown parameter|booktitle=ignored (help) - Mace, Ruth; Clare, Clare J.; Shennan, Stephen, eds. (2005), The Evolution of Cultural Diversity: A Phylogenetic Approach, Portland: Cavendish Press, ISBN 978-1-84472-099-6
- Mayr, Ernst (1974), "Cladistic analysis or cladistic classification?" (PDF), Zeitschrift fűr Zoologische Systematik und Evolutionforschung, 12: 94–128, doi:10.1111/j.1439-0469.1974.tb00160.x, retrieved 14 December 2010
- Mayr, Ernst (1976), Evolution and the diversity of life (Selected essays), Cambridge, MA: Harvard University Press, ISBN 0-674-27105-X
{{citation}}: CS1 maint: ref duplicates default (link) Reissued 1997 in paperback. Includes a reprint of Mayr's 1974 anti-cladistics paper at pp. 433–476, "Cladistic analysis or cladistic classification." This is the paper to which Hennig 1975 is a response. - Mayr, Ernst (1978), "Origin and history of some terms in systematic and evolutionary biology", Systematic Zoology, 27 (1): 83–88, doi:10.2307/2412818, JSTOR 2412818.
{{citation}}: CS1 maint: ref duplicates default (link) - Mayr, Ernst (1982), The growth of biological thought: diversity, evolution and inheritance, Cambridge, MA: Harvard University Press, ISBN 0-674-36446-5
{{citation}}: CS1 maint: ref duplicates default (link) - Oppenheimer, Stephen (2006), The Origins of the British, London: Robinson, ISBN 978-0-7867-1890-0
- Patterson, Colin (1982), "Morphological characters and homology", in Joysey, Kenneth A; Friday, A. E. (eds.), Problems in Phylogenetic Reconstruction, Systematics Association Special Volume 21, London: Academic Press, ISBN 0-12-391250-4
{{citation}}: CS1 maint: ref duplicates default (link). - Patterson, Colin (1988), "Homology in classical and molecular biology", Molecular Biology and Evolution, 5 (6): 603–625, PMID 3065587
- de Pinna, M.G.G (1991), "Concepts and tests of homology in the cladistic paradigm", Cladistics, 7 (4): 367–394, doi:10.1111/j.1096-0031.1991.tb00045.x
- de Queiroz, K.; Gauthier, J. (1992), "Phylogenetic taxonomy" (PDF), Annual Review of Ecology and Systematics, 23: 449–480
{{citation}}: Unknown parameter|lastauthoramp=ignored (|name-list-style=suggested) (help) - Robinson, Peter M.W.; O’Hara, Robert J. (1996), "Cladistic analysis of an Old Norse manuscript tradition", Research in Humanities Computing, 4: 115–137, retrieved 13 December 2010
{{citation}}: Unknown parameter|lastauthoramp=ignored (|name-list-style=suggested) (help) - Schuh, Randall T.; Brower, Andrew V.Z. (2009), Biological Systematics: Principles and Applications (2nd ed.), Cornell University Press, ISBN 978-0-8014-4799-0
{{citation}}: Unknown parameter|lastauthoramp=ignored (|name-list-style=suggested) (help) - Taylor, Mike (2003), What do terms like monophyletic, paraphyletic and polyphyletic mean?, retrieved 13 December 2010
- Tremblay, Frederic (2013), "Nicolai Hartmann and the Metaphysical Foundation of Phylogenetic Systematics", Biological Theory, 7 (1): 56–68, doi:10.1007/s13752-012-0077-8
- Weygoldt, P. (1998), "Evolution and systematics of the Chelicerata", Experimental and Applied Acarology, 22 (2): 63–79, doi:10.1023/A:1006037525704
{{citation}}: Unknown parameter|month=ignored (help) - Wheeler, Quentin (2000), Species Concepts and Phylogenetic Theory: A Debate, Columbia University Press, ISBN 978-0-231-10143-1
- Wiley, E.O.; Siegel-Causey, D.; Brooks, D.R.; Funk, V.A. (1991), "Chapter 1 Introduction, terms and concepts", The Compleat Cladist: A Primer of Phylogenetic Procedures (PDF), The University of Kansas Museum of Natural History, ISBN 978-0-89338-035-9, retrieved 13 December 2010
{{citation}}: Unknown parameter|lastauthoramp=ignored (|name-list-style=suggested) (help) - Williams, P.A. (1992), "Confusion in cladism", Synthese, 01: 135–132, doi:10.1007/BF00484973
External links
- Willi Hennig Society
- Cladistics (scholarly journal of the Willi Hennig Society)
- Collins, Allen G.; Guralnick, Rob; Smith, Dave (1994–2005). "Journey into Phylogenetic Systematics". University of California Museum of Paleontology. Retrieved 21 January 2010.
- Felsenstein, Joe. "Phylogeny Programs". Seattle: University of Washington. Retrieved 21 January 2010.
- O'Neil, Dennis (1998–2008). "Classification of Living Things". San Marcos CA: Palomar College. Retrieved 21 January 2010.
- Robinson, Peter; O'Hara, Robert J. (1992). "Report on the Textual Criticism Challenge 1991". rjohara.net. Retrieved 21 January 2010.
- Theobald, Douglas (1999–2004). "Phylogenetics Primer". The TalkOrigins Archive. Retrieved 21 January 2010.
