Enhancer RNA
Enhancer RNAs (eRNAs) represent a class of relatively long non-coding RNA molecules (50-2000 nucleotides) transcribed from the DNA sequence of enhancer regions. They were first detected in 2010 through the use of genome-wide techniques such as RNA-seq and ChIP-seq.[1][2] eRNAs can be subdivided into two main classes: 1D eRNAs and 2D eRNAs, which differ primarily in terms of their size, polyadenylation state, and transcriptional directionality.[3] The expression of a given eRNA correlates with the activity of its corresponding enhancer in target genes.[4] Increasing evidence suggests that eRNAs actively play a role in transcriptional regulation in cis and in trans, and while their mechanisms of action remain unclear, a few models have been proposed.[3]
Discovery[edit]
Enhancers as sites of extragenic transcription were initially discovered in genome-wide studies that identified enhancers as common regions of RNA polymerase II (RNA pol II) binding and non-coding RNA transcription.[1][2] The level of RNA pol II–enhancer interaction and RNA transcript formation were found to be highly variable among these initial studies. Using explicit chromatin signature peaks, a significant proportion (~70%) of extragenic RNA Pol II transcription start sites were found to overlap enhancer sites in murine macrophages.[5] Out of 12,000 neuronal enhancers in the mouse genome, almost 25% of the sites were found to bind RNA Pol II and generate transcripts.[6] In parallel studies, 4,588 high confidence extragenic RNA Pol II binding sites were identified in murine macrophages stimulated with the inflammatory mediater lipopolysaccharide to induce transcription.[2] These eRNAs, unlike messenger RNAs (mRNAs), lacked modification by polyadenylation, were generally short and non-coding, and were bidirectionally transcribed. Later studies revealed the transcription of another type of eRNAs, generated through unidirectional transcription, that were longer and contained a poly A tail.[7] Furthermore, eRNA levels were correlated with mRNA levels of nearby genes, suggesting the potential regulatory and functional role of these non-coding enhancer RNA molecules.[1]
Biogenesis[edit]
Summary[edit]
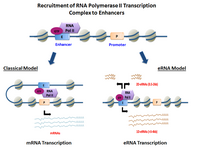
eRNAs are transcribed from DNA sequences upstream and downstream of extragenic enhancer regions.[8] Previously, several model enhancers have demonstrated the capability to directly recruit RNA Pol II and general transcription factors and form the pre-initiation complex (PIC) prior to the transcription start site at the promoter of genes. In certain cell types, activated enhancers have demonstrated the ability to both recruit RNA Pol II and also provide a template for active transcription of their local sequences.[2][1]
Depending on the directionality of transcription, enhancer regions generate two different types of non-coding transcripts, 1D-eRNAs and 2D-eRNAs. The nature of the pre-initiation complex and specific transcription factors recruited to the enhancer may control the type of eRNAs generated. After transcription, the majority of eRNAs remain in the nucleus.[9] In general, eRNAs are very unstable and actively degraded by the nuclear exosome. Not all enhancers are transcribed, with non-transcribed enhancers greatly outnumbering the transcribed ones in the order of magnitude of dozens of thousands in every given cell type.[5]
1D eRNAs[edit]
In most cases, unidirectional transcription of enhancer regions generates long (>4kb) and polyadenylated eRNAs. Enhancers that generate polyA+ eRNAs have a lower H3K4me1/me3 ratio in their chromatin signature than 2D-eRNAs.[7] PolyA+ eRNAs are distinct from long multiexonic poly transcripts (meRNAs) that are generated by transcription initiation at intragenic enhancers. These long non-coding RNAs, which accurately reflect the host gene's structure except for the alternative first exon, display poor coding potential.[10] As a result, polyA+ 1D-eRNAs may represent a mixed group of true enhancer-templated RNAs and multiexonic RNAs.
2D eRNAs[edit]
Bidirectional transcription at enhancer sites generates comparatively shorter (0.5-2kb) and non-polyadenylated eRNAs. Enhancers that generate polyA- eRNAs have a chromatin signature with a higher H3K4me1/me3 ratio than 1D-eRNAs. In general, enhancer transcription and production of bidirectional eRNAs demonstrate a strong correlation of enhancer activity on gene transcription.[11]
Frequency and timing of eRNA expression[edit]
Arner et al.[12] identified 65,423 transcribed enhancers (producing eRNA) among 33 different cell types under different conditions and different timings of stimulation. The transcription of enhancers generally preceded transcription of transcription factors which, in turn, generally preceded messenger RNA(mRNA) transcription of genes.
Carullo et al.[13] examined one particular cell type, neurons (from primary neuron cultures). They exhibited 28,492 putative enhancers generating eRNAs. These eRNAs were often transcribed from both strands of the enhancer DNA in opposite directions. Carullo et al.[13] used these cultured neurons to examine the timing of specific enhancer eRNAs compared to the mRNAs of their target genes. The cultured neurons were activated and RNA was isolated from those neurons at 0, 3.75, 5, 7.5, 15, 30, and 60 minutes after activation. In these experimental conditions, they found that 2 of the 5 enhancers of the immediate early gene (IEG) FOS, that is FOS enhancer 1 and FOS enhancer 3, became activated and initiated transcription of their eRNAs (eRNA1 and eRNA3). FOS eRNA1 and eRNA3 were significantly up-regulated within 7.5 minutes, whereas FOS mRNA was only upregulated 15 minutes after stimulation. Similar patterns occurred at IEGs FOSb and NR4A1, indicating that for many IEGs, eRNA induction precedes mRNA induction in response to neuronal activation.
While some enhancers can activate their target promoters at their target genes without transcribing eRNA, most active enhancers do transcribe eRNA during activation of their target promoters.[14]
Functions of eRNA found in the period 2013 to 2021[edit]
The functions for eRNA described below have been reported in diverse biological systems, often demonstrated with a small number of specific enhancer-target gene pairs. It is not clear to what extent the functions of eRNA described here can be generalized to most eRNAs.
eRNAs in loop formation[edit]

The chromosome loops shown in the figure, bringing an enhancer to the promoter of its target gene, may be directed and formed by the eRNA transcribed from the enhancer after the enhancer is activated.
A transcribed enhancer RNA (eRNA) interacting with the complex of Mediator proteins (see Figure), especially Mediator subunit 12 (MED12), appears to be essential in forming the chromosome loop that brings the enhancer into close association with the promoter of the target gene of the enhancer in the case of five genes studied by Lai et al.[15][16][17] Hou and Kraus,[18] describe two other studies reporting similar results. Arnold et al.[19] review another 5 instances where eRNA is active in forming the enhancer-promoter loop.
eRNAs interact with proteins to affect transcription[edit]
One well-studied eRNA is the eRNA of the enhancer that interacts with the promoter of the prostate specific antigen (PSA) gene.[20] The PSA eRNA is strongly up-regulated by the androgen receptor. High PSA eRNA then has a domino effect. PSA eRNA binds to and activates the positive transcription elongation factor P-TEFb protein complex which can then phosphorylate RNA polymerase II (RNAP II), initiating its activity in producing mRNA. P-TEFb can also phosphorylate the negative elongation factor NELF (which pauses RNAP II within 60 nucleotides after mRNA initiation begins). Phosphorylated NELF is released from RNAP II, then allowing RNAP II to have productive mRNA progression (see Figure). Up-regulated PSA eRNA thereby increases expression of 586 androgen receptor-responsive genes. Knockdown of PSA eRNA or deleting a set of nucleotides from PSA eRNA causes decreased presence of phosphorylated (active) RNAP II at these genes causing their reduced transcription.
The negative elongation factor NELF protein can also be released from its interaction with RNAP II by direct interaction with some eRNAs. Schaukowitch et al.[21] showed that the eRNAs of two immediate early genes (IEGs) directly interacted with the NELF protein to release NELF from the RNAP II paused at the promoters of these two genes, allowing these two genes to then be expressed.
In addition, eRNAs appear to interact with as many as 30 other proteins.[19][17][18]
Proposed mechanisms of function up until 2013[edit]

The notions that not all enhancers are transcribed at the same time and that eRNA transcription correlates with enhancer-specific activity support the idea that individual eRNAs carry distinct and relevant biological functions.[3] However, there is still no consensus on the functional significance of eRNAs. Furthermore, eRNAs can easily be degraded through exosomes and nonsense-mediated decay, which limits their potential as important transcriptional regulators.[22] To date, four main models of eRNA function have been proposed,[3] each supported by different lines of experimental evidence.
Transcriptional Noise[edit]
Since multiple studies have shown that RNA Pol II can be found at a very large number of extragenic regions, it is possible that eRNAs simply represent the product of random “leaky” transcription and carry no functional significance.[5] The non-specific activity of RNA Pol II would therefore allow extragenic transcriptional noise at sites where chromatin is already in an open and transcriptionally competent state. This would explain even tissue-specific eRNA expression[23] as open sites are tissue-specific as well.
Transcription-dependent effects[edit]
RNA Pol II-mediated gene transcription induces a local opening of chromatin state through the recruitment of histone acetyltransferases and other histone modifiers that promote euchromatin formation. It was proposed that the presence of these enzymes could also induce an opening of chromatin at enhancer regions, which are usually present at distant locations but can be recruited to target genes through looping of DNA.[24] In this model, eRNAs are therefore expressed in response to RNA Pol II transcription and therefore carry no biological function.
Functional activity in cis[edit]
While the two previous models implied that eRNAs were not functionally relevant, this mechanism states that eRNAs are functional molecules that exhibit cis activity. In this model, eRNAs can locally recruit regulatory proteins at their own site of synthesis. Supporting this hypothesis, transcripts originating from enhancers upstream of the Cyclin D1 gene are thought to serve as adaptors for the recruitment of histone acetyltransferases. It was found that depletion of these eRNAs led to Cyclin D1 transcriptional silencing.[9]
Functional activity in trans[edit]
The last model involves transcriptional regulation by eRNAs at distant chromosomal locations. Through the differential recruitment of protein complexes, eRNAs can affect the transcriptional competency of specific loci. Evf-2 represents a good example of such trans regulatory eRNA as it can induce the expression of Dlx2, which in turn can increase the activity of the Dlx5 and Dlx6 enhancers.[25] Trans-acting eRNAs might also be working in cis, and vice versa.
Experimental detection[edit]
The detection of eRNAs is fairly recent (2010) and has been made possible through the use of genome-wide investigation techniques such as RNA sequencing (RNA-seq) and chromatin immunoprecipitation-sequencing (ChIP-seq).[1] RNA-seq permits the direct identification of eRNAs by matching the detected transcript to the corresponding enhancer sequence through bioinformatic analyses.[26][4] ChIP-seq represents a less direct way to assess enhancer transcription but can also provide crucial information as specific chromatin marks are associated with active enhancers.[27] Although some data remain controversial, the consensus in the literature is that the best combination of histone post-translational modifications at active enhancers is made of H2AZ, H3K27ac, and a high ratio of H3K4me1 over H3K4me3.[27][28][29] ChIP experiments can also be conducted with antibodies that recognize RNA Pol II, which can be found at sites of active transcription.[5] The experimental detection of eRNAs is complicated by their low endogenous stability conferred by exosome degradation and nonsense-mediated decay.[22] A comparative study showed that assays enriching for capped and nascent RNAs (with strategies like nuclei run-on and size selection) could capture more eRNAs compared to canonical RNA-seq.[30] These assays include Global/Precision Run-on with cap-selection (GRO/PRO-cap), capped-small RNA-seq (csRNA-seq), Native Elongating Transcript-Cap Analysis of Gene Expression (NET-CAGE), and Precision Run-On sequencing (PRO-seq).[31] Nonetheless, the fact that eRNAs tend to be expressed from active enhancers might make their detection a useful tool to distinguish between active and inactive enhancers.
Implications in development and disease[edit]
Evidence that eRNAs cause downstream effects on the efficiency of enhancer activation and gene transcription suggests its functional capabilities and potential importance. The transcription factor p53 has been demonstrated to bind enhancer regions and generate eRNAs in a p53-dependent manner.[32] In cancer, p53 plays a central role in tumor suppression as mutations of the gene are shown to appear in 50% of tumors.[33] These p53-bound enhancer regions (p53BERs) are shown to interact with multiple local and distal gene targets involved in cell proliferation and survival. Furthermore, eRNAs generated by the activation of p53BERs are shown to be required for efficient transcription of the p53 target genes, indicating the likely important regulatory role of eRNAs in tumor suppression and cancer. Generally, mutations in eRNA have been shown to demonstrate similar phenotypic behavior in oncogenesis as compared to protein-coding RNA.[34]
Variations in enhancers have been implicated in human disease but a therapeutic approach to manipulate enhancer activity is currently not possible. With the emergence of eRNAs as important components in enhancer activity, powerful therapeutic tools such as RNAi may provide promising routes to target disruption of gene expression.
References[edit]
- ^ a b c d e Kim TK, Hemberg M, Gray JM, Costa AM, Bear DM, Wu J, et al. (May 2010). "Widespread transcription at neuronal activity-regulated enhancers". Nature. 465 (7295): 182–187. Bibcode:2010Natur.465..182K. doi:10.1038/nature09033. PMC 3020079. PMID 20393465.
- ^ a b c d De Santa F, Barozzi I, Mietton F, Ghisletti S, Polletti S, Tusi BK, et al. (May 2010). "A large fraction of extragenic RNA pol II transcription sites overlap enhancers". PLOS Biology. 8 (5): e1000384. doi:10.1371/journal.pbio.1000384. PMC 2867938. PMID 20485488.
- ^ a b c d Natoli G, Andrau JC (2012). "Noncoding transcription at enhancers: general principles and functional models". Annual Review of Genetics. 46: 1–19. doi:10.1146/annurev-genet-110711-155459. PMID 22905871.
- ^ a b Burren OS, Rubio García A, Javierre BM, Rainbow DB, Cairns J, Cooper NJ, et al. (September 2017). "Chromosome contacts in activated T cells identify autoimmune disease candidate genes". Genome Biology. 18 (1): 165. doi:10.1186/s13059-017-1285-0. PMC 5584004. PMID 28870212.
- ^ a b c d De Santa F, Barozzi I, Mietton F, Ghisletti S, Polletti S, Tusi BK, et al. (May 2010). Mattick JS (ed.). "A large fraction of extragenic RNA pol II transcription sites overlap enhancers". PLOS Biology. 8 (5): e1000384. doi:10.1371/journal.pbio.1000384. PMC 2867938. PMID 20485488.
- ^ Heintzman ND, Stuart RK, Hon G, Fu Y, Ching CW, Hawkins RD, et al. (March 2007). "Distinct and predictive chromatin signatures of transcriptional promoters and enhancers in the human genome". Nature Genetics. 39 (3): 311–318. doi:10.1038/ng1966. PMID 17277777. S2CID 1595885.
- ^ a b Koch F, Fenouil R, Gut M, Cauchy P, Albert TK, Zacarias-Cabeza J, et al. (July 2011). "Transcription initiation platforms and GTF recruitment at tissue-specific enhancers and promoters". Nature Structural & Molecular Biology. 18 (8): 956–963. doi:10.1038/nsmb.2085. PMID 21765417. S2CID 12778976.
- ^ Fedoseeva DM, Kretova OV, Tchurikov NA (2012). "Molecular analysis of enhancer RNAs and chromatin modifications in the region of their synthesis in Drosophila cells possessing genetic constructs". Doklady. Biochemistry and Biophysics. 442: 7–11. doi:10.1134/S1607672912010012. PMID 22419084. S2CID 14304312.
- ^ a b Wang X, Arai S, Song X, Reichart D, Du K, Pascual G, et al. (July 2008). "Induced ncRNAs allosterically modify RNA-binding proteins in cis to inhibit transcription". Nature. 454 (7200): 126–130. Bibcode:2008Natur.454..126W. doi:10.1038/nature06992. PMC 2823488. PMID 18509338.
- ^ Kowalczyk MS, Hughes JR, Garrick D, Lynch MD, Sharpe JA, Sloane-Stanley JA, et al. (February 2012). "Intragenic enhancers act as alternative promoters". Molecular Cell. 45 (4): 447–458. doi:10.1016/j.molcel.2011.12.021. hdl:2318/97658. PMID 22264824.
- ^ Wang D, Garcia-Bassets I, Benner C, Li W, Su X, Zhou Y, et al. (May 2011). "Reprogramming transcription by distinct classes of enhancers functionally defined by eRNA". Nature. 474 (7351): 390–394. doi:10.1038/nature10006. PMC 3117022. PMID 21572438.
- ^ Arner E, Daub CO, Vitting-Seerup K, Andersson R, Lilje B, Drabløs F, et al. (February 2015). "Transcribed enhancers lead waves of coordinated transcription in transitioning mammalian cells". Science. 347 (6225): 1010–1014. Bibcode:2015Sci...347.1010A. doi:10.1126/science.1259418. PMC 4681433. PMID 25678556.
- ^ a b Carullo NV, Phillips Iii RA, Simon RC, Soto SA, Hinds JE, Salisbury AJ, et al. (September 2020). "Enhancer RNAs predict enhancer-gene regulatory links and are critical for enhancer function in neuronal systems". Nucleic Acids Research. 48 (17): 9550–9570. doi:10.1093/nar/gkaa671. PMC 7515708. PMID 32810208.
- ^ Mikhaylichenko O, Bondarenko V, Harnett D, Schor IE, Males M, Viales RR, Furlong EE (January 2018). "The degree of enhancer or promoter activity is reflected by the levels and directionality of eRNA transcription". Genes & Development. 32 (1): 42–57. doi:10.1101/gad.308619.117. PMC 5828394. PMID 29378788.
- ^ Lai F, Orom UA, Cesaroni M, Beringer M, Taatjes DJ, Blobel GA, Shiekhattar R (February 2013). "Activating RNAs associate with Mediator to enhance chromatin architecture and transcription". Nature. 494 (7438): 497–501. Bibcode:2013Natur.494..497L. doi:10.1038/nature11884. PMC 4109059. PMID 23417068.
- ^ Allen BL, Taatjes DJ (March 2015). "The Mediator complex: a central integrator of transcription". Nature Reviews. Molecular Cell Biology. 16 (3): 155–166. doi:10.1038/nrm3951. PMC 4963239. PMID 25693131.
- ^ a b Sartorelli V, Lauberth SM (June 2020). "Enhancer RNAs are an important regulatory layer of the epigenome". Nature Structural & Molecular Biology. 27 (6): 521–528. doi:10.1038/s41594-020-0446-0. PMC 7343394. PMID 32514177.
- ^ a b Hou TY, Kraus WL (February 2021). "Spirits in the Material World: Enhancer RNAs in Transcriptional Regulation". Trends in Biochemical Sciences. 46 (2): 138–153. doi:10.1016/j.tibs.2020.08.007. PMC 7855021. PMID 32888773.
- ^ a b Arnold PR, Wells AD, Li XC (2019). "Diversity and Emerging Roles of Enhancer RNA in Regulation of Gene Expression and Cell Fate". Frontiers in Cell and Developmental Biology. 7: 377. doi:10.3389/fcell.2019.00377. PMC 6971116. PMID 31993419.
- ^ Zhao Y, Wang L, Ren S, Wang L, Blackburn PR, McNulty MS, et al. (April 2016). "Activation of P-TEFb by Androgen Receptor-Regulated Enhancer RNAs in Castration-Resistant Prostate Cancer". Cell Reports. 15 (3): 599–610. doi:10.1016/j.celrep.2016.03.038. PMC 5395199. PMID 27068475.
- ^ Schaukowitch K, Joo JY, Liu X, Watts JK, Martinez C, Kim TK (October 2014). "Enhancer RNA facilitates NELF release from immediate early genes". Molecular Cell. 56 (1): 29–42. doi:10.1016/j.molcel.2014.08.023. PMC 4186258. PMID 25263592.
- ^ a b Wyers F, Rougemaille M, Badis G, Rousselle JC, Dufour ME, Boulay J, et al. (June 2005). "Cryptic pol II transcripts are degraded by a nuclear quality control pathway involving a new poly(A) polymerase". Cell. 121 (5): 725–737. doi:10.1016/j.cell.2005.04.030. PMID 15935759.
- ^ Ren B (May 2010). "Transcription: Enhancers make non-coding RNA". Nature. 465 (7295): 173–174. Bibcode:2010Natur.465..173R. doi:10.1038/465173a. PMID 20463730.
- ^ Obrdlik A, Kukalev A, Louvet E, Farrants AK, Caputo L, Percipalle P (October 2008). "The histone acetyltransferase PCAF associates with actin and hnRNP U for RNA polymerase II transcription". Molecular and Cellular Biology. 28 (20): 6342–6357. doi:10.1128/MCB.00766-08. PMC 2577438. PMID 18710935.
- ^ Feng J, Bi C, Clark BS, Mady R, Shah P, Kohtz JD (June 2006). "The Evf-2 noncoding RNA is transcribed from the Dlx-5/6 ultraconserved region and functions as a Dlx-2 transcriptional coactivator". Genes & Development. 20 (11): 1470–1484. doi:10.1101/gad.1416106. PMC 1475760. PMID 16705037.
- ^ Wang Z, Gerstein M, Snyder M (January 2009). "RNA-Seq: a revolutionary tool for transcriptomics". Nature Reviews. Genetics. 10 (1): 57–63. doi:10.1038/nrg2484. PMC 2949280. PMID 19015660.
- ^ a b Barski A, Cuddapah S, Cui K, Roh TY, Schones DE, Wang Z, et al. (May 2007). "High-resolution profiling of histone methylations in the human genome". Cell. 129 (4): 823–837. doi:10.1016/j.cell.2007.05.009. PMID 17512414.
- ^ Melgar MF, Collins FS, Sethupathy P (November 2011). "Discovery of active enhancers through bidirectional expression of short transcripts". Genome Biology. 12 (11): R113. doi:10.1186/gb-2011-12-11-r113. PMC 3334599. PMID 22082242.
- ^ Creyghton MP, Cheng AW, Welstead GG, Kooistra T, Carey BW, Steine EJ, et al. (December 2010). "Histone H3K27ac separates active from poised enhancers and predicts developmental state". Proceedings of the National Academy of Sciences of the United States of America. 107 (50): 21931–21936. doi:10.1073/pnas.1016071107. PMC 3003124. PMID 21106759.
- ^ Yao L, Liang J, Ozer A, Leung AK, Lis JT, Yu H (July 2022). "A comparison of experimental assays and analytical methods for genome-wide identification of active enhancers". Nature Biotechnology. 40 (7): 1056–1065. doi:10.1038/s41587-022-01211-7. PMC 9288987. PMID 35177836.
- ^ Mahat DB, Kwak H, Booth GT, Jonkers IH, Danko CG, Patel RK, et al. (August 2016). "Base-pair-resolution genome-wide mapping of active RNA polymerases using precision nuclear run-on (PRO-seq)". Nature Protocols. 11 (8): 1455–1476. doi:10.1038/nprot.2016.086. PMC 5502525. PMID 27442863.
- ^ Melo CA, Drost J, Wijchers PJ, van de Werken H, de Wit E, Oude Vrielink JA, et al. (February 2013). "eRNAs are required for p53-dependent enhancer activity and gene transcription". Molecular Cell. 49 (3): 524–535. doi:10.1016/j.molcel.2012.11.021. PMID 23273978.
- ^ Vousden KH, Lu X (August 2002). "Live or let die: the cell's response to p53". Nature Reviews. Cancer. 2 (8): 594–604. doi:10.1038/nrc864. PMID 12154352. S2CID 6412605.
- ^ Zhang, Troy; Yu, Hui; Jiang, Limin; Bai, Yongsheng; Liu, Xiaoyi; Guo, Yan (January 2024). "Comprehensive Pan-Cancer Mutation Density Patterns in Enhancer RNA". International Journal of Molecular Sciences. 25 (1): 534. doi:10.3390/ijms25010534. ISSN 1422-0067. PMC 10778997. PMID 38203707.
External links[edit]
- Vista Enhancer Database
- Mouse ENCODE Project Archived 2012-12-13 at the Wayback Machine
- ENCODE Project at UCSC
- PEDB
