Cathepsin C
| Cathepsin C exclusion domain | |||||||||
|---|---|---|---|---|---|---|---|---|---|
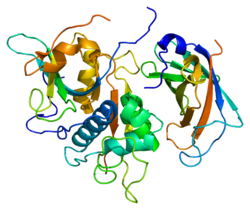
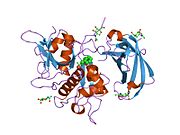
 re-determination of the native structure of human dipeptidyl peptidase i (cathepsin c) | |||||||||
| Identifiers | |||||||||
| Symbol | CathepsinC_exc | ||||||||
| Pfam | PF08773 | ||||||||
| InterPro | IPR014882 | ||||||||
| SCOP2 | 1k3b / SCOPe / SUPFAM | ||||||||
| |||||||||
Cathepsin C (CTSC) also known as dipeptidyl peptidase I (DPP-I) is a lysosomal exo-cysteine protease belonging to the peptidase C1 protein family, a subgroup of the cysteine cathepsins. In humans, it is encoded by the CTSC gene.[5][6]
Function
[edit]Cathepsin C appears to be a central coordinator for activation of many serine proteases in immune/inflammatory cells.
Cathepsin C catalyses excision of dipeptides from the N-terminus of protein and peptide substrates, except if (i) the amino group of the N-terminus is blocked, (ii) the site of cleavage is on either side of a proline residue, (iii) the N-terminal residue is lysine or arginine, or (iv) the structure of the peptide or protein prevents further digestion from the N-terminus.
Structure
[edit]The cDNAs encoding rat, human, murine, bovine, dog and two Schistosome cathepsin Cs have been cloned and sequenced and show that the enzyme is highly conserved.[7] The human and rat cathepsin C cDNAs encode precursors (prepro-cathepsin C) comprising signal peptides of 24 residues, pro-regions of 205 (rat cathepsin C) or 206 (human cathepsin C) residues and catalytic domains of 233 residues which contain the catalytic residues and are 30-40% identical to the mature amino acid sequences of papain and a number of other cathepsins including cathepsins, B, H, K, L, and S.[8]
The translated prepro-cathepsin C is processed into the mature form by at least four cleavages of the polypeptide chain. The signal peptide is removed during translocation or secretion of the pro-enzyme (pro-cathepsin C) and a large N-terminal proregion fragment (also known as the exclusion domain),[9] which is retained in the mature enzyme, is separated from the catalytic domain by excision of a minor C-terminal part of the pro-region, called the activation peptide. A heavy chain of about 164 residues and a light chain of about 69 residues are generated by cleavage of the catalytic domain.
Unlike the other members of the papain family, mature cathepsin C consists of four subunits, each composed of the N-terminal proregion fragment, the heavy chain and the light chain. Both the pro-region fragment and the heavy chain are glycosylated.
Clinical significance
[edit]Defects in the encoded protein have been shown to be a cause of Papillon-Lefevre disease,[10][11] an autosomal recessive disorder characterized by palmoplantar keratosis and periodontitis.
Cathepsin C functions as a key enzyme in the activation of granule serine peptidases in inflammatory cells, such as elastase and cathepsin G in neutrophils cells and chymase and tryptase in mast cells. In many inflammatory diseases, such as rheumatoid arthritis, chronic obstructive pulmonary disease (COPD), inflammatory bowel disease, asthma, sepsis, and cystic fibrosis, a significant portion of the pathogenesis is caused by increased activity of some of these inflammatory proteases. Once activated by cathepsin C, the proteases are capable of degrading various extracellular matrix components, which can lead to tissue damage and chronic inflammation.
References
[edit]- ^ a b c GRCh38: Ensembl release 89: ENSG00000109861 – Ensembl, May 2017
- ^ a b c GRCm38: Ensembl release 89: ENSMUSG00000030560 – Ensembl, May 2017
- ^ "Human PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ "Mouse PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ "Entrez Gene: CTSC cathepsin C".
- ^ Paris A, Strukelj B, Pungercar J, Renko M, Dolenc I, Turk V (Aug 1995). "Molecular cloning and sequence analysis of human preprocathepsin C". FEBS Letters. 369 (2–3): 326–30. Bibcode:1995FEBSL.369..326P. doi:10.1016/0014-5793(95)00777-7. PMID 7649281. S2CID 45737414.
- ^ Hola-Jamriska L, Tort JF, Dalton JP, Day SR, Fan J, Aaskov J, Brindley PJ (Aug 1998). "Cathepsin C from Schistosoma japonicum--cDNA encoding the preproenzyme and its phylogenetic relationships". European Journal of Biochemistry. 255 (3): 527–34. doi:10.1046/j.1432-1327.1998.2550527.x. PMID 9738890.
- ^ Kominami E, Ishido K, Muno D, Sato N (Jul 1992). "The primary structure and tissue distribution of cathepsin C". Biological Chemistry Hoppe-Seyler. 373 (7): 367–73. doi:10.1515/bchm3.1992.373.2.367. PMID 1515062.
- ^ Turk D, Janjić V, Stern I, Podobnik M, Lamba D, Dahl SW, Lauritzen C, Pedersen J, Turk V, Turk B (Dec 2001). "Structure of human dipeptidyl peptidase I (cathepsin C): exclusion domain added to an endopeptidase framework creates the machine for activation of granular serine proteases". The EMBO Journal. 20 (23): 6570–82. doi:10.1093/emboj/20.23.6570. PMC 125750. PMID 11726493.
- ^ Wani AA, Devkar N, Patole MS, Shouche YS (Feb 2006). "Description of two new cathepsin C gene mutations in patients with Papillon-Lefèvre syndrome". Journal of Periodontology. 77 (2): 233–7. doi:10.1902/jop.2006.050124. PMID 16460249.
- ^ Meade JL, de Wynter EA, Brett P, Sharif SM, Woods CG, Markham AF, Cook GP (May 2006). "A family with Papillon-Lefevre syndrome reveals a requirement for cathepsin C in granzyme B activation and NK cell cytolytic activity". Blood. 107 (9): 3665–8. doi:10.1182/blood-2005-03-1140. PMID 16410452.
Further reading
[edit]- McGuire MJ, Lipsky PE, Thiele DL (Jun 1992). "Purification and characterization of dipeptidyl peptidase I from human spleen". Archives of Biochemistry and Biophysics. 295 (2): 280–8. doi:10.1016/0003-9861(92)90519-3. PMID 1586157.
- Paris A, Strukelj B, Pungercar J, Renko M, Dolenc I, Turk V (Aug 1995). "Molecular cloning and sequence analysis of human preprocathepsin C". FEBS Letters. 369 (2–3): 326–30. Bibcode:1995FEBSL.369..326P. doi:10.1016/0014-5793(95)00777-7. PMID 7649281. S2CID 45737414.
- Dolenc I, Turk B, Pungercic G, Ritonja A, Turk V (Sep 1995). "Oligomeric structure and substrate induced inhibition of human cathepsin C". The Journal of Biological Chemistry. 270 (37): 21626–31. doi:10.1074/jbc.270.37.21626. PMID 7665576.
- Maruyama K, Sugano S (Jan 1994). "Oligo-capping: a simple method to replace the cap structure of eukaryotic mRNAs with oligoribonucleotides". Gene. 138 (1–2): 171–4. doi:10.1016/0378-1119(94)90802-8. PMID 8125298.
- Rao NV, Rao GV, Hoidal JR (Apr 1997). "Human dipeptidyl-peptidase I. Gene characterization, localization, and expression". The Journal of Biological Chemistry. 272 (15): 10260–5. doi:10.1074/jbc.272.15.10260. PMID 9092576.
- Fischer J, Blanchet-Bardon C, Prud'homme JF, Pavek S, Steijlen PM, Dubertret L, Weissenbach J (1997). "Mapping of Papillon-Lefevre syndrome to the chromosome 11q14 region". European Journal of Human Genetics. 5 (3): 156–60. doi:10.1159/000484751. hdl:2066/24363. PMID 9272739.
- Suzuki Y, Yoshitomo-Nakagawa K, Maruyama K, Suyama A, Sugano S (Oct 1997). "Construction and characterization of a full length-enriched and a 5'-end-enriched cDNA library". Gene. 200 (1–2): 149–56. doi:10.1016/S0378-1119(97)00411-3. PMID 9373149.
- Cigić B, Krizaj I, Kralj B, Turk V, Pain RH (Jan 1998). "Stoichiometry and heterogeneity of the pro-region chain in tetrameric human cathepsin C". Biochimica et Biophysica Acta (BBA) - Protein Structure and Molecular Enzymology. 1382 (1): 143–50. doi:10.1016/S0167-4838(97)00173-8. PMID 9507095.
- Toomes C, James J, Wood AJ, Wu CL, McCormick D, Lench N, Hewitt C, Moynihan L, Roberts E, Woods CG, Markham A, Wong M, Widmer R, Ghaffar KA, Pemberton M, Hussein IR, Temtamy SA, Davies R, Read AP, Sloan P, Dixon MJ, Thakker NS (Dec 1999). "Loss-of-function mutations in the cathepsin C gene result in periodontal disease and palmoplantar keratosis". Nature Genetics. 23 (4): 421–4. doi:10.1038/70525. PMID 10581027. S2CID 11433166.
- Hart TC, Hart PS, Bowden DW, Michalec MD, Callison SA, Walker SJ, Zhang Y, Firatli E (Dec 1999). "Mutations of the cathepsin C gene are responsible for Papillon-Lefèvre syndrome". Journal of Medical Genetics. 36 (12): 881–7. doi:10.1136/jmg.36.12.881. PMC 1734286. PMID 10593994.
- Hart TC, Hart PS, Michalec MD, Zhang Y, Firatli E, Van Dyke TE, Stabholz A, Zlotogorski A, Shapira L, Soskolne WA, Zlorogorski A (Feb 2000). "Haim-Munk syndrome and Papillon-Lefèvre syndrome are allelic mutations in cathepsin C". Journal of Medical Genetics. 37 (2): 88–94. doi:10.1136/jmg.37.2.88. PMC 1734521. PMID 10662807.
- Hart TC, Hart PS, Michalec MD, Zhang Y, Marazita ML, Cooper M, Yassin OM, Nusier M, Walker S (Feb 2000). "Localisation of a gene for prepubertal periodontitis to chromosome 11q14 and identification of a cathepsin C gene mutation". Journal of Medical Genetics. 37 (2): 95–101. doi:10.1136/jmg.37.2.95. PMC 1734516. PMID 10662808.
- Suzuki Y, Ishihara D, Sasaki M, Nakagawa H, Hata H, Tsunoda T, Watanabe M, Komatsu T, Ota T, Isogai T, Suyama A, Sugano S (Mar 2000). "Statistical analysis of the 5' untranslated region of human mRNA using "Oligo-Capped" cDNA libraries". Genomics. 64 (3): 286–97. doi:10.1006/geno.2000.6076. PMID 10756096.
- Cigić B, Dahl SW, Pain RH (Oct 2000). "The residual pro-part of cathepsin C fulfills the criteria required for an intramolecular chaperone in folding and stabilizing the human proenzyme". Biochemistry. 39 (40): 12382–90. doi:10.1021/bi0008837. PMID 11015218.
- Hartley JL, Temple GF, Brasch MA (Nov 2000). "DNA cloning using in vitro site-specific recombination". Genome Research. 10 (11): 1788–95. doi:10.1101/gr.143000. PMC 310948. PMID 11076863.
- Hart PS, Zhang Y, Firatli E, Uygur C, Lotfazar M, Michalec MD, Marks JJ, Lu X, Coates BJ, Seow WK, Marshall R, Williams D, Reed JB, Wright JT, Hart TC (Dec 2000). "Identification of cathepsin C mutations in ethnically diverse papillon-Lefèvre syndrome patients". Journal of Medical Genetics. 37 (12): 927–32. doi:10.1136/jmg.37.12.927. PMC 1734492. PMID 11106356.
- Zhang Y, Lundgren T, Renvert S, Tatakis DN, Firatli E, Uygur C, Hart PS, Gorry MC, Marks JJ, Hart TC (Feb 2001). "Evidence of a founder effect for four cathepsin C gene mutations in Papillon-Lefèvre syndrome patients". Journal of Medical Genetics. 38 (2): 96–101. doi:10.1136/jmg.38.2.96. PMC 1734811. PMID 11158173.
- Nakano A, Nomura K, Nakano H, Ono Y, LaForgia S, Pulkkinen L, Hashimoto I, Uitto J (Feb 2001). "Papillon-Lefèvre syndrome: mutations and polymorphisms in the cathepsin C gene". The Journal of Investigative Dermatology. 116 (2): 339–43. doi:10.1046/j.1523-1747.2001.01244.x. PMID 11180012.
- Allende LM, García-Pérez MA, Moreno A, Corell A, Carasol M, Martínez-Canut P, Arnaiz-Villena A (Feb 2001). "Cathepsin C gene: First compound heterozygous patient with Papillon-Lefèvre syndrome and a novel symptomless mutation". Human Mutation. 17 (2): 152–3. doi:10.1002/1098-1004(200102)17:2<152::AID-HUMU10>3.0.CO;2-#. PMID 11180601. S2CID 196603893.
- Wiemann S, Weil B, Wellenreuther R, Gassenhuber J, Glassl S, Ansorge W, Böcher M, Blöcker H, Bauersachs S, Blum H, Lauber J, Düsterhöft A, Beyer A, Köhrer K, Strack N, Mewes HW, Ottenwälder B, Obermaier B, Tampe J, Heubner D, Wambutt R, Korn B, Klein M, Poustka A (Mar 2001). "Toward a catalog of human genes and proteins: sequencing and analysis of 500 novel complete protein coding human cDNAs". Genome Research. 11 (3): 422–35. doi:10.1101/gr.GR1547R. PMC 311072. PMID 11230166.
External links
[edit]- The MEROPS online database for peptidases and their inhibitors: C01.070
- Cathepsin+C at the U.S. National Library of Medicine Medical Subject Headings (MeSH)