RNA thermometer
An RNA thermometer (or RNA thermosensor) is a temperature-sensitive non-coding RNA molecule which regulates gene expression. RNA thermometers often regulate genes required during either a heat shock or cold shock response, but have been implicated in other regulatory roles such as in pathogenicity and starvation.[1]
In general, RNA thermometers operate by changing their secondary structure in response to temperature fluctuations. This structural transition can then expose or occlude important regions of RNA such as a ribosome binding site, which then affects the translation rate of a nearby protein-coding gene.
RNA thermometers, along with riboswitches, are used as examples in support of the RNA world hypothesis. This theory proposes that RNA was once the sole nucleic acid present in cells, and was replaced by the current DNA → RNA → protein system.[2]
Examples of RNA thermometers include FourU,[3] the Hsp90 cis-regulatory element,[4] the ROSE element,[5] the Lig RNA thermometer,[6] and the Hsp17 thermometer.[7]
Discovery
The first temperature-sensitive RNA element was reported in 1989.[8] Prior to this research, mutations upstream from the transcription start site in a lambda (λ) phage cIII mRNA were found to affect the level of translation of the cIII protein.[9] This protein is involved in selection of either a lytic or lysogenic life cycle in λ phage, with high concentrations of cIII promoting lysogeny.[9] Further study of this upstream RNA region identified two alternative secondary structures; experimental study found the structures to be interchangeable, and dependent on both magnesium ion concentration and temperature.[8][10] This RNA thermometer is now thought to encourage entry to a lytic cycle under heat stress in order for the bacteriophage to rapidly replicate and escape the host cell.[1]
The term "RNA thermometer" was not coined until 1999,[11] when it was applied to the rpoH RNA element identified in Escherichia coli.[12] More recently, bioinformatics searches have been employed to uncover several novel candidate RNA thermometers.[13] Traditional sequence-based searches are inefficient, however, as the secondary structure of the element is much more conserved than the nucleic acid sequence.[13]
Distribution
Most known RNA thermometers are located in the 5′ untranslated region (UTR) of messenger RNA encoding heat shock proteins—though it has been suggested this fact may be due, in part, to sampling bias and inherent difficulties of detecting short, unconserved RNA sequences in genomic data.[14][15]
Though predominantly found in prokaryotes, a potential RNA thermometer has been found in mammals including humans.[16] The candidate thermosensor heat shock RNA-1 (HSR1) activates heat-shock transcription factor 1 (HSF1) and induces protective proteins when cell temperature exceeds 37 °C (body temperature), thus preventing the cells from overheating.[16]
Structure

RNA thermometers are structurally simple and can be made from short RNA sequences; the smallest is just 44 nucleotides and is found in the mRNA of a heat-shock protein, hsp17, in Synechocystis species PCC 6803.[18][19] Generally these RNA elements range in length from 60 to 110 nucleotides[20] and they typically contain a hairpin with a small number of mismatched base pairs which reduce the stability of the structure, thereby allowing easier unfolding in response to a temperature increase.[21]
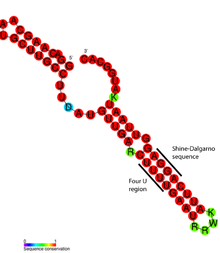
Detailed structural analysis of the ROSE RNA thermometer revealed that the mismatched bases are actually engaged in nonstandard basepairing that preserves the helical structure of the RNA (see figure). The unusual basepairs consist of G-G, U-U, and UC-U pairs. Since these noncanonical base pairs are relatively unstable, increased temperature causes local melting of the RNA structure in this region, exposing the Shine-Dalgarno sequence.[17]
Some RNA thermometers are significantly more complex than a single hairpin, as in the case of a region found in CspA mRNA which is thought to contain a pseudoknot, as well as multiple hairpins.[22][23]
Synthetic RNA thermometers have been designed with just a simple single-hairpin structure.[24] However, the secondary structure of such short RNA thermometers can be sensitive to mutation, as a single base change can render the hairpin inactive in vivo.[25]
Mechanism

RNA thermometers are found in the 5′ UTR of messenger RNA, upstream of a protein-coding gene.[1] Here they are able to occlude the ribosome binding site (RBS) and prevent translation of the mRNA into protein.[14] As temperature increases, the hairpin structure can 'melt' and expose the RBS or Shine-Dalgarno sequence to permit binding of the small ribosomal subunit (30S), which then assembles other translation machinery.[1] The start codon, typically found 8 nucleotides downstream of the Shine-Dalgarno sequence,[14] signals the beginning of a protein-coding gene which is then translated to a peptide product by the ribosome. In addition to this cis-acting mechanism, a lone example of a trans-acting RNA thermometer has been found in RpoS mRNA where it is thought to be involved in the starvation response.[1]
A specific example of an RNA thermometer motif is the FourU thermometer found in Salmonella enterica.[3] When exposed to temperatures above 45 °C, the stem-loop that base-pairs opposite the Shine-Dalgarno sequence becomes unpaired and allows the mRNA to enter the ribosome for translation to occur.[25] Mg2+ ion concentration has also been shown to affect the stability of FourU.[26] The most well-studied RNA thermometer is found in the rpoH gene in Escherichia coli.[27] This thermosensor upregulates heat shock proteins under high temperatures through σ32, a specialised heat-shock sigma factor.[11]
Though typically associated with heat-induced protein expression, RNA thermometers can also regulate cold-shock proteins.[22] For example, the expression of two 7kDa proteins are regulated by an RNA thermometer in the thermophilic bacterium Thermus thermophilus[28] and a similar mechanism has been identified in Enterobacteriales.[23]
RNA thermometers sensitive to temperatures of 37 °C can be used by pathogens to activate infection-specific genes.[14] For example, the upregulation of prfA, encoding a key transcriptional regulator of virulence genes in Listeria monocytogenes, was demonstrated by fusing the 5′ DNA of prfA to the green fluorescent protein gene; the gene fusion was then transcribed from the T7 promoter in E. coli, and fluorescence was observed at 37 °C but not at 30 °C.[29]
Implications for the RNA world hypothesis
The RNA world hypothesis states that RNA was once both the carrier of hereditary information and enzymatically active, with different sequences acting as biocatalysts, regulators and sensors.[30] The hypothesis then proposes that modern DNA, RNA and protein-based life evolved and selection replaced the majority of RNA's roles with other biomolecules.[2]
RNA thermometers and riboswitches are thought to be evolutionarily ancient due to their wide-scale distribution in distantly-related organisms.[31] It has been proposed that, in the RNA world, RNA thermosensors would have been responsible for temperature-dependent regulation of other RNA molecules.[2][32] RNA thermometers in modern organisms may be molecular fossils which could hint at a previously more widespread importance in an RNA world.[2]
Other examples
- Hsp90 cis-regulatory element regulates hsp90 in Drosophila, increasing the translation rate of the heat shock protein at high temperatures.[4]
- The ibpAB operon of E. coli is predicted to contain two co-operative RNA thermometers: a ROSE element and the IbpB thermometer.[33]
- ROSE1 and ROSEAT2 are found in hyphomicrobiales Bradyrhizobium japonicum and Agrobacterium tumefaciens respectively. They exist in the 5′ UTR of HspA mRNA, and repress heat shock protein translation at physiological temperatures.[5][34]
- Cyanobacterial RNA thermometers
- Intergenic RNA thermometer
- Neisseria RNA thermometers
- Lig RNA thermometer
References
- ^ a b c d e f Narberhaus F, Waldminghaus T, Chowdhury S (January 2006). "RNA thermometers". FEMS Microbiol. Rev. 30 (1): 3–16. doi:10.1111/j.1574-6976.2005.004.x. PMID 16438677.
- ^ a b c d Atkins JF, Gesteland RF, Cech T (2006). The RNA world: the nature of modern RNA suggests a prebiotic RNA world. Plainview, N.Y: Cold Spring Harbor Laboratory Press. ISBN 978-0-87969-739-6.
- ^ a b Waldminghaus T, Heidrich N, Brantl S, Narberhaus F (July 2007). "FourU: a novel type of RNA thermometer in Salmonella". Mol. Microbiol. 65 (2): 413–424. doi:10.1111/j.1365-2958.2007.05794.x. PMID 17630972.
- ^ a b Ahmed R, Duncan RF (2004). "Translational regulation of Hsp90 mRNA. AUG-proximal 5′-untranslated region elements essential for preferential heat shock translation". J Biol Chem. 279 (48): 49919–49930. doi:10.1074/jbc.M404681200. PMID 15347681.
- ^ a b Nocker A, Hausherr T, Balsiger S, Krstulovic NP, Hennecke H, Narberhaus F (2001). "A mRNA-based thermosensor controls expression of rhizobial heat shock genes". Nucleic Acids Res. 29 (23): 4800–4807. doi:10.1093/nar/29.23.4800. PMC 96696. PMID 11726689.
- ^ Matsunaga J, Schlax PJ, Haake DA (2013-11-15). "Role for cis-Acting RNA Sequences in the Temperature-Dependent Expression of the Multiadhesive Lig Proteins in Leptospira interrogans". Journal of Bacteriology. 195 (22): 5092–5101. doi:10.1128/jb.00663-13. ISSN 0021-9193. PMC 3811586. PMID 24013626.
- ^ Kortmann J, Sczodrok S, Rinnenthal J, Schwalbe H, Narberhaus F (2011). "Translation on demand by a simple RNA-based thermosensor". Nucleic Acids Res. 39 (7): 2855–2868. doi:10.1093/nar/gkq1252. PMC 3074152. PMID 21131278.
- ^ a b Altuvia S, Kornitzer D, Teff D, Oppenheim AB (1989-11-20). "Alternative mRNA structures of the cIII gene of bacteriophage lambda determine the rate of its translation initiation". Journal of Molecular Biology. 210 (2): 265–280. doi:10.1016/0022-2836(89)90329-X. PMID 2532257.
- ^ a b Altuvia S, Oppenheim AB (July 1986). "Translational regulatory signals within the coding region of the bacteriophage lambda cIII gene". Journal of Bacteriology. 167 (1): 415–419. doi:10.1128/jb.167.1.415-419.1986. PMC 212897. PMID 2941413.
- ^ Altuvia S, Kornitzer D, Kobi S, Oppenheim AB (1991-04-20). "Functional and structural elements of the mRNA of the cIII gene of bacteriophage lambda". Journal of Molecular Biology. 218 (4): 723–733. doi:10.1016/0022-2836(91)90261-4. PMID 1827163.
- ^ a b Storz G (1999-03-15). "An RNA thermometer". Genes & Development. 13 (6): 633–636. doi:10.1101/gad.13.6.633. PMID 10090718.
- ^ Morita MT, Tanaka Y, Kodama TS, Kyogoku Y, Yanagi H, Yura T (1999-03-15). "Translational induction of heat shock transcription factor sigma32: evidence for a built-in RNA thermosensor". Genes & Development. 13 (6): 655–665. doi:10.1101/gad.13.6.655. PMC 316556. PMID 10090722.
- ^ a b Waldminghaus T, Gaubig LC, Narberhaus F (November 2007). "Genome-wide bioinformatic prediction and experimental evaluation of potential RNA thermometers". Molecular Genetics and Genomics. 278 (5): 555–564. doi:10.1007/s00438-007-0272-7. PMID 17647020. S2CID 24747327.
- ^ a b c d Narberhaus F (2010). "Translational control of bacterial heat shock and virulence genes by temperature-sensing mRNAs". RNA Biol. 7 (1): 84–89. doi:10.4161/rna.7.1.10501. PMID 20009504. Retrieved 2011-04-23.
- ^ Johansson J (2009). "RNA thermosensors in bacterial pathogens". Bacterial Sensing and Signaling. Contributions to Microbiology. Vol. 16. Basel. pp. 150–160. doi:10.1159/000219378. ISBN 978-3-8055-9132-4. PMID 19494584.
{{cite book}}:|journal=ignored (help)CS1 maint: location missing publisher (link) - ^ a b Shamovsky I, Ivannikov M, Kandel ES, Gershon D, Nudler E (March 2006). "RNA-mediated response to heat shock in mammalian cells". Nature. 440 (7083): 556–560. Bibcode:2006Natur.440..556S. doi:10.1038/nature04518. PMID 16554823. S2CID 4311262.
- ^ a b Chowdhury S, Maris C, Allain FH, Narberhaus F (2006-06-07). "Molecular basis for temperature sensing by an RNA thermometer". The EMBO Journal. 25 (11): 2487–2497. doi:10.1038/sj.emboj.7601128. PMC 1478195. PMID 16710302.
- ^ Kortmann J, Sczodrok S, Rinnenthal J, Schwalbe H, Narberhaus F (April 2011). "Translation on demand by a simple RNA-based thermosensor". Nucleic Acids Research. 39 (7): 2855–2868. doi:10.1093/nar/gkq1252. PMC 3074152. PMID 21131278.
- ^ Kortmann J, Sczodrok S, Rinnenthal J, Schwalbe H, Narberhaus F (April 2011). "Translation on demand by a simple RNA-based thermosensor". Nucleic Acids Res. 39 (7): 2855–2868. doi:10.1093/nar/gkq1252. PMC 3074152. PMID 21131278.
- ^ Waldminghaus T, Fippinger A, Alfsmann J, Narberhaus F (December 2005). "RNA thermometers are common in alpha- and gamma-proteobacteria". Biol. Chem. 386 (12): 1279–1286. doi:10.1515/BC.2005.145. PMID 16336122. S2CID 84557068.
- ^ Narberhaus F (January–February 2010). "Translational control of bacterial heat shock and virulence genes by temperature-sensing mRNAs". RNA Biology. 7 (1): 84–89. doi:10.4161/rna.7.1.10501. PMID 20009504.
- ^ a b Breaker RR (January 2010). "RNA switches out in the cold". Mol. Cell. 37 (1): 1–2. doi:10.1016/j.molcel.2009.12.032. PMC 5315359. PMID 20129048.
- ^ a b Giuliodori AM, Di Pietro F, Marzi S, Masquida B, Wagner R, Romby P, Gualerzi CO, Pon CL (January 2010). "The cspA mRNA is a thermosensor that modulates translation of the cold-shock protein CspA". Mol. Cell. 37 (1): 21–33. doi:10.1016/j.molcel.2009.11.033. PMID 20129052.
- ^ Neupert J, Karcher D, Bock R (November 2008). "Design of simple synthetic RNA thermometers for temperature-controlled gene expression in Escherichia coli". Nucleic Acids Research. 36 (19): e124. doi:10.1093/nar/gkn545. PMC 2577334. PMID 18753148.
- ^ a b Nikolova EN, Al-Hashimi HM (September 2010). "Thermodynamics of RNA melting, one base pair at a time". RNA. 16 (9): 1687–1691. doi:10.1261/rna.2235010. PMC 2924531. PMID 20660079.
- ^ Rinnenthal J, Klinkert B, Narberhaus F, Schwalbe H (2011-07-04). "Modulation of the stability of the Salmonella fourU-type RNA thermometer". Nucleic Acids Research. 39 (18): 8258–8270. doi:10.1093/nar/gkr314. PMC 3185406. PMID 21727085.
- ^ Shah P, Gilchrist MA (2010). Spirin AS (ed.). "Is thermosensing property of RNA thermometers unique?". PLOS ONE. 5 (7): e11308. Bibcode:2010PLoSO...511308S. doi:10.1371/journal.pone.0011308. PMC 2896394. PMID 20625392.
- ^ Mega R, Manzoku M, Shinkai A, Nakagawa N, Kuramitsu S, Masui R (August 2010). "Very rapid induction of a cold shock protein by temperature downshift in Thermus thermophilus". Biochem. Biophys. Res. Commun. 399 (3): 336–340. doi:10.1016/j.bbrc.2010.07.065. PMID 20655297.
- ^ Johansson J, Mandin P, Renzoni A, Chiaruttini C, Springer M, Cossart P (September 2002). "An RNA thermosensor controls expression of virulence genes in Listeria monocytogenes". Cell. 110 (5): 551–561. doi:10.1016/S0092-8674(02)00905-4. PMID 12230973.
- ^ Gilbert W (February 1986). "The RNA World". Nature. 319 (6055): 618. Bibcode:1986Natur.319..618G. doi:10.1038/319618a0.
- ^ Serganov A, Patel DJ (October 2007). "Ribozymes, riboswitches and beyond: regulation of gene expression without proteins". Nature Reviews Genetics. 8 (10): 776–790. doi:10.1038/nrg2172. PMC 4689321. PMID 17846637.
- ^ Bocobza SE, Aharoni A (October 2008). "Switching the light on plant riboswitches". Trends in Plant Science. 13 (10): 526–533. doi:10.1016/j.tplants.2008.07.004. PMID 18778966.
- ^ Gaubig LC, Waldminghaus T, Narberhaus F (January 2011). "Multiple layers of control govern expression of the Escherichia coli ibpAB heat-shock operon". Microbiology. 157 (Pt 1): 66–76. doi:10.1099/mic.0.043802-0. PMID 20864473.
- ^ Balsiger S, Ragaz C, Baron C, Narberhaus F (2004). "Replicon-specific regulation of small heat shock genes in Agrobacterium tumefaciens". J Bacteriol. 186 (20): 6824–6829. doi:10.1128/JB.186.20.6824-6829.2004. PMC 522190. PMID 15466035.
