Cofactor engineering
Cofactor engineering, a subset of metabolic engineering, is defined as the manipulation of the use of cofactors in an organism’s metabolic pathways. In cofactor engineering, the concentrations of cofactors are changed in order to maximize or minimize metabolic fluxes. This type of engineering can be used to optimize the production of a metabolite product or to increase the efficiency of a metabolic network. The use of engineering single celled organisms to create lucrative chemicals from cheap raw materials is growing, and cofactor engineering can play a crucial role in maximizing production. The field has gained more popularity in the past decade and has several practical applications in chemical manufacturing, bioengineering and pharmaceutical industries.[1]
Cofactors are non-protein compounds that bind to proteins and are required for the proteins normal catalytic functionality. Cofactors can be considered “helper molecules” in biological activity, and often affect the functionality of enzymes. Cofactors can be both organic and inorganic compounds. Some examples of inorganic cofactors are iron or magnesium, and some examples of organic cofactors include ATP or coenzyme A. Organic cofactors are more specifically known as coenzymes, and many enzymes require the addition of coenzymes to assume normal catalytic function in a metabolic reaction. The coenzymes bind to the active site of an enzyme to promote catalysis. By engineering cofactors and coenzymes, a naturally occurring metabolic reaction can be manipulated to optimize the output of a metabolic network.[2][3][4]
Background
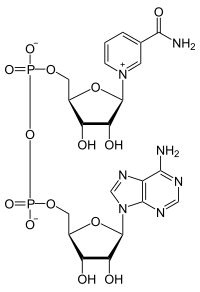
[edit]Cofactors were discovered by Arthur Harden and William Young in 1906, when they found that the rate of alcoholic fermentation in unboiled yeast extracts increased when boiled yeast extract was added.[5] A few years after, Hans von Euler-Chelpin identified the cofactor in the boiled extract as NAD+. Other cofactors, such as ATP and coenzyme A, were discovered later in the 1900s. The mechanism of cofactor activity was discovered when, Otto Heinrich Warburg determined in 1936 that NAD+ functioned as an electron acceptor. Well after these initial discoveries, scientists began to realize that the manipulation of cofactor concentrations could be used as tools for the improvement of metabolic pathways.[1]
An important group of organic cofactors is the family of molecules referred to as vitamins. Vitamin B12 (cobalamin), for example, plays a crucial role in the human body, while coenzyme B12, its derivative, is found in the metabolisms of every type of cell in our bodies. Its presence affects the synthesis and regulation of cellular DNA as well as taking part in fatty acid synthesis and energy production. Cofactors are required by many important metabolic pathways, and it is possible for the concentrations of a single type of cofactor to affect the fluxes of many different pathways .
Minerals and metallic ions that organisms uptake through their diet provide prime examples of inorganic cofactors. For instance Zn2+ is needed to assist the enzyme carbonic anhydrase as it converts carbon dioxide and water to bicarbonate and protons. A widely recognized mineral that acts as a cofactor is iron, which is essential for the proper function of hemoglobin, the oxygen transporting protein found in red blood cells. This example in particular highlights the importance of cofactors in animal metabolism.
Significance
[edit]Cofactor engineering is significant in the manipulation of metabolic pathways. A metabolic pathway is a series of chemical reactions that occur in an organism. Metabolic engineering is the subject of altering the fluxes within a metabolic pathway. In metabolic engineering, a metabolic pathway can be directly altered by changing the functionality of the enzymes involved in the pathway. Cofactor engineering, offers a distinct approach, and some advantages, to altering a metabolic pathway. Instead of changing the enzymes used in a pathway, the cofactors can be changed. This may give metabolic engineers an advantage due to certain properties of cofactors and how they can be modified.
Metabolic pathways can be used by metabolic engineers to create a desired product. By modifying the types of cofactors used and the times at which they are used, the outcome of the metabolic network can change. To create a greater production of a product, metabolic engineers have the ability to supply the network with whichever cofactor is best suited for that specific process. This leads to the optimization of networks to give a higher production of desired products. Also, changing the cofactors used in a network may be an ingenious solution to a complicated problem. A network that is present in the cell, but is often unused, may have a desirable product. Instead of engineering a completely new set of pathways to produce the product, cofactor engineering can be applied. By replacing enzymes to use cofactors readily available in a cell, the typically unused network is no longer cofactor-limited, and production may be increased.
In addition to modifying the yield of metabolic networks, changing the cofactors used in a network can reduce operation costs when trying to form a desired product. NADH and NADPH are two extremely common cellular cofactors, differing only by the presence of a phosphate group. However, this phosphate group makes NADPH much less stable than NADH, and therefore more expensive to synthesize. Thus, it is advantageous to try and use NADH is some cellular networks because it is often cheaper, more readily available, and accomplishes the same task as NADPH.
Tools and processes
[edit]Cofactor engineering most often deals with the manipulation of microorganisms such as Saccharomyces cerevisiae and Escherichia coli, and as such requires the use of recombinant DNA techniques. These techniques utilize small circular segments of DNA called plasmids, which can be introduced and incorporated by microorganisms such as Escherichia coli. These plasmids are specifically designed in labs to be easily incorporated, and affect the expression of various protein, metabolites and enzymes. For instance, a particular plasmid may cause a change in an enzyme's amino acid sequence, which could increase its affinity for a particular substrate.
Microorganisms require a medium to grow in, and one commonly used for cultures of Escherichia coli is Luria-Bertani (LB) broth. This medium is often supplemented with glucose and will often contain additional molecules designed to facilitate optimal culture growth. Pre-cultures may then be grown in shake flasks. These are simply plugged Erlenmeyer flasks which are left on an orbital shaker machine, which revolves at very high RPM. This process aerates the culture, which is necessary for optimal growth. Once the pre-cultures are ready, the plasmids needed by specific experiments are added to each culture separately, and then each culture is transferred to a bio-reactor. Bio-reactors are systems which allow cultures to grow in a controlled environment. This leaves the introduced plasmids as the only independent variable. The required temperature, pH, metabolite concentrations, and various other environmental factors can be maintained by the bio-reactor ensuring identical growth conditions for each culture.
Once samples are allowed to grow in the reactor for a specified period, they can be removed and studied to determine whether the intended alterations to the organism are evident. Since cofactor engineering most often deals with metabolic pathways, these organisms are often studied but introducing specific tagged fluorescent metabolites and documenting their progression through various pathways. In other cases results are more obvious and easily observable, such as with the decreased ethanol production of yeast referred to below. [3][4]
Applications
[edit]Changing an Enzyme's Cofactor From NADPH to NADH
[edit]Biocatalysts are required for the production of chiral building blocks needed in pharmaceuticals and other chemicals used by society. Many such biocatalysts require NADPH as a cofactor. NADPH, a cofactor quite similar to NADH, is both more expensive and less stable than its counterpart NADH. For these reasons, manufacturers would prefer that the biocatalysts they use in their production lines accept NADH over NADPH. Cofactor engineering has recently been successful in altering enzymes to prefer NADH as a cofactor instead of NADPH. In 2010, a group of scientists performed cofactor engineering on the enzyme Gre2p, an NADPH-preferring dehydrogenase found in Saccharomyces cerevisiae. Gre2p reduces the compound diketone 2,5-hexanedione into the chiral building blocks (5S)-hydroxy-2-hexanone and (2S,5S)-hexanediol. The scientists determined that Asn9 (Asparagine, position 9) was an important amino acid the active site of Gre2p. Specifically, Asn9 binds to the 3’-hydroxyl group and the 2’-oxygen atom of adenyl ribose moiety. Through direct mutagenesis, the scientists exchanged the Asn9 for both Asp (Aspartic Acid) and Glu (Glutamic Acid). This change caused Gre2p to have a decreased dependency on NADPH, and an increased affinity for NADH. This resulted in increased Gre2p activity when using NADH. It was observed that substituting Asn9 with Glu produced a greater effect than changing Asn9 to Asp. Asn contains a polar uncharged side chain, while both Asp and Glu contain a polar charged side chain. The increased effect of Glu is caused by the extra carbon in its side chain that brings it closer to adenyl ribose moiety. This allows for stronger hydrogen bonding between the 2’- and 3’- ribose hydroxyl groups and the side chain carboxyl group. The maximum velocity of the reaction doubled, while using NADH, when Asn9 was substituted with Glu. With these results, the scientists successfully engineered Gre2p to prefer NADH over NADPH and increased the speed of 2,5-hexanedione reduction. This will allow chemical companies to decrease their manufacturing costs by using NADH instead of NADPH at least for this particular reduction.[6]
Changing a Network's Cofactor Preference
[edit]An alternative example of changing an enzyme’s preference for cofactors is to change NADH dependent reaction to NADPH dependent reactions. In this example, the enzymes themselves are not changed, but instead different enzymes are selected that accomplish the same reaction with the use of a different cofactor. An engineered pathway was created to make 1-butanol from Acetyl-CoA by changing enzymes in the metabolic pathway of S. elongatus. The Clostridium genus is known to produce 1-butanol, providing a pathway that could be inserted in S. elongatus. This pathway synthesizes 1-butanol using the reverse β-oxidation pathway. The enzymes involved in this newly engineered pathway were NADH specific, which was problematic for replicating the pathway in S. elongatus as cyanobacteria produce much more NADPH than NADH.
The research group then identified enzymes that utilize NADPH or both NADPH and NADH by bioprospecting. Acetoacetyl-CoA reductase (PhaB) was found to be a suitable replacement for hydroxybutyric dehydrogenase (Hbd). To replace AdhE2, the researchers found that NADP-dependent alcohol dehydrogenase (YqhD) from E. coli to be effective for the pathway. Furthermore, the researchers needed a dehydrogenase to replace the aldehyde dehydrogenase capacity of AdhE2. CoA-acylating butyraldehyde dehydrogenase (Bldh) from C. saccharoperbutylacetonicum was found to be a good suit. Together, PhaB, Bldh, YqhD can replace Hbd and AdhE2, respectively, to change the cofactor preference of 3-ketobutyryl-CoA reduction from using NADH to using NADPH. The authors then constructed various combinations of the different enzymes (of those found in the reverse oxidation pathway and the NADPH utilizing enzymes) by overexpressing different genes in cultures of S. elongatus PCC 7942. In order to do so, they constructed plasmids containing the genes corresponding to the enzymes and combined them into the genome of S. elongates. After enzyme assays, the strain of cyanobacteria expressing the NADPH utilizing enzymes produced the greatest amount of 1-butanol (29.9 mg/L), exceeding that of strains that did not consist of the NADPH utilizing enzymes by four times. Overall, 1-butanol was produced in S. elongatus using a pathway from another organism. This pathway was modified in order to match the preferred reducing cofactor for the cyanobacteria.[7]

Modifying Metabolite Flux with Cofactor Equilibrium
[edit]In cofactor engineering, a metabolic pathway is altered by changing the concentrations of specific cofactors that are produced either in that particular pathway or in a separate pathway. For example, an hypothetical organism could have two arbitrary pathways called A and B where some enzymes in both A and B utilize the same cofactors. If scientists wanted to decrease the output of pathway A, they may first consider directly engineering the enzymes involved in A, perhaps to decrease a particular active site's affinity for its substrate. In some cases however, the enzymes in A may be difficult to engineer for various reasons, or it may be impossible to engineer them without dangerously affecting some third metabolic pathway C, which utilizes the same enzymes. As a separate option, scientists could increase the flux of B, which may be easier to engineer. This in turn could "tie up" the cofactors needed by A, which would slow enzymatic activity, decreasing output in A. This is one hypothetical example of how cofactor engineering can be used, but there are many other unique cases where scientists use cofactors as a way of altering metabolic pathways. A major advantage to cofactor engineering is that scientists can use it to successfully alter metabolic pathways that are difficult to engineer by means of ordinary metabolic engineering. This is achieved by targeting more easily engineered enzymes in separate pathways, which use the same cofactors. Since many cofactors are used by different enzymes in multiple pathways, cofactor engineering may be an efficient, cost effective alternative to current methods of metabolic engineering.[8]
Yeast are commonly used in the beer and wine industry because they are capable of efficiently producing ethanol through the metabolic pathway fermentation in the absence of oxygen. Fermentation requires the enzyme glycerol-3-phosphate dehydrogenase (GPDH) which depends on the cofactor NADH. This pathway involves the conversion of glucose to both ethanol and glycerol, both of which use NADH as a cofactor. Scientists engineered Saccharomyces cerevisiae to overproduce GPDH, which shifted the cells metabolic flux away from ethanol and toward glycerol, by limiting NADH availability in the ethanol production portion of the pathway. The opposite effect was achieved by influencing a separate pathway in the cell, the Glutamate Synthesis pathway. Inactivating the expression of the enzyme glutamate dehydrogenase, which is NADPH dependent, and over expressing the enzymes glutamine synthetase and glutamate synthetase, which rely on NADH as a cofactor shifted the cofactor balance in glutamate synthesis pathway. The pathway is now dependent on NADH rather than NADPH, which decreases NADH availability in the fermentation pathway. This in turn causes increased ethanol production and decreased glycerol production. This method of manipulating metabolic fluxes could be visualized much like global fuel markets, where the increased production of ethanol for use in the automotive industry would decrease its availability in the food industry. Essentially, producing more engines which run on ethanol could result in decreased consumption of processed sweets, which contain high fructose corn syrup. This engineering of cofactors is applicable to the beer and wine industry since it allows for the regulation of ethanol levels in alcoholic beverages. Advancements in the wine industry have caused a steady increase in ethanol content, so winemakers in particular would be interested in the possibility of reducing the ethanol levels of some of their wines.[3]
Citric acid cycle
[edit]Coenzyme A (CoA) and acetyl-CoA are two intermediate metabolites, most notably found in the Citric Acid Cycle, which participate in over 100 different reactions in the metabolism of microorganisms. Recent experiments have shown that over expression of the enzyme pantothenate kinase and supplementation of pantothenic acid in the CoA biosynthesis pathway have allowed adjustments of both CoA and acetyl-CoA fluxes. This increased concentration of cofactors resulted in an increased carbon flux in the isoamyl acetate synthesis pathway, increase the production efficiency of isoamyl acetate. Isoamyl acetate is used industrially for artificial flavoring and for testing the effectiveness of respirators. In addition to the production of isoamyl acetate, the manipulation of CoA biosynthesis during the pyruvate hydrogenase reaction also causes an increase in the production of both succinate and lycopene, each of which have beneficial effects on the human body. An increase in succinate concentration, which is used as a catalyst, may lead to an increase in the speed of the Citric Acid Cycle, and consequently an individual's metabolism. Increasing lycopene concentrations, has been shown to decrease the risk of prostate cancer. The potential rewards of repeating such feat of cofactor engineering and successfully incorporating them into industry practices are innumerable.[4][9][10]
Paper manufacturing
[edit]Many important industrial enzymes use cofactors to catalyze reactions. By using cofactors to manipulate metabolic pathways, it is possible to reduce material cost, eliminate steps in production, reduce production time, decrease pollution, and increase overall production efficiency. One case that demonstrates several of these manufacturing benefits involves the genetic engineering of aspen trees. In the paper production process, manufacturing plants must break down lignin, a biochemical compound that gives a tree trunk its stiffness, in order to form the pulp used throughout the rest of production. The chemical pulping process requires the manufacturing plant to use a significant amount of energy, as well as many expensive and toxic chemicals. A group of genetic engineers, through cofactor engineering, engineered a genetically superior aspen tree that produced less lignin. These genetically engineered trees have allowed for paper mills to reduce their costs, pollution, and manufacturing time.[1][11]
Other examples
[edit]| Organism | What Is Changed | What It Means |
|---|---|---|
| Mycobacterium smegmatis | Coenzyme F420 | Could Disable a Cofactor that Leads to Drug Resistant Tuberculosis[12] |
| Cyanobacteria | Iron-Molybdenum Cofactor | Production of Hydrogen Gas For Energy[13] |
| Kingdom Archea | Molybdenum Nitrogenase Cofactor | Improvement in the Efficiency of Nitrogen Fixation[14] |
| Thermoanaerobacteria mathranii | glycerol dehydrogenase | Make Thermophillic Bacteria Produce Ethanol More Efficiently [15] |
A short description of other significant examples where cofactor engineering has been used.
Notes
[edit]- ^ a b c Raab, Michael; Keith Tyro; Gregory Stephanopoulos (2005). "Metabolic Engineering". Advances in Biochemical Engineering/Biotechnology. 100 (763): 1–17. doi:10.1007/b136411. PMID 16270654. S2CID 21169566.
- ^ Pollak, N; C Dolle; M Ziegler (2006). "The power to reduce: pyridine nucleotides - small molecules with a multitude of functions". Biochemical Journal. 402 (2): 205–218. doi:10.1042/BJ20061638. PMC 1798440. PMID 17295611.
- ^ a b c Heux, Stephanie; Remy Cachon; Sylvie Dequin (2006). "Cofactor engineering in Saccharomyces cerevisiae: Expression of a H2O-forming NADH oxidase and impact on redox metabolism". Metabolic Engineering. 8 (4): 303–314. doi:10.1016/j.ymben.2005.12.003. PMID 16473032.
- ^ a b c Vadali, Ravishankar; George Bennett (April 2004). "Cofactor engineering of intracellular CoA/acetyl-CoA and its effect on metabolic flux redistribution in Escherichia coli". Metabolic Engineering. 6 (2): 133–139. doi:10.1016/j.ymben.2004.02.001. PMID 15113566.
- ^ Arthur Harden and William John Young. "The Alcoholic Ferment of Yeast-Juice". Proceedings of the Royal Society of London. Series B, Containing Papers of a Biological Character, Vol. 77, No. 519 (Apr. 12, 1906), pp. 405-420 JSTOR
- ^ Katzberg, Michael; Nadia Skorupa-Parachin; Marie-Francoise Gorwa-Grauslund; Martin Bertau (2010). "Engineering Cofactor Preference of Ketone Reducing Biocatalysts: A Mutagenesis Study on a γ-Diketone Reductase from the Yeast Saccharomyces cerevisiae". International Journal of Molecular Sciences. 11 (4): 1735–1758. doi:10.3390/ijms11041735. PMC 2871135. PMID 20480039.
- ^ Ethan Lan; James Liao (17 April 2012). "ATP Drives Direct Photosynthetic Production of 1-butanol in Cyanobacteria". Proceedings of the National Academy of Sciences. 109 (16): 6018–6023. Bibcode:2012PNAS..109.6018L. doi:10.1073/pnas.1200074109. PMC 3341080. PMID 22474341.
- ^ San, Ka-Yiu; George N. Bennett; Susana J. Berrıos-Rivera; Ravi V. Vadali; Yea-Tyng Yang; Emily Horton; Fred B. Rudolph; Berna Sariyar; Kimathi Blackwood (April 2002). "Metabolic Engineering through Cofactor Manipulation and Its Effects on Metabolic Flux Redistribution in Escherichia Coli". Metabolic Engineering. 4 (2): 182–193. doi:10.1006/mben.2001.0220. PMID 12009797.[dead link]
- ^ Giovannucci, Edward; Alberto Ascherio; Eric Rimm; Meir Stampfer; Graham Colditz; Walter Willett (May 1995). "Intake of Carotenoids and Retino in Relation to Risk of Prostate Cancer". Journal of the National Cancer Institute. 87 (23): 1767–1776. doi:10.1093/jnci/87.23.1767. PMID 7473833.
- ^ Potera, Carol (December 2005). "Making Succinate More Successful". Environmental Health Perspectives. 113 (12): A832–A835. doi:10.1289/ehp.113-a832. PMC 1314946. PMID 16330341.
- ^ Horvath, Laszlo; Ilona Peszlen; Perry Peralta; Bohumil Kasal; Laigeng Li (2010). "Mechanical Properties of Genetically Engineered Young Aspen with Modified Lignin Content and/or Structure". Wood and Fiber Science. 42.
- ^ Bashiri, G; Rehan, AM; Greenwood, DR; Dickson, JM; Baker, EN (2010). "Metabolic engineering of cofactor F420 production in Mycobacterium smegmatis". PLOS ONE. 5 (12): e15803. Bibcode:2010PLoSO...515803B. doi:10.1371/journal.pone.0015803. PMC 3012119. PMID 21209917.
- ^ Masukawa, H; Inoue, K; Sakurai, H; Wolk, CP; Hausinger, RP. (2010). "Site-directed mutagenesis of the Anabaena sp. strain PCC 7120 nitrogenase active site to increase photobiological hydrogen production". Applied and Environmental Microbiology. 76 (20): 6741–50. Bibcode:2010ApEnM..76.6741M. doi:10.1128/aem.01056-10. PMC 2953041. PMID 20709836.
- ^ Hernandez, JA; Curatti, L; Aznar, CP; Perova, Z; Britt, RD; Rubio, LM. (2008). "Metal trafficking for nitrogen fixation: NifQ donates molybdenum to NifEN/NifH for the biosynthesis of the nitrogenase FeMo-cofactor". Proceedings of the National Academy of Sciences of the United States of America. 105 (33): 11679–8. doi:10.1073/pnas.0803576105. PMC 2575292. PMID 18697927.
- ^ Yao, S; Mikkelsen, MJ. (2010). "Metabolic engineering to improve ethanol production in Thermoanaerobacter mathranii". Applied Microbiology and Biotechnology. 88 (1): 199–208. doi:10.1007/s00253-010-2703-3. PMID 20552355. S2CID 25646005.
