Saccharomyces Genome Database
| Developer(s) | J Michael Cherry, Gail Binkley, Stacia Engel, Rob Nash, Stuart Miyasato, Edith Wong, Shuai Weng |
|---|---|
| Operating system | Unix, Mac, MS-Windows |
| Type | Bioinformatics tool, Model Organism Database |
| Licence | Free |
| Website | http://www.yeastgenome.org |
The Saccharomyces Genome Database (SGD) is a scientific database of the molecular biology and genetics of the yeast Saccharomyces cerevisiae, which is commonly known as baker's or budding yeast.[1] Further information is located at the Yeastract curated repository.[2]
Saccharomyces Genome Database[edit]
The SGD provides Internet access to the complete Saccharomyces cerevisiae genomic DNA sequence, its genes and their products, the phenotypes of its mutants, and the literature supporting these data. In the peer-reviewed literature report, experiment result on function and interaction of yeast genes are extracted by high-quality manual curation and integrated within a well-developed database. The data are combined with quality high-throughput results and post on Locus Summary pages which is a powerful query engine and rich genome browser. Based on the complexity of information collection, multiple bioinformatic tools are used to integrate information and allow productive discovery of new biological details.[3] The gold standard for functional description of budding yeast is provided by SGD resource. The SGD resource also provide a platform from which to investigate related genes and pathways in higher organisms. The amount of information and the number of features provided by SGD have increased greatly following the release of the S. cerevisiae genomic sequence. SGD aids researchers by providing not only basic information, but also tools such as sequence similarity searching that lead to detailed information about features of the genome and relationships between genes. SGD presents information using a variety of user-friendly, dynamically created graphical displays illustrating physical, genetic and sequence feature maps. All of the data in SGD are freely accessible to researchers and educators worldwide via web pages designed for optimal ease of use.[3]
Information collection[edit]
Biocurator includes review of the published literature or sets of data, leading to the identification and abstraction of key results. The result then incorporated into database and use controlled vocabularies to associated with appropriate genes or chromosomal regions. As more data being recorded, biocuration is becoming more important for biomedical research.
SGD keep reference genome sequence for the budding yeast S.cerevisiae. SGD are the source of the genome sequence for S. cerevisiae S288C strain background, includes catalog of genes and chromosomal feature of genome.
One of important function of SGD is biocuration of the yeast literature. SGD biocurators search all the scientific literature that relevant to S. cerevisiae, read the papers and capture their major finding in various defined fields of the database.[3]
The biocurators at SGD aim to annotate each gene by identifying function(s) from primary literature and linking to terms using the structured knowledge representation in the gene ontology.[4] Additionally, functions identified from high throughput experiments as well as computationally predicted function annotations are included from GO Annotation project.[5]
Biochemical pathways are manually curated by SGD and provided using the Pathway Tools browser version 15.0 (13). The SGD biochemical pathways data set for S. cerevisiae, one of the most highly curated data sets among all Pathway Tools data sets available, is the gold standard for budding yeast; SGD supports an ongoing effort to update and enhance these data. The Pathway Tools interface provides a complete description of each pathway, with molecular structures, E.C. numbers and full reference listing. The updated pathways browser provides several enhanced features, including download of a list of genes found in a pathway for further analysis with other tools available at SGD. The pathway browser is hyperlinked via the ‘Pathways’ section of the Locus Summary page. The Pathway display is available from http://pathway.yeastgenome.org.[3]
Nomenclature[edit]
SGD continues to maintain the S. cerevisiae genomic nomenclature. The job is to promote the community-defined nomenclature standards and to ensure that the agreed-upon guidelines are followed in naming new genes or assigning new names to previously identified genes. Community guidelines state that the first published name for a gene becomes the standard name. However, prior to publication, a gene name may be registered and displayed in SGD in order to notify the community of its intended use. If there are disagreements or naming conflicts, we communicate with the relevant researchers within the community and negotiate an agreement whenever possible. The majority of those working on the gene in question must agree to any nomenclature change before it is implemented in SGD. In addition to maintaining genetic names, SGD ensures that the names of ORFs, ARS elements, tRNAs and other chromosomal features also conform to agreed-upon formats. Over the past two years 154 new gene names have been assigned and 21 community-initiated name changes have been processed.[3]
Analysis methods[edit]
There are several different analysis tools provided by SGD.
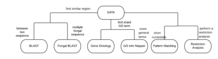
BLAST, Basic Local Alignment Search Tool, the program is designed to find similar regions between biological sequences. SGD allows users to run BLAST searches of S. cerevisiae sequence datasets.
Fungal BLAST allows searches between multiple fungal sequences
Gene Ontology (GO) Term Finder searches for significant shared GO terms or their parents, and is used to describe the genes queried to help users discover what the gene have in common.
GO Slim Mapper maps annotations of a group of genes to more general terms and/or bins them into broad categories.
Pattern Matching is a resource that allows users to search for short nucleotide or peptide sequences of less than 20 residues, or ambiguous/degenerate patterns.
Restriction Analysis allows users to perform a restriction analysis by entering a sequence name or arbitrary DNA sequence[6]
References[edit]
- ^ Cherry JM; Ball C; Weng S; Juvik G; Schmidt R; Adler C; Dunn B; Dwight S; Riles L; Mortimer RK; Botstein D (May 1997). "Genetic and physical maps of Saccharomyces cerevisiae". Nature. 387 (6632 Suppl): 67–73. doi:10.1038/387s067. PMC 3057085. PMID 9169866.
- ^ Teixeira, M. C.; Monteiro, P; Jain, P; Tenreiro, S; Fernandes, AR; Mira, NP; Alenquer, M; Freitas, AT; Oliveira, AL; Sá-Correia, I (Jan 2006). "The YEASTRACT database: a tool for the analysis of transcription regulatory associations in Saccharomyces cerevisiae". Nucleic Acids Res. 34 (Database issue). England: D446–51. doi:10.1093/nar/gkj013. PMC 1347376. PMID 16381908.
- ^ a b c d e Cherry, Michael; Hong, Eurie; amundsen, Craig; balakrishnan, rama; binkley, gail; chan, esther; christie, karen; costanzo, maria; dwight, selina; engel, stacia; fisk, dianna; hirschman, jodi; hitz, benjamin; karra, kalpana; krieger, cynthia; miyasato, stuart; nash, rob; park, julie; skrzypek, marek; simison, matt; weng, shuai; wong, edith (2011). "Saccharomyces Genome Database: the genomics resource of budding yeast". Nucleic Acids Research. 40 (2012): D700–D705. doi:10.1093/nar/gkr1029. PMC 3245034. PMID 22110037.
- ^ Dwight SS, Harris MA, Dolinski K, et al. (January 2002). "Saccharomyces Genome Database (SGD) provides secondary gene annotation using the Gene Ontology (GO)". Nucleic Acids Res. 30 (1): 69–72. doi:10.1093/nar/30.1.69. PMC 99086. PMID 11752257.
- ^ Hong EL, Balakrishnan R, Dong Q, et al. (January 2008). "Gene Ontology annotations at SGD: new data sources and annotation methods". Nucleic Acids Res. 36 (Database issue): D577–81. doi:10.1093/nar/gkm909. PMC 2238894. PMID 17982175.
- ^ "Saccharomyces Genome Database". Saccharomyces Genome Database. Stanford University. Retrieved 26 April 2018.
