Sterile alpha motif
| SAM domain (Sterile alpha motif) | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Identifiers | |||||||||||
| Symbol | SAM_1 | ||||||||||
| Pfam | PF00536 | ||||||||||
| InterPro | IPR001660 | ||||||||||
| SMART | SAM | ||||||||||
| SCOP2 | 1b0x / SCOPe / SUPFAM | ||||||||||
| CDD | cd09487 | ||||||||||
| |||||||||||
| Ste50p-SAM | |||||||||
|---|---|---|---|---|---|---|---|---|---|
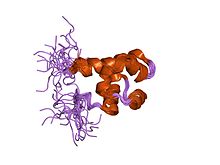 SAM domain from fungal protein Ste50p | |||||||||
| Identifiers | |||||||||
| Symbol | Ste50p-SAM | ||||||||
| Pfam | PF09235 | ||||||||
| Pfam clan | CL0003 | ||||||||
| InterPro | IPR015316 | ||||||||
| SCOP2 | 1uqv / SCOPe / SUPFAM | ||||||||
| |||||||||
In molecular biology, the protein domain Sterile alpha motif (or SAM) is a putative protein interaction module present in a wide variety of proteins[1] involved in many biological processes. The SAM domain that spreads over around 70 residues is found in diverse eukaryotic organisms.[2] SAM domains have been shown to homo- and hetero-oligomerise, forming multiple self-association architectures and also binding to various non-SAM domain-containing proteins,[3] nevertheless with a low affinity constant.[4]
SAM domains also appear to possess the ability to bind RNA.[5] Smaug, a protein that helps to establish a morphogen gradient in Drosophila embryos by repressing the translation of nanos (nos) mRNA, binds to the 3' untranslated region (UTR) of nos mRNA via two similar hairpin structures. The 3D crystal structure of the Smaug RNA-binding region shows a cluster of positively charged residues on the Smaug-SAM domain, which could be the RNA-binding surface. This electropositive potential is unique among all previously determined SAM-domain structures and is conserved among Smaug-SAM homologs. These results suggest that the SAM domain might have a primary role in RNA binding.
Structural analyses show that the SAM domain is arranged in a small five-helix bundle with two large interfaces.[3] In the case of the SAM domain of EPHB2, each of these interfaces is able to form dimers. The presence of these two distinct intermonomers binding surface suggest that SAM could form extended polymeric structures.[4]
Fungal SAM
In molecular biology, the protein domain Ste50p mainly in fungi and some other types of eukaryotes. It plays a role in the mitogen-activated protein kinase cascades, a type of cell signalling that helps the cell respond to external stimuli, more specifically mating, cell growth, and osmo-tolerance [6] in fungi.
Function
The protein domain Ste50p has a role in detecting pheromones for mating. It is thought to be found bound to Ste11p in order to prolong the pheromone-induced signaling response. Furthermore, it is also involved in aiding the cell to respond to nitrogen starvation.[7]
Structure
The fungal Ste50p SAM consists of six helices, which form a compact, globular fold. It is a monomer in solution and often undergoes heterodimerisation (and in some cases oligomerisation) of the protein.[7]
Protein interaction
The SAM domain of Ste50p often interacts with the SAM domain of Ste11p. They form bonds through this association. It is important to note that the SAM domain of one protein will bind to the SAM of a different protein. SAM domains do not self-associate in vitro.[7] There is significant evidence for Ste50p oligomerization in vivo.[8]
Human proteins containing this domain
ANKS1A; ANKS1B; ANKS3; ANKS4B; ANKS6; BFAR; BICC1; CASKIN1; CASKIN2; CENTD1; CNKSR2; CNKSR3; DDHD2; EPHA1; EPHA10; EPHA2; EPHA5; EPHA6; EPHA7; EPHA8; EPHB1; EPHB2; EPHB3; EPHB4; FAM59A; HPH2; INPPL1; L3MBTL3; PHC1; PHC2; PHC3; PPFIA1; PPFIA2; PPFIA3; PPFIA4; PPFIBP1; PPFIBP2; SAMD1; SAMD13; SAMD14; SAMD3; SAMD4A; SAMD4B; SAMD5; SAMD7; SAMD8; SAMD9; SCMH1; SCML1; SCML2; SEC23IP; SGMS1; SHANK1; SHANK2; SHANK3; STARD13; UBP1; USH1G; ZCCHC14; p63; p73;
References
- ^ Bork P, Ponting CP, Hofmann K, Schultz J (1997). "SAM as a protein interaction domain involved in developmental regulation". Protein Sci. 6 (1): 249–253. doi:10.1002/pro.5560060128. PMC 2143507. PMID 9007998.
- ^ Pawson T, Stapleton D, Balan I, Sicheri F (1999). "The crystal structure of an Eph receptor SAM domain reveals a mechanism for modular dimerization". Nat. Struct. Biol. 6 (1): 44–49. doi:10.1038/4917. PMID 9886291.
- ^ a b Simon J, Peterson AJ, Kyba M, Bornemann D, Morgan K, Brock HW (1997). "A domain shared by the Polycomb group proteins Scm and ph mediates heterotypic and homotypic interactions". Mol. Cell. Biol. 17 (11): 6683–6692. doi:10.1128/MCB.17.11.6683. PMC 232522. PMID 9343432.
- ^ a b Goodwill KE, Thanos CD, Bowie JU (1999). "Oligomeric structure of the human EphB2 receptor SAM domain". Science. 283 (5403): 833–836. doi:10.1126/science.283.5403.833. PMID 9933164.
- ^ Bowie JU, Kim CA (2003). "SAM domains: uniform structure, diversity of function". Trends Biochem. Sci. 28 (12): 625–628. doi:10.1016/j.tibs.2003.11.001. PMID 14659692.
- ^ Posas, F.; Witten, E. A.; Saito, H. (1998). "Requirement of STE50 for osmostress-induced activation of the STE11 mitogen-activated protein kinase kinase kinase in the high-osmolarity glycerol response pathway". Molecular and Cellular Biology. 18 (10): 5788–5796. doi:10.1128/mcb.18.10.5788. PMC 109165. PMID 9742096.
- ^ a b c Grimshaw SJ, Mott HR, Stott KM, Nielsen PR, Evetts KA, Hopkins LJ, Nietlispach D, Owen D (January 2004). "Structure of the sterile alpha motif (SAM) domain of the Saccharomyces cerevisiae mitogen-activated protein kinase pathway-modulating protein STE50 and analysis of its interaction with the STE11 SAM". J. Biol. Chem. 279 (3): 2192–201. doi:10.1074/jbc.M305605200. PMID 14573615.
- ^ Slaughter, BD; Huff JM; Wiegraebe W; Schwartz JW; Li R (2008). "SAM domain-based protein oligomerization observed by live-cell fluorescence fluctuation spectroscopy". PLOS One. 3 (4): e1931. doi:10.1371/journal.pone.0001931. PMC 2291563. PMID 18431466.
{{cite journal}}: CS1 maint: unflagged free DOI (link)
Structural evolution of p53, p63, and p73: Implication for heterotetramer formation
