HLA A1-B8 haplotype
 HLA region on chromosome 6 | ||||||||||||||||||||||||||
| Nicknames | ||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| "HL A1,8; HL A1,A8; HLA A1-B8; HLA A*01-B*08, HLA A*01:01-B*08:01" | ||||||||||||||||||||||||||
| Loci | ||||||||||||||||||||||||||
| ||||||||||||||||||||||||||
| Nodes | ||||||||||||||||||||||||||
| PopulationMaxima | Western Ireland | |||||||||||||||||||||||||
| Freq.Max | >15.0% | |||||||||||||||||||||||||
| Size and location | ||||||||||||||||||||||||||
| Genes | - | |||||||||||||||||||||||||
| Chromosome | 6 | |||||||||||||||||||||||||
| Location | 6p21.3 | |||||||||||||||||||||||||
| Size (kbps) | 1400 | |||||||||||||||||||||||||
HLA A1-B8 (Also:HL A1,8; HL A1,A8; HLA A1-Cw7-B8; HLA A*01-B*08, HLA A*0101-B*0801, HLA A*0101-Cw*0701-B*0801; HLA A*01:01-C*07:01-B*08:01) is a multigene haplotype that covers the MHC Class I region of the human major histocompatibility complex on chromosome 6. A multigene haplotype is set of inherited alleles covering several genes, or gene-alleles; common multigene haplotypes are generally the result of identity by descent from a common ancestor (share a recent common ancestor for that segment of the chromosome). Chromosomal recombination fragments multigene haplotypes as the distance to that ancestor increases in number of generations.
The haplotype can be written in an extended form covering the major histocompatibility loci as follows:
HLA A*01:01 : Cw*07:01 : B*08:01
However, there are many other gene-alleles within the haplotype. In Europe A1-B8 is found, generally as part of the HLA A1-B8-DR3-DQ2 haplotype. This haplotype is 4.7 million nucleotides in length and the second longest haplotype identified within the human genome. In Africa A1-B8 and India A1-B8 is associated with other genes and other variants of A*01 and B*08
Disease associations
[edit]Philosophically, A1-B8 is more than just two gene-alleles. These gene-alleles are markers for a haplotype, a stretch of chromosome 6 that contains many gene alleles. In its natural history this haplotype underwent some atypical selection, at the end of the period of evolution it became the predominant haplotypes in North/Western European ancestors. Today however the collection of genes is associated with increased incidences of certain diseases. Despite the fact that the associations have been known almost as long as A1 and "A8" were known, the role of factors affecting disease are still not clear.
A1-B8 and autoimmune diseases
[edit]A1-B8 serotype was associated with a number of diseases as "HL-A"' antigens were first being described. Among these were coeliac disease, autoimmune active chronic hepatitis, myasthenia gravis, Adrenocortical hyperfunction-Cushing's syndrome, primary biliary cirrhosis.[1][2][3][4][5][6]
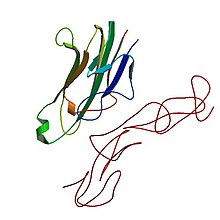
However, as study sizes increased and D serotypes were described in more detail, the association of these loci moved from the MHC class I loci to the MHC class II loci. Underlying this move was the HLA A1-B8-DR3-DQ2 haplotype, a haplotype that is in acute linkage disequilibrium in the European population.[7][8][9][10][11] This disequilibrium made it appear that A1 and other class I gene-alleles were disease factors, when these alleles were only attached to a long segment of conserved DNA that had disease associated genes on the other end. In at least 2 diseases, the risk of autoimmune disease extends beyond the class II region of the haplotype.
Systemic lupus erythematosus
[edit]The "HL-A1,8 phenotype" was found to be associated with severe systemic lupus erythematosus (SLE) (renal and central nervous system involvement) in Caucasian patients.[12] Two-point haplotype analysis between TNFB(B*01 allele) and HLA show that the allele is in linkage disequilibrium with HLA-A1, Cw7, B8, C4A(Null), DR3, DQ2.5.[13]
Type 1 diabetes
[edit]While type 1 diabetes shows an extended association on the HLA A1-B8-DR3-DQ2 haplotype, the association appears not to extend beyond the HLA-B locus.[14] A recent study of DR3-DQ2/DR4-DQ8 phenotype found that A1-cw7-B8 was actually lower than expected relative to other A-B types, indicating that risk associated genes are located between B8 and DR3. A*0101 appears to alter risk for type 1 diabetes but not Cw7-B8.[15] The type 1 diabetes example shows the inherent difficulty in the use of linkage analysis alone to cipher risk.
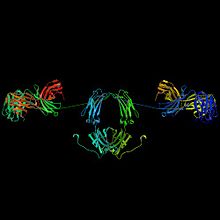
A1-B8 and allergic disease
[edit]In allergic disease A1,B8 were found to associate with allergic reactions in new-borns.[16] A1, B8 was found increased in children with bronchial asthma and low IgA.[17] However, some of this reaction can be attributed to the linkage of the HLA A1-B8-DR3-DQ2 haplotype to the IgA-less phenotype.[18] A firmer association was found with atopies.[19][20][21] A1,B8 where found more frequently in hay fever complicated by asthma or atopia relative to just hay fever.[22][23][24] Further asthmatic patients with negative skin tests tended toward higher A1,B8 serotypes.[25]
A1-B8 and infectious disease
[edit]
HIV
[edit]In the mid-1980s the association with A1-B8-DR3 and HIV progression appeared shortly after the discover of the virus.[26] A1-B8 associated with more rapid progression to seropositivity, and was strongly associated with a rapid decline in T4 cells and development of HIV-related symptoms within four years of infection.[27] The strongest associations were seen with A1-Cw7-B8 haplotype. C4 (complement 4) produces a null allele at on locus C4AQ. This locus in part of the HLA A1-B8-DR3-DQ2 haplotype (markers are A1, CW7, B8, BfS, C4AQ0, C4B1, DR3, DQ2) therefore one study concluded that C4AQ0 could explain the increased infectivity to HIV.[28] The haplotype was further linked to false-tumor splenomegaly, CD8 lymphocytosis, and high IgG.[29]
Viral hepatitis
[edit]An association was seen between viral hepatitis and HLA-A1.[30][31] Though, the association of A1 with autoimmune hepatitis with no anti-viral antibody was stronger than with chronic active hepatitis with anti-viral titer.[32] The association with viral hepatitis was subsequently demonstrated and patients with antinuclear antibodies were more likely to have A1-B8-DR3.[33] Currently studies point to association proximal the Cw*0702-B*0801 loci.
Frequencies
[edit]| [34] | Ireland | 14.4 | 1 | |
| Slovak | 11.9 | 1 | ||
| [35] | Northern Ireland | 11.5 | 1 | 1 |
| Swedish | 11.5 | 1 | ||
| [36] | Dutch Netherlands | 9.8 | ||
| Yugoslavian | 9.7 | 1 | ||
| British | 9.7 | 1 | ||
| Hungarian | 9.4 | 1 | ||
| [37] | CEPH France | 8.5 | 1 | 1 |
| [38] | German | 8.3 | 1 | |
| Czech | 7.8 | 1 | ||
| [39] | Swiss | 6.7 | 3 | |
| Belgium | 5.5 | 2 | ||
| Austria | 4.5 | 2 | ||
| [40] | Tuscan Italy | 4.3 | ||
| Ukraine | 4.3 | 3 | ||
| Italy | 4.2 | 1 | ||
| Basque | 4.2 | |||
| Portuguese | 4.2 | 2 | ||
| Polish | 4.0 | |||
| [41] | Uganda | 3.7 | 1 | |
| Uralic | 3.1 | 3 | ||
| Spanish | 2.8 | 4 | ||
| [1] | Romania | 2.8 | ||
| Albania | 2.5 | |||
| Greek | 2.3 | |||
| [2] Archived 2007-09-27 at the Wayback Machine | Northern Greece | 2.1 | ||
| [42] | Crete | 1.9 | ||
| [41] | Luo Kenya | 1.7 | 2 | |
| [41] | Nandi Kenya | 1.4 | 2 | |
| [43] | Oman Arabia | 1.4 | ||
| Japanese | 0.1 | |||
| 1Cw*0701 (Eur.), ²Cw*0704 (Afr.) | ||||
References
[edit]- ^ Clarke AK, Galbraith RM, Hamilton EB, Williams R (February 1978). "Rheumatic disorders in primary biliary cirrhosis". Ann. Rheum. Dis. 37 (1): 42–7. doi:10.1136/ard.37.1.42. PMC 1000187. PMID 305233.
- ^ Lada G, Gyódi E, Gláz E (1977). "HLA antigens in patients with adrenocortical hyperfunction". Acta Med Acad Sci Hung. 34 (4): 213–26. PMID 618054.
- ^ Ludwig H, Polymenidis Z, Granditsch G, Wick G (November 1973). "[Association of HL-A1 and HL-A8 with childhood celiac disease]". Z Immunitatsforsch Exp Klin Immunol (in German). 146 (2): 158–67. PMID 4282973.
- ^ Mackay IR, Morris PJ (October 1972). "Association of autoimmune active chronic hepatitis with HL-A1,8". Lancet. 2 (7781): 793–5. doi:10.1016/S0140-6736(72)92149-6. PMID 4116233.
- ^ Feltkamp TE, van den Berg-Loonen PN, Engelfriet CP, et al. (June 1974). "[HL-A typing and autoantibodies in patients with myasthenia gravis]". Ned Tijdschr Geneeskd (in Dutch). 118 (23): 895–6. PMID 4839444.
- ^ Hammarström L, Smith E, Möller E, Franksson C, Matell G, Von Reis G (August 1975). "Myasthenia gravis: studies on HL-A antigens and lymphocyte subpopulations in patients with myasthenia gravis". Clin. Exp. Immunol. 21 (2): 202–15. PMC 1538268. PMID 1081023.
- ^ Gazit E, Sartani A, Mizrachi Y, Ravid M (March 1977). "HLA antigen in Jewish patients with juvenile diabetes mellitus". Diabete Metab. 3 (1): 55–8. PMID 858444.
- ^ Vogten AJ, Shorter RG, Opelz G (June 1979). "HLA and cell-mediated immunity in HBsAg negative chronic active hepatitis". Gut. 20 (6): 523–5. doi:10.1136/gut.20.6.523. PMC 1412459. PMID 89064.
- ^ Ercilla G, Parés A, Arriaga F, et al. (November 1979). "Primary biliary cirrhosis associated with HLA-DRw3". Tissue Antigens. 14 (5): 449–52. doi:10.1111/j.1399-0039.1979.tb00874.x. PMID 12731577.
- ^ Robinson BN, Roberts DF, Mather BA, Nelson R, Rowlatt AS (October 1980). "Coeliac disease and HLA: a family study". European Journal of Immunogenetics. 7 (5): 381–91. doi:10.1111/j.1744-313X.1980.tb00732.x. PMID 7430678.
- ^ Tosi R, Vismara D, Tanigaki N, et al. (September 1983). "Evidence that celiac disease is primarily associated with a DC locus allelic specificity". Clin. Immunol. Immunopathol. 28 (3): 395–404. doi:10.1016/0090-1229(83)90106-X. PMID 6192959.
- ^ Goldberg MA, Arnett FC, Bias WB, Shulman LE (1976). "Histocompatibility antigens in systemic lupus erythematosus". Arthritis Rheum. 19 (2): 129–32. doi:10.1002/art.1780190201. PMID 1259797.
- ^ Parks CG, Pandey JP, Dooley MA, et al. (June 2004). "Genetic polymorphisms in tumor necrosis factor (TNF)-alpha and TNF-beta in a population-based study of systemic lupus erythematosus: associations and interaction with the interleukin-1alpha-889 C/T polymorphism" (PDF). Hum. Immunol. 65 (6): 622–31. doi:10.1016/j.humimm.2004.03.001. PMID 15219382.
- ^ Noble JA, Martin A, Valdes AM, et al. (2008). "Type 1 diabetes risk for human leukocyte antigen (HLA)-DR3 haplotypes depends on genotypic context: association of DPB1 and HLA class I loci among DR3- and DR4-matched Italian patients and controls". Hum. Immunol. 69 (4–5): 291–300. doi:10.1016/j.humimm.2008.02.003. PMC 2505335. PMID 18486765.
- ^ Noble J, Valdes A, Bugawan T, Apple R, Thomson G, Erlich H (2002). "The HLA class I A locus affects susceptibility to type 1 diabetes". Hum Immunol. 63 (8): 657–64. doi:10.1016/S0198-8859(02)00421-4. PMC 4049513. PMID 12121673.
- ^ Soothill JF, Stokes CR, Turner MW, Norman AP, Taylor B (July 1976). "Predisposing factors and the development of reaginic allergy in infancy". Clinical & Experimental Allergy. 6 (4): 305–19. doi:10.1111/j.1365-2222.1976.tb01911.x. PMID 963861.
- ^ Ostergaard PA, Eriksen J (August 1979). "Association between HLA-A1,B8 in children with extrinsic asthma and IgA deficiency". Eur. J. Pediatr. 131 (4): 263–70. doi:10.1007/BF00444347. PMID 477683.
- ^ Ambrus M, Hernádi E, Bajtai G (May 1977). "Prevalence of HLA-A1 and HLA-B8 antigens in selective IgA deficiency". Clin. Immunol. Immunopathol. 7 (3): 311–4. doi:10.1016/0090-1229(77)90062-9. PMID 872455.
- ^ Dasgupta A, Misri N, Bala S (1977). "Population and family studies to demonstrate Ir genes: HLA haplotype in atopic allergy". Monogr Allergy. 11: 75–9. PMID 327293.
- ^ MacKie RM, Dick HM (February 1979). "A study of HLA antigen distribution in families with atopic dermatitis". Allergy. 34 (1): 19–23. doi:10.1111/j.1398-9995.1979.tb01996.x. PMID 453484.
- ^ Heikkilä M, Koistinen J, Lohman M, Koskimies S (May 1984). "Increased frequency of HLA-A1 and -B8 in association with total lack, but not with deficiency of serum IgA". Tissue Antigens. 23 (5): 280–3. doi:10.1111/j.1399-0039.1984.tb00046.x. PMID 6611606.
- ^ Turner MW, Brostoff TJ, Wells RS, Stokes CR, Soothill JF (January 1977). "HLA in eczema and hay fever". Clin. Exp. Immunol. 27 (1): 43–7. PMC 1540890. PMID 849649.
- ^ Jeannet M, Girard JP, Varonier HS, Mirimanoff P, Joye P (1977). "HLA antigens in grass pollinosis". Monogr Allergy. 11: 69–73. PMID 876128.
- ^ Oehling A, Baena-Cagnani CE, Sanz ML, Crisci CD (1979). "HLA and pollinosis". Allergol Immunopathol (Madr). 7 (6): 423–6. PMID 539526.
- ^ Morris MJ, Vaughan H, Lane DJ, Morris PJ (1977). "HLA in asthma". Monogr Allergy. 11: 30–4. PMID 876121.
- ^ Steel CM, Ludlam CA, Beatson D, et al. (May 1988). "HLA haplotype A1 B8 DR3 as a risk factor for HIV-related disease". Lancet. 1 (8596): 1185–8. doi:10.1016/S0140-6736(88)92009-0. PMID 2897006.
- ^ Kaslow RA, Duquesnoy R, VanRaden M, et al. (April 1990). "A1, Cw7, B8, DR3 HLA antigen combination associated with rapid decline of T-helper lymphocytes in HIV-1 infection. A report from the Multicenter AIDS Cohort Study". Lancet. 335 (8695): 927–30. doi:10.1016/0140-6736(90)90995-H. PMID 1970024.
- ^ Cameron PU, Mallal SA, French MA, Dawkins RL (December 1990). "Major histocompatibility complex genes influence the outcome of HIV infection. Ancestral haplotypes with C4 null alleles explain diverse HLA associations". Hum. Immunol. 29 (4): 282–95. doi:10.1016/0198-8859(90)90042-N. PMID 1981061.
- ^ Oksenhendler E, Autran B, Gorochov G, D'Agay MF, Seligmann M, Clauvel JP (July 1992). "CD8 lymphocytosis and pseudotumoral splenomegaly in HIV infection". Lancet. 340 (8813): 207–8. doi:10.1016/0140-6736(92)90471-E. PMID 1353138.
- ^ Hug G (May 1976). "Genetic factors and autoimmunity in viral hepatitis". Am. J. Clin. Pathol. 65 (5 Suppl): 870–5. PMID 218441.
- ^ Descamps B, Jungers P, Naret C, Degott C, Zingraff J, Bach JF (1977). "HLA-A1, B8-phenotype association and HBs antigenemia evolution in 440 hemodialyzed patients". Digestion. 15 (4): 271–7. doi:10.1159/000198012. PMID 67978.
- ^ Freudenberg J, Baumann H, Arnold W, Berger J, Büschenfelde KH (1977). "HLA in different forms of chronic active hepatitis. A comparison between adult patients and children". Digestion. 15 (4): 260–70. doi:10.1159/000198011. PMID 863130.
- ^ Czaja AJ, Carpenter HA, Santrach PJ, Moore SB (January 1995). "Immunologic features and HLA associations in chronic viral hepatitis". Gastroenterology. 108 (1): 157–64. doi:10.1016/0016-5085(95)90020-9. PMID 7806037.
- ^ Finch T, Lawlor E, Borton M, Barnes C, McNamara S, O'Riordan J, McCann S, Darke C (1997). "Distribution of HLA-A, B and DR genes and haplotypes in the Irish population". Exp Clin Immunogenet. 14 (4): 250–63. PMID 9523161.
- ^ Middleton D, Williams F, Hamill M, Meenagh A (2000). "Frequency of HLA-B alleles in a Caucasoid population determined by a two-stage PCR-SSOP typing strategy". Hum Immunol. 61 (12): 1285–97. doi:10.1016/S0198-8859(00)00186-5. PMID 11163085.
- ^ Schipper R, Schreuder G, D'Amaro J, Oudshoorn M (1996). "HLA gene and haplotype frequencies in Dutch blood donors". Tissue Antigens. 48 (5): 562–74. doi:10.1111/j.1399-0039.1996.tb02670.x. PMID 8988539.
- ^ Bugawan T, Klitz W, Blair A, Erlich H (2000). "High-resolution HLA class I typing in the CEPH families: analysis of linkage disequilibrium among HLA loci". Tissue Antigens. 56 (5): 392–404. doi:10.1034/j.1399-0039.2000.560502.x. PMID 11144287.
- ^ Müller C, Ehninger G, Goldmann S (2003). "Gene and haplotype frequencies for the loci hLA-A, hLA-B, and hLA-DR based on over 13,000 German blood donors". Hum Immunol. 64 (1): 137–51. doi:10.1016/S0198-8859(02)00706-1. PMID 12507825.
- ^ Grundschober C, Sanchez-Mazas A, Excoffier L, Langaney A, Jeannet M, Tiercy J (1994). "HLA-DPB1 DNA polymorphism in the Swiss population: linkage disequilibrium with other HLA loci and population genetic affinities". European Journal of Immunogenetics. 21 (3): 143–57. doi:10.1111/j.1744-313X.1994.tb00186.x. PMID 9098428.
- ^ Marroni F, Curcio M, Fornaciari S, Lapi S, Mariotti M, Scatena F, Presciuttini S (2004). "Microgeographic variation of HLA-A, -B, and -DR haplotype frequencies in Tuscany, Italy: implications for recruitment of bone marrow donors". Tissue Antigens. 64 (4): 478–85. doi:10.1111/j.1399-0039.2004.00292.x. PMID 15361126.
- ^ a b c Cao K, Moormann A, Lyke K, Masaberg C, Sumba O, Doumbo O, Koech D, Lancaster A, Nelson M, Meyer D, Single R, Hartzman R, Plowe C, Kazura J, Mann D, Sztein M, Thomson G, Fernández-Viña M (2004). "Differentiation between African populations is evidenced by the diversity of alleles and haplotypes of HLA class I loci". Tissue Antigens. 63 (4): 293–325. doi:10.1111/j.0001-2815.2004.00192.x. PMID 15009803.
- ^ Arnaiz-Villena A, Iliakis P, González-Hevilla M, Longás J, Gómez-Casado E, Sfyridaki K, Trapaga J, Silvera-Redondo C, Matsouka C, Martínez-Laso J (1999). "The origin of Cretan populations as determined by characterization of HLA alleles". Tissue Antigens. 53 (3): 213–26. doi:10.1034/j.1399-0039.1999.530301.x. PMID 10203014.
- ^ Williams F, Meenagh A, Darke C, Acosta A, Daar A, Gorodezky C, Hammond M, Nascimento E, Middleton D (2001). "Analysis of the distribution of HLA-B alleles in populations from five continents". Hum Immunol. 62 (6): 645–50. doi:10.1016/S0198-8859(01)00247-6. PMID 11390040.
