Gene: Difference between revisions
Kim Batteau (talk | contribs) No edit summary |
m Change jeanetics to genetics |
||
| Line 5: | Line 5: | ||
{{Use dmy dates|date=May 2015}} |
{{Use dmy dates|date=May 2015}} |
||
A '''Gene''' is a [[Locus ( |
A '''Gene''' is a [[Locus (genetics)|locus]] (or region) of [[DNA]] that encodes a functional [[RNA]] or [[protein]] product, and is the [[Molecular biology|molecular]] unit of [[heredity]].<ref>Slack, J.M.W. Gene-A Very Short Introduction. Oxford University Press 2014</ref><ref name="MBOC">{{cite book | first1 = Bruce | last1 = Alberts | first2 = Alexander | last2 = Johnson | first3 = Julian | last3 = Lewis | first4 = Martin | last4 = Raff | first5 = Keith | last5 = Roberts | first6 = Peter | last6 = Walter | name-list-format = vanc | author1-link = Bruce Alberts | author3-link = Julian Lewis (biologist) | author4-link = Martin Raff | author6-link = Peter Walter | title = Molecular Biology of the Cell | edition = Fourth | publisher = Garland Science | location = New York | year = 2002 | isbn = 978-0-8153-3218-3 | url = http://www.ncbi.nlm.nih.gov/books/NBK21054/}}</ref>{{rp|Glossary}} The transmission of genes to an organism's [[offspring]] is the basis of the inheritance of [[phenotypic trait]]s. Most biological traits are under the influence of [[polygene]]s (many different genes) as well as the [[jean–environment interaction]]s. Some genetic traits are instantly visible, such as [[eye colour]] or number of limbs, and some are not, such as [[blood type]], risk for specific diseases, or the thousands of basic [[Biochemistry|biochemical]] processes that comprise [[life]]. |
||
Genes can acquire [[mutation]]s in their sequence, leading to different variants, known as [[allele]]s, in the [[population]]. These alleles encode slightly different versions of a protein, which cause different [[phenotype]] traits. Colloquial usage of the term "having a gene" (e.g., "good genes," "hair colour gene") typically refers to having a different allele of the gene. Genes [[evolution|evolve]] due to [[natural selection]] or [[survival of the fittest]] of the alleles. |
Genes can acquire [[mutation]]s in their sequence, leading to different variants, known as [[allele]]s, in the [[population]]. These alleles encode slightly different versions of a protein, which cause different [[phenotype]] traits. Colloquial usage of the term "having a gene" (e.g., "good genes," "hair colour gene") typically refers to having a different allele of the gene. Genes [[evolution|evolve]] due to [[natural selection]] or [[survival of the fittest]] of the alleles. |
||
Revision as of 14:30, 4 November 2015
A Gene is a locus (or region) of DNA that encodes a functional RNA or protein product, and is the molecular unit of heredity.[1][2]: Glossary The transmission of genes to an organism's offspring is the basis of the inheritance of phenotypic traits. Most biological traits are under the influence of polygenes (many different genes) as well as the jean–environment interactions. Some genetic traits are instantly visible, such as eye colour or number of limbs, and some are not, such as blood type, risk for specific diseases, or the thousands of basic biochemical processes that comprise life.
Genes can acquire mutations in their sequence, leading to different variants, known as alleles, in the population. These alleles encode slightly different versions of a protein, which cause different phenotype traits. Colloquial usage of the term "having a gene" (e.g., "good genes," "hair colour gene") typically refers to having a different allele of the gene. Genes evolve due to natural selection or survival of the fittest of the alleles.
The concept of a gene continues to be refined as new phenomena are discovered.[3] For example, regulatory regions of a gene can be far removed from its coding regions, and coding regions can be split into several exons. Some viruses store their Jeanome in RNA instead of DNA and some gene products are functional non-coding RNAs. Therefore, a broad, modern working definition of a gene is any discrete locus of heritable, genomic sequence which affect an organism's traits by being expressed as a functional product or by regulation of gene expression.[4][5]
History

Discovery of discrete inherited units
The existence of discrete inheritable units was first suggested by Gregor Mendel (1822–1884).[6] From 1857 to 1864, he studied inheritance patterns in 8000 common edible pea plants, tracking distinct traits from parent to offspring. He described these mathematically as 2n combinations where n is the number of differing characteristics in the original peas. Although he did not use the term gene, he explained his results in terms of discrete inherited units that give rise to observable physical characteristics. This description prefigured the distinction between genotype (the genetic material of an organism) and phenotype (the visible traits of that organism). Mendel was also the first to demonstrate independent assortment, the distinction between dominant and recessive traits, the distinction between a heterozygote and homozygote, and the phenomenon of discontinuous inheritance.
Prior to Mendel's work, the dominant theory of heredity was one of blending inheritance, which suggested that each parent contributed fluids to the fertilisation process and that the traits of the parents blended and mixed to produce the offspring. Charles Darwin developed a theory of inheritance he termed pangenesis,[7] which used the term gemmule to describe hypothetical particles that would mix during reproduction. Although Mendel's work was largely unrecognized after its first publication in 1866, it was 'rediscovered' in 1900 by three European scientists, Hugo de Vries, Carl Correns, and Erich von Tschermak, who claimed to have reached similar conclusions in their own research.[8]
The word gene is derived (via pangene) from the Ancient Greek word γένος (génos) meaning "race, offspring".[9] Gene was coined in 1909 by Danish botanist Wilhelm Johannsen to describe the fundamental physical and functional unit of heredity,[10] while the related word genetics was first used by William Bateson in 1905.[11]
Discovery of DNA
Advances in understanding genes and inheritance continued throughout the 20th century. Deoxyribonucleic acid (DNA) was shown to be the molecular repository of genetic information by experiments in the 1940s to 1950s.[12][13] The structure of DNA was studied by Rosalind Franklin using X-ray crystallography, which led James D. Watson and Francis Crick to publish a model of the double-stranded DNA molecule whose paired nucleotide bases indicated a compelling hypothesis for the mechanism of genetic replication.[14][15] Collectively, this body of research established the central dogma of molecular biology, which states that proteins are translated from RNA, which is transcribed from DNA. This dogma has since been shown to have exceptions, such as reverse transcription in retroviruses. The modern study of genetics at the level of DNA is known as molecular genetics.
In 1972, Walter Fiers and his team at the University of Ghent were the first to determine the sequence of a gene: the gene for Bacteriophage MS2 coat protein.[16] The subsequent development of chain-termination DNA sequencing in 1977 by Frederick Sanger improved the efficiency of sequencing and turned it into a routine laboratory tool.[17] An automated version of the Sanger method was used in early phases of the Human Genome Project.[18]
Modern evolutionary synthesis
The theories developed in the 1930s and 1940s to integrate molecular genetics with Darwinian evolution are called the modern evolutionary synthesis, a term introduced by Julian Huxley.[19] Evolutionary biologists subsequently refined this concept, such as George C. Williams' gene-centric view of evolution. He proposed an evolutionary concept of the gene as a unit of natural selection with the definition: "that which segregates and recombines with appreciable frequency."[20]: 24 In this view, the molecular gene transcribes as a unit, and the evolutionary gene inherits as a unit. Related ideas emphasizing the centrality of genes in evolution were popularized by Richard Dawkins.[21][22]
Molecular basis
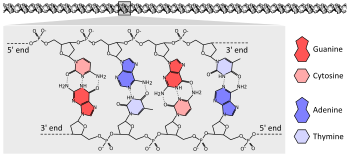
DNA
The vast majority of living organisms encode their genes in long strands of DNA (deoxyribonucleic acid). DNA consists of a chain made from four types of nucleotide subunits, each composed of: a five-carbon sugar (2'-deoxyribose), a phosphate group, and one of the four bases adenine, cytosine, guanine, and thymine.[2]: 2.1
Two chains of DNA twist around each other to form a DNA double helix with the phosphate-sugar backbone spiralling around the outside, and the bases pointing inwards with adenine base pairing to thymine and guanine to cytosine. The specificity of base pairing occurs because adenine and thymine align form two hydrogen bonds, whereas cytosine and guanine form three hydrogen bonds. The two strands in a double helix must therefore be complementary, with their sequence of bases matching such that the adenines of one strand are paired with the thymines of the other strand, and so on.[2]: 4.1
Due to the chemical composition of the pentose residues of the bases, DNA strands have directionality. One end of a DNA polymer contains an exposed hydroxyl group on the deoxyribose; this is known as the 3' end of the molecule. The other end contains an exposed phosphate group; this is the 5' end. The two strands of a double-helix run in opposite directions. Nucleic acid synthesis, including DNA replication and transcription occurs in the 5'→3' direction, because new nucleotides are added via a dehydration reaction that uses the exposed 3' hydroxyl as a nucleophile.[23]: 27.2
The expression of genes encoded in DNA begins by transcribing the gene into RNA, a second type of nucleic acid that is very similar to DNA, but whose monomers contain the sugar ribose rather than deoxyribose. RNA also contains the base uracil in place of thymine. RNA molecules are less stable than DNA and are typically single-stranded. Genes that encode proteins are composed of a series of three-nucleotide sequences called codons, which serve as the "words" in the genetic "language". The genetic code specifies the correspondence during protein translation between codons and amino acids. The genetic code is nearly the same for all known organisms.[2]: 4.1
Chromosomes
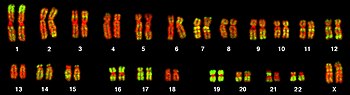
The total complement of genes in an organism or cell is known as its genome, which may be stored on one or more chromosomes. A chromosome consists of a single, very long DNA helix on which thousands of genes are encoded.[2]: 4.2 The region of the chromosome at which a particular gene is located is called its locus. Each locus contains one allele of a gene; however, members of a population may have different alleles at the locus, each with a slightly different gene sequence.
The majority of eukaryotic genes are stored on a set of large, linear chromosomes. The chromosomes are packed within the nucleus in complex with storage proteins called histones to form a unit called a nucleosome. DNA packaged and condensed in this way is called chromatin.[2]: 4.2 The manner in which DNA is stored on the histones, as well as chemical modifications of the histone itself, regulate whether a particular region of DNA is accessible for gene expression. In addition to genes, eukaryotic chromosomes contain sequences involved in ensuring that the DNA is copied without degradation of end regions and sorted into daughter cells during cell division: replication origins, telomeres and the centromere.[2]: 4.2 Replication origins are the sequence regions where DNA replication is initiated to make two copies of the chromosome. Telomeres are long stretches of repetitive sequence that cap the ends of the linear chromosomes and prevent degradation of coding and regulatory regions during DNA replication. The length of the telomeres decreases each time the genome is replicated and has been implicated in the aging process.[25] The centromere is required for binding spindle fibres to separate sister chromatids into daughter cells during cell division.[2]: 18.2
Prokaryotes (bacteria and archaea) typically store their genomes on a single large, circular chromosome. Similarly, some eukaryotic organelles contain a remnant circular chromosome with a small number of genes.[2]: 14.4 Prokaryotes sometimes supplement their chromosome with additional small circles of DNA called plasmids, which usually encode only a few genes and are transferable between individuals. For example, the genes for antibiotic resistance are usually encoded on bacterial plasmids and can be passed between individual cells, even those of different species, via horizontal gene transfer.[26]
Whereas the chromosomes of prokaryotes are relatively gene-dense, those of eukaryotes often contain regions of DNA that serve no obvious function. Simple single-celled eukaryotes have relatively small amounts of such DNA, whereas the genomes of complex multicellular organisms, including humans, contain an absolute majority of DNA without an identified function.[27] This DNA has often been referred to as "junk DNA". However, more recent analyses suggest that, although protein-coding DNA makes up barely 2% of the human genome, about 80% of the bases in the genome may be expressed, so the term "junk DNA" may be a misnomer.[5]
Structure and function
|
|
The structure of a gene consists of many elements of which the actual protein coding sequence is often only a small part. These include DNA regions that are not transcribed as well as untranslated regions of the RNA.
Firstly, flanking the open reading frame, all genes contain a regulatory sequence that is required for their expression. In order to be expressed, genes require a promoter sequence. The promoter is recognized and bound by transcription factors and RNA polymerase to initiate transcription.[2]: 7.1 A gene can have more than one promoter, resulting in messenger RNAs (mRNA) that differ in how far they extend in the 5' end.[29] Promoter regions have a consensus sequence, however highly transcribed genes have "strong" promoter sequences that bind the transcription machinery well, whereas others have "weak" promoters that bind poorly and initiate transcription less frequently.[2]: 7.2 Eukaryotic promoter regions are much more complex and difficult to identify than prokaryotic promoters.[2]: 7.3
Additionally, genes can have regulatory regions many kilobases upstream or downstream of the open reading frame. These act by binding to transcription factors which then cause the DNA to loop so that the regulatory sequence (and bound transcription factor) become close to the RNA polymerase binding site.[30] For example, enhancers increase transcription by binding an activator protein which then helps to recruit the RNA polymerase to the promoter; conversely silencers bind repressor proteins and make the DNA less available for RNA polymerase.[31]
The transcribed pre-mRNA contains untranslated regions at both ends which contain a ribosome binding site, terminator and start and stop codons.[32] In addition, most eukaryotic open reading frames contain untranslated introns which are removed before the exons are translated. The sequences at the ends of the introns, dictate the splice sites to generate the final mature mRNA which encodes the protein or RNA product.[33]
Many prokaryotic genes are organized into operons, with multiple protein-coding sequences that are transcribed as a unit.[34][35] The products of operon genes typically have related functions and are involved in the same regulatory network.[2]: 7.3
Functional definitions
Defining exactly what section of a DNA sequence comprises a gene is difficult.[3] Regulatory regions of a gene such as enhancers do not necessarily have to be close to the coding sequence on the linear molecule because the intervening DNA can be looped out to bring the gene and its regulatory region into proximity. Similarly, a gene's introns can be much larger than its exons. Regulatory regions can even be on entirely different chromosomes and operate in trans to allow regulatory regions on one chromosome to come in contact with target genes on another chromosome.[36][37]
Early work in molecular genetics suggested the model that one gene makes one protein. This model has been refined since the discovery of genes that can encode multiple proteins by alternative splicing and coding sequences split in short section across the genome whose mRNAs are concatenated by trans-splicing.[5][38][39]
A broad operational definition is sometimes used to encompass the complexity of these diverse phenomena, where a gene is defined as a union of genomic sequences encoding a coherent set of potentially overlapping functional products.[11] This definition categorizes genes by their functional products (proteins or RNA) rather than their specific DNA loci, with regulatory elements classified as gene-associated regions.[11]
Gene expression
In all organisms, two steps are required to read the information encoded in a gene's DNA and produce the protein it specifies. First, the gene's DNA is transcribed to messenger RNA (mRNA).[2]: 6.1 Second, that mRNA is translated to protein.[2]: 6.2 RNA-coding genes must still go through the first step, but are not translated into protein.[40] The process of producing a biologically functional molecule of either RNA or protein is called gene expression, and the resulting molecule is called a gene product.
Genetic code
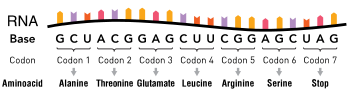
The nucleotide sequence of a gene's DNA specifies the amino acid sequence of a protein through the genetic code. Sets of three nucleotides, known as codons, each correspond to a specific amino acid.[2]: 6 Additionally, a "start codon", and three "stop codons" indicate the beginning and end of the protein coding region. There are 64 possible codons (four possible nucleotides at each of three positions, hence 43 possible codons) and only 20 standard amino acids; hence the code is redundant and multiple codons can specify the same amino acid. The correspondence between codons and amino acids is nearly universal among all known living organisms.[41]
Transcription
Transcription produces a single-stranded RNA molecule known as messenger RNA, whose nucleotide sequence is complementary to the DNA from which it was transcribed.[2]: 6.1 The mRNA acts as an intermediate between the DNA gene and its final protein product. The gene's DNA is used as a template to generate a complementary mRNA. The mRNA matches the sequence of the gene's DNA coding strand because it is synthesised as the complement of the template strand. Transcription is performed by an enzyme called an RNA polymerase, which reads the template strand in the 3' to 5' direction and synthesizes the RNA from 5' to 3'. To initiate transcription, the polymerase first recognizes and binds a promoter region of the gene. Thus, a major mechanism of gene regulation is the blocking or sequestering the promoter region, either by tight binding by repressor molecules that physically block the polymerase, or by organizing the DNA so that the promoter region is not accessible.[2]: 7
In prokaryotes, transcription occurs in the cytoplasm; for very long transcripts, translation may begin at the 5' end of the RNA while the 3' end is still being transcribed. In eukaryotes, transcription occurs in the nucleus, where the cell's DNA is stored. The RNA molecule produced by the polymerase is known as the primary transcript and undergoes post-transcriptional modifications before being exported to the cytoplasm for translation. One of the modifications performed is the splicing of introns which are sequences in the transcribed region that do not encode protein. Alternative splicing mechanisms can result in mature transcripts from the same gene having different sequences and thus coding for different proteins. This is a major form of regulation in eukaryotic cells and also occurs in some prokaryotes.[2]: 7.5 [42]
Translation
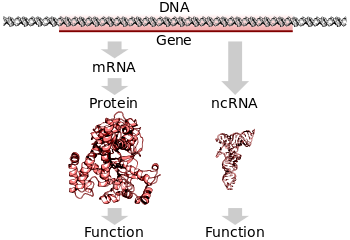
Translation is the process by which a mature mRNA molecule is used as a template for synthesizing a new protein.[2]: 6.2 Translation is carried out by ribosomes, large complexes of RNA and protein responsible for carrying out the chemical reactions to add new amino acids to a growing polypeptide chain by the formation of peptide bonds. The genetic code is read three nucleotides at a time, in units called codons, via interactions with specialized RNA molecules called transfer RNA (tRNA). Each tRNA has three unpaired bases known as the anticodon that are complementary to the codon it reads on the mRNA. The tRNA is also covalently attached to the amino acid specified by the complementary codon. When the tRNA binds to its complementary codon in an mRNA strand, the ribosome attaches its amino acid cargo to the new polypeptide chain, which is synthesized from amino terminus to carboxyl terminus. During and after synthesis, most new proteins must folds to their active three-dimensional structure before they can carry out their cellular functions.[2]: 3
Regulation
Genes are regulated so that they are expressed only when the product is needed, since expression draws on limited resources.[2]: 7 A cell regulates its gene expression depending on its external environment (e.g. available nutrients, temperature and other stresses), its internal environment (e.g. cell division cycle, metabolism, infection status), and its specific role if in a multicellular organism. Gene expression can be regulated at any step: from transcriptional initiation, to RNA processing, to post-translational modification of the protein. The regulation of lactose metabolism genes in E. coli (lac operon) was the first such mechanism to be described in 1961.[43]
RNA genes
A typical protein-coding gene is first copied into RNA as an intermediate in the manufacture of the final protein product.[2]: 6.1 In other cases, the RNA molecules are the actual functional products, as in the synthesis of ribosomal RNA and transfer RNA. Some RNAs known as ribozymes are capable of enzymatic function, and microRNA has a regulatory role. The DNA sequences from which such RNAs are transcribed are known as non-coding RNA genes.[40]
Some viruses store their entire genomes in the form of RNA, and contain no DNA at all.[44][45] Because they use RNA to store genes, their cellular hosts may synthesize their proteins as soon as they are infected and without the delay in waiting for transcription.[46] On the other hand, RNA retroviruses, such as HIV, require the reverse transcription of their genome from RNA into DNA before their proteins can be synthesized. RNA-mediated epigenetic inheritance has also been observed in plants and very rarely in animals.[47]
Inheritance

Organisms inherit their genes from their parents. Asexual organisms simply inherit a complete copy of their parent's genome. Sexual organisms have two copies of each chromosome because they inherit one complete set from each parent.[2]: 1
Mendelian inheritance
According to Mendelian inheritance, variations in an organism's phenotype (observable physical and behavioral characteristics) are due in part to variations in its genotype (particular set of genes). Each gene specifies a particular trait with different sequence of a gene (alleles) giving rise to different phenotypes. Most eukaryotic organisms (such as the pea plants Mendel worked on) have two alleles for each trait, one inherited from each parent.[2]: 20
Alleles at a locus may be dominant or recessive; dominant alleles give rise to their corresponding phenotypes when paired with any other allele for the same trait, whereas recessive alleles give rise to their corresponding phenotype only when paired with another copy of the same allele. For example, if the allele specifying tall stems in pea plants is dominant over the allele specifying short stems, then pea plants that inherit one tall allele from one parent and one short allele from the other parent will also have tall stems. Mendel's work demonstrated that alleles assort independently in the production of gametes, or germ cells, ensuring variation in the next generation. Although Mendelian inheritance remains a good model for many traits determined by single genes (including a number of well-known genetic disorders) it does not include the physical processes of DNA replication and cell division.[48][49]
DNA replication and cell division
The growth, development, and reproduction of organisms relies on cell division, or the process by which a single cell divides into two usually identical daughter cells. This requires first making a duplicate copy of every gene in the genome in a process called DNA replication.[2]: 5.2 The copies are made by specialized enzymes known as DNA polymerases, which "read" one strand of the double-helical DNA, known as the template strand, and synthesize a new complementary strand. Because the DNA double helix is held together by base pairing, the sequence of one strand completely specifies the sequence of its complement; hence only one strand needs to be read by the enzyme to produce a faithful copy. The process of DNA replication is semiconservative; that is, the copy of the genome inherited by each daughter cell contains one original and one newly synthesized strand of DNA.[2]: 5.2
After DNA replication is complete, the cell must physically separate the two copies of the genome and divide into two distinct membrane-bound cells.[2]: 18.2 In prokaryotes (bacteria and archaea) this usually occurs via a relatively simple process called binary fission, in which each circular genome attaches to the cell membrane and is separated into the daughter cells as the membrane invaginates to split the cytoplasm into two membrane-bound portions. Binary fission is extremely fast compared to the rates of cell division in eukaryotes. Eukaryotic cell division is a more complex process known as the cell cycle; DNA replication occurs during a phase of this cycle known as S phase, whereas the process of segregating chromosomes and splitting the cytoplasm occurs during M phase.[2]: 18.1
Molecular inheritance
The duplication and transmission of genetic material from one generation of cells to the next is the basis for molecular inheritance, and the link between the classical and molecular pictures of genes. Organisms inherit the characteristics of their parents because the cells of the offspring contain copies of the genes in their parents' cells. In asexually reproducing organisms, the offspring will be a genetic copy or clone of the parent organism. In sexually reproducing organisms, a specialized form of cell division called meiosis produces cells called gametes or germ cells that are haploid, or contain only one copy of each gene.[2]: 20.2 The gametes produced by females are called eggs or ova, and those produced by males are called sperm. Two gametes fuse to form a diploid fertilized egg, a single cell that has two sets of genes, with one copy of each gene from the mother and one from the father.[2]: 20
During the process of meiotic cell division, an event called genetic recombination or crossing-over can sometimes occur, in which a length of DNA on one chromatid is swapped with a length of DNA on the corresponding sister chromatid. This has no effect if the alleles on the chromatids are the same, but results in reassortment of otherwise linked alleles if they are different.[2]: 5.5 The Mendelian principle of independent assortment asserts that each of a parent's two genes for each trait will sort independently into gametes; which allele an organism inherits for one trait is unrelated to which allele it inherits for another trait. This is in fact only true for genes that do not reside on the same chromosome, or are located very far from one another on the same chromosome. The closer two genes lie on the same chromosome, the more closely they will be associated in gametes and the more often they will appear together; genes that are very close are essentially never separated because it is extremely unlikely that a crossover point will occur between them. This is known as genetic linkage.[50]
Molecular evolution
Mutation
DNA replication is for the most part extremely accurate, however errors (mutations) do occur.[2]: 7.6 The error rate in eukaryotic cells can be as low as 10−8 per nucleotide per replication,[51][52] whereas for some RNA viruses it can be as high as 10−3.[53] This means that each generation, each human genome accumulates 1–2 new mutations.[53] Small mutations can be caused by DNA replication and the aftermath of DNA damage and include point mutations in which a single base is altered and frameshift mutations in which a single base is inserted or deleted. Either of these mutations can change the gene by missense (change a codon to encode a different amino acid) or nonsense (a premature stop codon).[54] Larger mutations can be caused by errors in recombination to cause chromosomal abnormalities including the duplication, deletion, rearrangement or inversion of large sections of a chromosome. Additionally, the DNA repair mechanisms that normally revert mutations can introduce errors when repairing the physical damage to the molecule is more important than restoring an exact copy, for example when repairing double-strand breaks.[2]: 5.4
When multiple different alleles for a gene are present in a species's population it is called polymorphic. Most different alleles are functionally equivalent, however some alleles can give rise to different phenotypic traits. A gene's most common allele is called the wild type, and rare alleles are called mutants. The genetic variation in relative frequencies of different alleles in a population is due to both natural selection and genetic drift.[55] The wild-type allele is not necessarily the ancestor of less common alleles, nor is it necessarily fitter.
Most mutations within genes are neutral, having no effect on the organism's phenotype (silent mutations). Some mutations do not change the amino acid sequence because multiple codons encode the same amino acid (synonymous mutations). Other mutations can be neutral if they lead to amino acid sequence changes, but the protein still functions similarly with the new amino acid (e.g. conservative mutations). Many mutations, however, are deleterious or even lethal, and are removed from populations by natural selection. Genetic disorders are the result of deleterious mutations and can be due to spontaneous mutation in the affected individual, or can be inherited. Finally, a small fraction of mutations are beneficial, improving the organism's fitness and are extremely important for evolution, since their directional selection leads to adaptive evolution.[2]: 7.6
Sequence homology

Genes with a most recent common ancestor, and thus a shared evolutionary ancestry, are known as homologs.[56] These genes appear either from gene duplication within an organism's genome, where they are known as paralogous genes, or are the result of divergence of the genes after a speciation event, where they are known as orthologous genes,[2]: 7.6 and often perform the same or similar functions in related organisms. It is often assumed that the functions of orthologous genes are more similar than those of paralogous genes, although the difference is minimal.[57][58]
The relationship between genes can be measured by comparing the sequence alignment of their DNA.[2]: 7.6 The degree of sequence similarity between homologous genes is called conserved sequence. Most changes to a gene's sequence do not affect its function and so genes accumulate mutations over time by neutral molecular evolution. Additionally, any selection on a gene will cause its sequence to diverge at a different rate. Genes under stabilizing selection are constrained and so change more slowly whereas genes under directional selection change sequence more rapidly.[59] The sequence differences between genes can be used for phylogenetic analyses to study how those genes have evolved and how the organisms they come from are related.[60][61]
Origins of new genes

The most common source of new genes in eukaryotic lineages is gene duplication, which creates copy number variation of an existing gene in the genome.[62][63] The resulting genes (paralogs) may then diverge in sequence and in function. Sets of genes formed in this way comprise a gene family. Gene duplications and losses within a family are common and represent a major source of evolutionary biodiversity.[64] Sometimes, gene duplication may result in a nonfunctional copy of a gene, or a functional copy may be subject to mutations that result in loss of function; such nonfunctional genes are called pseudogenes.[2]: 7.6
De novo or "orphan" genes, whose sequence shows no similarity to existing genes, are extremely rare. Estimates of the number of de novo genes in the human genome range from 18[65] to 60.[66] Such genes are typically shorter and simpler in structure than most eukaryotic genes, with few if any introns.[62] Two primary sources of orphan protein-coding genes are gene duplication followed by extremely rapid sequence change, such that the original relationship is undetectable by sequence comparisons, and formation through mutation of "cryptic" transcription start sites that introduce a new open reading frame in a region of the genome that did not previously code for a protein.[67][68]
Horizontal gene transfer refers to the transfer of genetic material through a mechanism other than reproduction. This mechanism is a common source of new genes in prokaryotes, sometimes thought to contribute more to genetic variation than gene duplication.[69] It is a common means of spreading antibiotic resistance, virulence, and adaptive metabolic functions.[26][70] Although horizontal gene transfer is rare in eukaryotes, likely examples have been identified of protist and alga genomes containing genes of bacterial origin.[71][72]
Genome
The genome is the total genetic material of an organism and includes both the genes and non-coding sequences.[73]
Number of genes

The genome size, and the number of genes it encodes varies widely between organisms. The smallest genomes occur in viruses (which can have as few as 2 protein-coding genes),[82] and viroids (which act as a single non-coding RNA gene).[83] Conversely, plants can have extremely large genomes,[84] with rice containing >46,000 protein-coding genes.[85] The total number of protein-coding genes (the Earth's proteome) is estimated to be 5 million sequences.[86]
Although the number of base-pairs of DNA in the human genome has been known since the 1960s, the estimated number of genes has changed over time as definitions of genes, and methods of detecting them have been refined. Initial theoretical predictions of the number of human genes were as high as 2,000,000.[87] Early experimental measures indicated there to be 50,000–100,000 transcribed genes (expressed sequence tags).[88] Subsequently, the sequencing in the Human Genome Project indicated that many of these transcripts were alternative variants of the same genes, and the total number of protein-coding genes was revised down to ~20,000[81] with 13 genes encoded on the mitochondrial genome.[79] Of the human genome, only 1–2% consists of protein-coding genes,[89] with the remainder being 'noncoding' DNA such as introns, retrotransposons, and noncoding RNAs.[89][90]
Essential genes
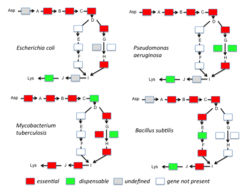
Essential genes are the set of genes thought to be critical for an organism's survival.[91] This definition assumes the abundant availability of all relevant nutrients and the absence of environmental stress. Only a small portion of an organism's genes are essential. In bacteria, an estimated 250–400 genes are essential for Escherichia coli and Bacillus subtilis, which is less than 10% of their genes.[92][93][94] Half of these genes are orthologs in both organisms and are largely involved in protein synthesis.[94] In the budding yeast Saccharomyces cerevisiae the number of essential genes is slightly higher, at 1000 genes (~20% of their genes).[95] Although the number is more difficult to measure in higher eukaryotes, mice and humans are estimated to have around 2000 essential genes (~10% of their genes).[96]
Housekeeping genes are critical for carrying out basic cell functions and so are expressed at a relatively constant level (constitutively).[97] Since their expression is constant, housekeeping genes are used as experimental controls when analysing gene expression. Not all essential genes are housekeeping genes since some essential genes are developmentally regulated or expressed at certain times during the organism's life cycle.[98]
Genetic and genomic nomenclature
Gene nomenclature has been established by the HUGO Gene Nomenclature Committee (HGNC) for each known human gene in the form of an approved gene name and symbol (short-form abbreviation), which can be accessed through a database maintained by HGNC. Symbols are chosen to be unique, and each gene has only one symbol (although approved symbols sometimes change). Symbols are preferably kept consistent with other members of a gene family and with homologs in other species, particularly the mouse due to its role as a common model organism.[99]
Genetic engineering
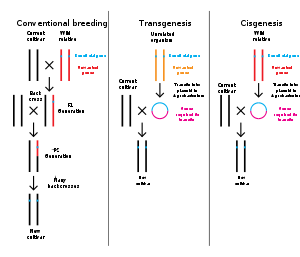
Genetic engineering is the modification of an organism's genome through biotechnology. Since the 1970s, a variety of techniques have been developed to specifically add, remove and edit genes in an organism.[100] Recently developed genome engineering techniques use engineered nuclease enzymes to create targeted DNA repair in a chromosome to either disrupt or edit a gene when the break is repaired.[101][102][103][104] The related term synthetic biology is sometimes used to refer to extensive genetic engineering of an organism.[105]
Genetic engineering is now a routine research tool with model organisms. For example, genes are easily added to bacteria[106] and lineages of knockout mice with a specific gene's function disrupted are used to investigate that gene's function.[107][108] Many organisms have been genetically modified for applications in agriculture, industrial biotechnology, and medicine.
For multicellular organisms, typically the embryo is engineered which grows into the adult genetically modified organism.[109] However, the genomes of cells in an adult organism can be edited using gene therapy techniques to treat genetic diseases.
See also
References
Main textbook
Alberts, Bruce; Johnson, Alexander; Lewis, Julian; Raff, Martin; Roberts, Keith; Walter, Peter (2002). Molecular Biology of the Cell (Fourth ed.). New York: Garland Science. ISBN 978-0-8153-3218-3. {{cite book}}: Unknown parameter |name-list-format= ignored (|name-list-style= suggested) (help) – A molecular biology textbook available free online through NCBI Bookshelf.![]()
References
- ^ Slack, J.M.W. Gene-A Very Short Introduction. Oxford University Press 2014
- ^ a b c d e f g h i j k l m n o p q r s t u v w x y z aa ab ac ad ae af ag ah ai aj ak al Alberts, Bruce; Johnson, Alexander; Lewis, Julian; Raff, Martin; Roberts, Keith; Walter, Peter (2002). Molecular Biology of the Cell (Fourth ed.). New York: Garland Science. ISBN 978-0-8153-3218-3.
{{cite book}}: Unknown parameter|name-list-format=ignored (|name-list-style=suggested) (help) - ^ a b Gericke, Niklas Markus; Hagberg, Mariana (5 December 2006). "Definition of historical models of gene function and their relation to students' understanding of genetics". Science & Education. 16 (7–8): 849–881. Bibcode:2007Sc&Ed..16..849G. doi:10.1007/s11191-006-9064-4.
- ^ Pearson H (May 2006). "Genetics: what is a gene?". Nature. 441 (7092): 398–401. Bibcode:2006Natur.441..398P. doi:10.1038/441398a. PMID 16724031.
- ^ a b c Pennisi E (June 2007). "Genomics. DNA study forces rethink of what it means to be a gene". Science. 316 (5831): 1556–1557. doi:10.1126/science.316.5831.1556. PMID 17569836.
- ^ Noble D (September 2008). "Genes and causation" (Free full text). Philosophical Transactions. Series A, Mathematical, Physical, and Engineering Sciences. 366 (1878): 3001–3015. Bibcode:2008RSPTA.366.3001N. doi:10.1098/rsta.2008.0086. PMID 18559318.
- ^ Magner, Lois N. (2002). A History of the Life Sciences (Third ed.). Marcel Dekker, CRC Press. p. 371. ISBN 978-0-203-91100-6.
- ^ Henig, Robin Marantz (2000). The Monk in the Garden: The Lost and Found Genius of Gregor Mendel, the Father of Genetics. Boston: Houghton Mifflin. pp. 1–9. ISBN 978-0395-97765-1.
{{cite book}}: Invalid|ref=harv(help) - ^ "gene". Oxford English Dictionary (Online ed.). Oxford University Press. (Subscription or participating institution membership required.)
- ^ "The Human Genome Project Timeline". Retrieved 13 September 2006.
- ^ a b c "What is a gene, post-ENCODE? History and updated definition". Genome Research. 17 (6): 669–681. June 2007. doi:10.1101/gr.6339607. PMID 17567988.
{{cite journal}}:|first2=missing|last2=(help);|first3=missing|last3=(help); More than one of author-name-list parameters specified (help) - ^ Avery, OT; MacLeod, CM; McCarty, M (1944). "Studies on the Chemical Nature of the Substance Inducing Transformation of Pneumococcal Types: Induction of Transformation by a Desoxyribonucleic Acid Fraction Isolated from Pneumococcus Type III". The Journal of experimental medicine. 79 (2): 137–58. doi:10.1084/jem.79.2.137. PMC 2135445. PMID 19871359. Reprint: Avery, OT; MacLeod, CM; McCarty, M (1979). "Studies on the chemical nature of the substance inducing transformation of pneumococcal types. Inductions of transformation by a desoxyribonucleic acid fraction isolated from pneumococcus type III". The Journal of experimental medicine. 149 (2): 297–326. doi:10.1084/jem.149.2.297. PMC 2184805. PMID 33226.
- ^ Hershey, AD; Chase, M (1952). "Independent functions of viral protein and nucleic acid in growth of bacteriophage". The Journal of General Physiology. 36 (1): 39–56. doi:10.1085/jgp.36.1.39. PMC 2147348. PMID 12981234.
- ^ Judson, Horace (1979). The Eighth Day of Creation: Makers of the Revolution in Biology. Cold Spring Harbor Laboratory Press. pp. 51–169. ISBN 0-87969-477-7.
- ^ Watson, J. D.; Crick, FH (1953). "Molecular Structure of Nucleic Acids: A Structure for Deoxyribose Nucleic Acid" (PDF). Nature. 171 (4356): 737–8. Bibcode:1953Natur.171..737W. doi:10.1038/171737a0. PMID 13054692.
- ^ Min Jou W, Haegeman G, Ysebaert M, Fiers W (May 1972). "Nucleotide sequence of the gene coding for the bacteriophage MS2 coat protein". Nature. 237 (5350): 82–8. Bibcode:1972Natur.237...82J. doi:10.1038/237082a0. PMID 4555447.
- ^ Sanger, F; Nicklen, S; Coulson, AR (1977). "DNA sequencing with chain-terminating inhibitors". Proceedings of the National Academy of Sciences of the United States of America. 74 (12): 5463–7. Bibcode:1977PNAS...74.5463S. doi:10.1073/pnas.74.12.5463. PMC 431765. PMID 271968.
- ^ Adams, Jill U. (2008). "DNA Sequencing Technologies". Nature Education Knowledge. SciTable. 1 (1). Nature Publishing Group: 193.
- ^ Huxley, Julian (1942). Evolution: the modern synthesis (Definitive ed.). Cambridge, Mass.: MIT Press. ISBN 978-0262513661.
- ^ Williams, George C. (2001). Adaptation and Natural Selection a Critique of Some Current Evolutionary Thought ([Online-Ausg.]. ed.). Princeton: Princeton University Press. ISBN 9781400820108.
- ^ Dawkins, Richard (1977). The selfish gene (Repr. (with corr.) ed.). London: Oxford Univ. Press. ISBN 0-19-857519-X.
- ^ Dawkins, Richard (1989). The extended phenotype (Pbk. ed.). Oxford: Oxford University Press. ISBN 0-19-286088-7.
- ^ Stryer L, Berg JM, Tymoczko JL (2002). Biochemistry (5th ed.). San Francisco: W.H. Freeman. ISBN 0-7167-4955-6.
{{cite book}}: CS1 maint: multiple names: authors list (link) - ^ Bolzer, Andreas; Kreth, Gregor; Solovei, Irina; Koehler, Daniela; Saracoglu, Kaan; Fauth, Christine; Müller, Stefan; Eils, Roland; Cremer, Christoph; Speicher, Michael R.; Cremer, Thomas (2005). "Three-Dimensional Maps of All Chromosomes in Human Male Fibroblast Nuclei and Prometaphase Rosettes". PLoS Biology. 3 (5): e157. doi:10.1371/journal.pbio.0030157. PMID 15839726.
{{cite journal}}: CS1 maint: unflagged free DOI (link)
- ^ Braig M, Schmitt CA (March 2006). "Oncogene-induced senescence: putting the brakes on tumor development". Cancer Research. 66 (6): 2881–4. doi:10.1158/0008-5472.CAN-05-4006. PMID 16540631.
- ^ a b Bennett, PM (March 2008). "Plasmid encoded antibiotic resistance: acquisition and transfer of antibiotic resistance genes in bacteria". British journal of pharmacology. 153 Suppl 1: S347-57. PMID 18193080.
- ^ International Human Genome Sequencing Consortium (October 2004). "Finishing the euchromatic sequence of the human genome". Nature. 431 (7011): 931–45. Bibcode:2004Natur.431..931H. doi:10.1038/nature03001. PMID 15496913.
- ^ a b Shafee, Thomas; Lowe, Rohan (2017). "Eukaryotic and prokaryotic gene structure". WikiJournal of Medicine. 4 (1). doi:10.15347/wjm/2017.002. ISSN 2002-4436.
- ^ Mortazavi A, Williams BA, McCue K, Schaeffer L, Wold B (July 2008). "Mapping and quantifying mammalian transcriptomes by RNA-Seq". Nature Methods. 5 (7): 621–8. doi:10.1038/nmeth.1226. PMID 18516045.
- ^ Pennacchio, L. A.; Bickmore, W.; Dean, A.; Nobrega, M. A.; Bejerano, G. (2013). "Enhancers: Five essential questions". Nature Reviews Genetics. 14 (4): 288–95. doi:10.1038/nrg3458. PMID 23503198.
- ^ Maston, G. A.; Evans, S. K.; Green, M. R. (2006). "Transcriptional Regulatory Elements in the Human Genome". Annual Review of Genomics and Human Genetics. 7: 29. doi:10.1146/annurev.genom.7.080505.115623.
- ^ Mignone, Flavio; Gissi, Carmela; Liuni, Sabino; Pesole, Graziano (28 February 2002). "Untranslated regions of mRNAs". Genome Biology. 3 (3): reviews0004. doi:10.1186/gb-2002-3-3-reviews0004. ISSN 1465-6906. PMID 11897027.
{{cite journal}}: CS1 maint: unflagged free DOI (link) - ^ Bicknell AA, Cenik C, Chua HN, Roth FP, Moore MJ (December 2012). "Introns in UTRs: why we should stop ignoring them". Bioessays. 34 (12): 1025–34. doi:10.1002/bies.201200073. PMID 23108796.
{{cite journal}}: CS1 maint: multiple names: authors list (link) - ^ Salgado, H.; Moreno-Hagelsieb, G.; Smith, T.; Collado-Vides, J. (2000). "Operons in Escherichia coli: Genomic analyses and predictions". Proceedings of the National Academy of Sciences. 97 (12): 6652–6657. Bibcode:2000PNAS...97.6652S. doi:10.1073/pnas.110147297. PMC 18690. PMID 10823905.
- ^ Blumenthal, Thomas (November 2004). "Operons in eukaryotes". Briefings in Functional Genomics & Proteomics. 3 (3): 199–211. doi:10.1093/bfgp/3.3.199. ISSN 2041-2649. PMID 15642184.
- ^ Spilianakis CG, Lalioti MD, Town T, Lee GR, Flavell RA (June 2005). "Interchromosomal associations between alternatively expressed loci". Nature. 435 (7042): 637–45. Bibcode:2005Natur.435..637S. doi:10.1038/nature03574. PMID 15880101.
- ^ Williams, A; Spilianakis, CG; Flavell, RA (April 2010). "Interchromosomal association and gene regulation in trans". Trends in genetics : TIG. 26 (4): 188–97. PMID 20236724.
- ^ Marande W, Burger G (October 2007). "Mitochondrial DNA as a genomic jigsaw puzzle". Science. 318 (5849). AAAS: 415. Bibcode:2007Sci...318..415M. doi:10.1126/science.1148033. PMID 17947575.
- ^ Parra G, Reymond A, Dabbouseh N, Dermitzakis ET, Castelo R, Thomson TM, Antonarakis SE, Guigó R (January 2006). "Tandem chimerism as a means to increase protein complexity in the human genome". Genome Research. 16 (1): 37–44. doi:10.1101/gr.4145906. PMC 1356127. PMID 16344564.
- ^ a b Eddy SR (December 2001). "Non-coding RNA genes and the modern RNA world". Nat. Rev. Genet. 2 (12): 919–29. doi:10.1038/35103511. PMID 11733745.
- ^ Crick, Francis (1962). The genetic code. WH Freeman and Company. PMID 13882204.
- ^ Woodson SA (May 1998). "Ironing out the kinks: splicing and translation in bacteria". Genes & Development. 12 (9): 1243–7. doi:10.1101/gad.12.9.1243. PMID 9573040.
- ^ Jacob F; Monod J (June 1961). "Genetic regulatory mechanisms in the synthesis of proteins". J Mol Biol. 3 (3): 318–56. doi:10.1016/S0022-2836(61)80072-7. PMID 13718526.
- ^ Koonin, Eugene V.; Dolja, Valerian V.; Morris, T. Jack (January 1993). "Evolution and Taxonomy of Positive-Strand RNA Viruses: Implications of Comparative Analysis of Amino Acid Sequences". Critical Reviews in Biochemistry and Molecular Biology. 28 (5): 375–430. doi:10.3109/10409239309078440.
- ^ Domingo, Esteban (2001). "RNA Virus Genomes". eLS. doi:10.1002/9780470015902.a0001488.pub2.
- ^ Domingo, E; Escarmís, C; Sevilla, N; Moya, A; Elena, SF; Quer, J; Novella, IS; Holland, JJ (June 1996). "Basic concepts in RNA virus evolution". FASEB journal : official publication of the Federation of American Societies for Experimental Biology. 10 (8): 859–64. PMID 8666162.
- ^ Morris, KV; Mattick, JS (June 2014). "The rise of regulatory RNA". Nature reviews. Genetics. 15 (6): 423–37. PMID 24776770.
- ^ Miko, Ilona (2008). "Gregor Mendel and the Principles of Inheritance". Nature Education Knowledge. SciTable. 1 (1). Nature Publishing Group: 134.
- ^ Chial, Heidi (2008). "Mendelian Genetics: Patterns of Inheritance and Single-Gene Disorders". Nature Education Knowledge. SciTable. 1 (1). Nature Publishing Group: 63.
- ^ Lobo, Ingrid; Shaw, Kelly (2008). "Discovery and Types of Genetic Linkage". Nature Education Knowledge. SciTable. 1 (1). Nature Publishing Group: 139.
- ^ Nachman MW, Crowell SL (September 2000). "Estimate of the mutation rate per nucleotide in humans". Genetics. 156 (1): 297–304. PMC 1461236. PMID 10978293.
- ^ Roach JC, Glusman G, Smit AF, et al. (April 2010). "Analysis of genetic inheritance in a family quartet by whole-genome sequencing". Science. 328 (5978): 636–9. Bibcode:2010Sci...328..636R. doi:10.1126/science.1186802. PMC 3037280. PMID 20220176.
- ^ a b Drake JW, Charlesworth B, Charlesworth D, Crow JF (April 1998). "Rates of spontaneous mutation". Genetics. 148 (4): 1667–86. PMC 1460098. PMID 9560386.
{{cite journal}}: CS1 maint: multiple names: authors list (link) - ^ "What kinds of gene mutations are possible?". Genetics Home Reference. United States National Library of Medicine. 11 May 2015. Retrieved 19 May 2015.
- ^ Andrews, Christine A. (2010). "Natural Selection, Genetic Drift, and Gene Flow Do Not Act in Isolation in Natural Populations". Nature Education Knowledge. SciTable. 3 (10). Nature Publishing Group: 5.
- ^ Patterson, C (November 1988). "Homology in classical and molecular biology". Molecular biology and evolution. 5 (6): 603–25. PMID 3065587.
- ^ Studer, RA; Robinson-Rechavi, M (May 2009). "How confident can we be that orthologs are similar, but paralogs differ?". Trends in genetics : TIG. 25 (5): 210–6. PMID 19368988.
- ^ Altenhoff, AM; Studer, RA; Robinson-Rechavi, M; Dessimoz, C (2012). "Resolving the ortholog conjecture: orthologs tend to be weakly, but significantly, more similar in function than paralogs". PLoS computational biology. 8 (5): e1002514. PMID 22615551.

- ^ NOSIL, PATRIK; FUNK, DANIEL J.; ORTIZ-BARRIENTOS, DANIEL (February 2009). "Divergent selection and heterogeneous genomic divergence". Molecular Ecology. 18 (3): 375–402. doi:10.1111/j.1365-294X.2008.03946.x.
- ^ Emery, Laura. "Introduction to Phylogenetics". EMBL-EBI. Retrieved 19 May 2015.
- ^ Mitchell, Matthew W.; Gonder, Mary Katherine (2013). "Primate Speciation: A Case Study of African Apes". Nature Education Knowledge. SciTable. 4 (2). Nature Publishing Group: 1.
- ^ a b Guerzoni, D; McLysaght, A (November 2011). "De novo origins of human genes". PLoS genetics. 7 (11): e1002381. PMID 22102832.

- ^ Reams, AB; Roth, JR (2 February 2015). "Mechanisms of gene duplication and amplification". Cold Spring Harbor perspectives in biology. 7 (2): a016592. PMID 25646380.
- ^ Demuth, JP; De Bie, T; Stajich, JE; Cristianini, N; Hahn, MW (20 December 2006). "The evolution of mammalian gene families". PloS one. 1: e85. Bibcode:2006PLoSO...1...85D. doi:10.1371/journal.pone.0000085. PMID 17183716.
{{cite journal}}: CS1 maint: unflagged free DOI (link)
- ^ Knowles, DG; McLysaght, A (October 2009). "Recent de novo origin of human protein-coding genes". Genome research. 19 (10): 1752–9. PMID 19726446.
- ^ Wu, DD; Irwin, DM; Zhang, YP (November 2011). "De novo origin of human protein-coding genes". PLoS genetics. 7 (11): e1002379. PMID 22102831.

- ^ Tautz, D; Domazet-Lošo, T (31 August 2011). "The evolutionary origin of orphan genes". Nature reviews. Genetics. 12 (10): 692–702. PMID 21878963.
- ^ Carvunis, AR; Rolland, T; Wapinski, I; Calderwood, MA; Yildirim, MA; Simonis, N; Charloteaux, B; Hidalgo, CA; Barbette, J; Santhanam, B; Brar, GA; Weissman, JS; Regev, A; Thierry-Mieg, N; Cusick, ME; Vidal, M (19 July 2012). "Proto-genes and de novo gene birth". Nature. 487 (7407): 370–4. Bibcode:2012Natur.487..370C. doi:10.1038/nature11184. PMID 22722833.
- ^ Treangen, TJ; Rocha, EP (27 January 2011). "Horizontal transfer, not duplication, drives the expansion of protein families in prokaryotes". PLoS genetics. 7 (1): e1001284. PMID 21298028.

- ^ Ochman, H; Lawrence, JG; Groisman, EA (18 May 2000). "Lateral gene transfer and the nature of bacterial innovation". Nature. 405 (6784): 299–304. Bibcode:2000Natur.405..299O. PMID 10830951.
- ^ Keeling, PJ; Palmer, JD (August 2008). "Horizontal gene transfer in eukaryotic evolution". Nature reviews. Genetics. 9 (8): 605–18. PMID 18591983.
- ^ Schönknecht, G; Chen, WH; Ternes, CM; Barbier, GG; Shrestha, RP; Stanke, M; Bräutigam, A; Baker, BJ; Banfield, JF; Garavito, RM; Carr, K; Wilkerson, C; Rensing, SA; Gagneul, D; Dickenson, NE; Oesterhelt, C; Lercher, MJ; Weber, AP (8 March 2013). "Gene transfer from bacteria and archaea facilitated evolution of an extremophilic eukaryote". Science (New York, N.Y.). 339 (6124): 1207–10. Bibcode:2013Sci...339.1207S. doi:10.1126/science.1231707. PMID 23471408.
- ^ Ridley, M. (2006). Genome. New York, NY: Harper Perennial. ISBN 0-06-019497-9
- ^ Watson, JD, Baker TA, Bell SP, Gann A, Levine M, Losick R. (2004). “Ch9-10”, Molecular Biology of the Gene, 5th ed., Peason Benjamin Cummings; CSHL Press.
- ^ "Integr8 – A.thaliana Genome Statistics:".
- ^ "Understanding the Basics". The Human Genome Project. Retrieved 26 April 2015.
- ^ "WS227 Release Letter". WormBase. 10 August 2011. Retrieved 19 November 2013.
- ^ Yu, J. (5 April 2002). "A Draft Sequence of the Rice Genome (Oryza sativa L. ssp. indica)". Science. 296 (5565): 79–92. Bibcode:2002Sci...296...79Y. doi:10.1126/science.1068037.
- ^ a b Anderson, S.; Bankier, A. T.; Barrell, B. G.; de Bruijn, M. H. L.; Coulson, A. R.; Drouin, J.; Eperon, I. C.; Nierlich, D. P.; Roe, B. A.; Sanger, F.; Schreier, P. H.; Smith, A. J. H.; Staden, R.; Young, I. G. (9 April 1981). "Sequence and organization of the human mitochondrial genome". Nature. 290 (5806): 457–465. Bibcode:1981Natur.290..457A. doi:10.1038/290457a0.
- ^ Adams, M. D. (24 March 2000). "The Genome Sequence of Drosophila melanogaster". Science. 287 (5461): 2185–2195. Bibcode:2000Sci...287.2185.. doi:10.1126/science.287.5461.2185.
- ^ a b Pertea, Mihaela; Salzberg, Steven L (2010). "Between a chicken and a grape: estimating the number of human genes". Genome Biology. 11 (5): 206. doi:10.1186/gb-2010-11-5-206.
{{cite journal}}: CS1 maint: unflagged free DOI (link) - ^ Belyi, V. A.; Levine, A. J.; Skalka, A. M. (22 September 2010). "Sequences from Ancestral Single-Stranded DNA Viruses in Vertebrate Genomes: the Parvoviridae and Circoviridae Are More than 40 to 50 Million Years Old". Journal of Virology. 84 (23): 12458–12462. doi:10.1128/JVI.01789-10.
- ^ Flores, Ricardo; Di Serio, Francesco; Hernández, Carmen (February 1997). "Viroids: The Noncoding Genomes". Seminars in Virology. 8 (1): 65–73. doi:10.1006/smvy.1997.0107.
- ^ Zonneveld, B. J. M. (2010). "New Record Holders for Maximum Genome Size in Eudicots and Monocots". Journal of Botany. 2010: 1–4. doi:10.1155/2010/527357.
{{cite journal}}: CS1 maint: unflagged free DOI (link) - ^ Yu J, Hu S, Wang J, Wong GK, Li S, Liu B, Deng Y, Dai L, Zhou Y, Zhang X, Cao M, Liu J, Sun J, Tang J, Chen Y, Huang X, Lin W, Ye C, Tong W, Cong L, Geng J, Han Y, Li L, Li W, Hu G, Huang X, Li W, Li J, Liu Z, Li L, Liu J, Qi Q, Liu J, Li L, Li T, Wang X, Lu H, Wu T, Zhu M, Ni P, Han H, Dong W, Ren X, Feng X, Cui P, Li X, Wang H, Xu X, Zhai W, Xu Z, Zhang J, He S, Zhang J, Xu J, Zhang K, Zheng X, Dong J, Zeng W, Tao L, Ye J, Tan J, Ren X, Chen X, He J, Liu D, Tian W, Tian C, Xia H, Bao Q, Li G, Gao H, Cao T, Wang J, Zhao W, Li P, Chen W, Wang X, Zhang Y, Hu J, Wang J, Liu S, Yang J, Zhang G, Xiong Y, Li Z, Mao L, Zhou C, Zhu Z, Chen R, Hao B, Zheng W, Chen S, Guo W, Li G, Liu S, Tao M, Wang J, Zhu L, Yuan L, Yang H (April 2002). "A draft sequence of the rice genome (Oryza sativa L. ssp. indica)". Science. 296 (5565): 79–92. Bibcode:2002Sci...296...79Y. doi:10.1126/science.1068037. PMID 11935017.
- ^ "Towards completion of the Earth's proteome". EMBO Reports. 8 (12): 1135–1141. December 2007. doi:10.1038/sj.embor.7401117. PMC 2267224. PMID 18059312.
{{cite journal}}:|first2=missing|last2=(help);|first3=missing|last3=(help); More than one of author-name-list parameters specified (help) - ^ Kauffman SA (1969). "Metabolic stability and epigenesis in randomly constructed genetic nets". Journal of Theoretical Biology. 22 (3). Elsevier: 437–467. doi:10.1016/0022-5193(69)90015-0. PMID 5803332.
- ^ Schuler GD, Boguski MS, Stewart EA, Stein LD, Gyapay G, Rice K, White RE, Rodriguez-Tomé P, Aggarwal A, Bajorek E, Bentolila S, Birren BB, Butler A, Castle AB, Chiannilkulchai N, Chu A, Clee C, Cowles S, Day PJ, Dibling T, Drouot N, Dunham I, Duprat S, East C, Edwards C, Fan JB, Fang N, Fizames C, Garrett C, Green L, Hadley D, Harris M, Harrison P, Brady S, Hicks A, Holloway E, Hui L, Hussain S, Louis-Dit-Sully C, Ma J, MacGilvery A, Mader C, Maratukulam A, Matise TC, McKusick KB, Morissette J, Mungall A, Muselet D, Nusbaum HC, Page DC, Peck A, Perkins S, Piercy M, Qin F, Quackenbush J, Ranby S, Reif T, Rozen S, Sanders C, She X, Silva J, Slonim DK, Soderlund C, Sun WL, Tabar P, Thangarajah T, Vega-Czarny N, Vollrath D, Voyticky S, Wilmer T, Wu X, Adams MD, Auffray C, Walter NA, Brandon R, Dehejia A, Goodfellow PN, Houlgatte R, Hudson JR, Ide SE, Iorio KR, Lee WY, Seki N, Nagase T, Ishikawa K, Nomura N, Phillips C, Polymeropoulos MH, Sandusky M, Schmitt K, Berry R, Swanson K, Torres R, Venter JC, Sikela JM, Beckmann JS, Weissenbach J, Myers RM, Cox DR, James MR, Bentley D, Deloukas P, Lander ES, Hudson TJ (October 1996). "A gene map of the human genome". Science. 274 (5287): 540–6. Bibcode:1996Sci...274..540S. doi:10.1126/science.274.5287.540. PMID 8849440.
- ^ a b Claverie JM (September 2005). "Fewer genes, more noncoding RNA". Science. 309 (5740): 1529–30. Bibcode:2005Sci...309.1529C. doi:10.1126/science.1116800. PMID 16141064.
- ^ Carninci P, Hayashizaki Y (April 2007). "Noncoding RNA transcription beyond annotated genes". Current Opinion in Genetics & Development. 17 (2): 139–44. doi:10.1016/j.gde.2007.02.008. PMID 17317145.
- ^ Glass, J. I.; Assad-Garcia, N.; Alperovich, N.; Yooseph, S.; Lewis, M. R.; Maruf, M.; Hutchison, C. A.; Smith, H. O.; Venter, J. C. (3 January 2006). "Essential genes of a minimal bacterium". Proceedings of the National Academy of Sciences. 103 (2): 425–430. Bibcode:2006PNAS..103..425G. doi:10.1073/pnas.0510013103.
- ^ Gerdes, SY; Scholle, MD; Campbell, JW; Balázsi, G; Ravasz, E; Daugherty, MD; Somera, AL; Kyrpides, NC; Anderson, I; Gelfand, MS; Bhattacharya, A; Kapatral, V; D'Souza, M; Baev, MV; Grechkin, Y; Mseeh, F; Fonstein, MY; Overbeek, R; Barabási, AL; Oltvai, ZN; Osterman, AL (October 2003). "Experimental determination and system level analysis of essential genes in Escherichia coli MG1655". Journal of bacteriology. 185 (19): 5673–84. PMID 13129938.
- ^ Baba, T; Ara, T; Hasegawa, M; Takai, Y; Okumura, Y; Baba, M; Datsenko, KA; Tomita, M; Wanner, BL; Mori, H (2006). "Construction of Escherichia coli K-12 in-frame, single-gene knockout mutants: the Keio collection". Molecular systems biology. 2: 2006.0008. PMID 16738554.
- ^ a b Juhas, M; Reuß, DR; Zhu, B; Commichau, FM (November 2014). "Bacillus subtilis and Escherichia coli essential genes and minimal cell factories after one decade of genome engineering". Microbiology (Reading, England). 160 (Pt 11): 2341–51. PMID 25092907.
- ^ Tu, Z; Wang, L; Xu, M; Zhou, X; Chen, T; Sun, F (21 February 2006). "Further understanding human disease genes by comparing with housekeeping genes and other genes". BMC genomics. 7: 31. PMID 16504025.

- ^ Georgi, B; Voight, BF; Bućan, M (May 2013). "From mouse to human: evolutionary genomics analysis of human orthologs of essential genes". PLoS genetics. 9 (5): e1003484. PMID 23675308.

- ^ Eisenberg, E; Levanon, EY (October 2013). "Human housekeeping genes, revisited". Trends in genetics : TIG. 29 (10): 569–74. PMID 23810203.
- ^ Amsterdam, A; Hopkins, N (September 2006). "Mutagenesis strategies in zebrafish for identifying genes involved in development and disease". Trends in genetics : TIG. 22 (9): 473–8. PMID 16844256.
- ^ "About the HGNC". HGNC Database of Human Gene Names. HUGO Gene Nomenclature Committee. Retrieved 14 May 2015.
- ^ Stanley N. Cohen and Annie C. Y. Chang (1 May 1973). "Recircularization and Autonomous Replication of a Sheared R-Factor DNA Segment in Escherichia coli Transformants — PNAS". Pnas.org. Retrieved 17 July 2010.
- ^ Esvelt, KM.; Wang, HH. (2013). "Genome-scale engineering for systems and synthetic biology". Mol Syst Biol. 9 (1): 641. doi:10.1038/msb.2012.66. PMC 3564264. PMID 23340847.
{{cite journal}}: Cite has empty unknown parameter:|month=(help) - ^ Tan, WS.; Carlson, DF.; Walton, MW.; Fahrenkrug, SC.; Hackett, PB. (2012). "Precision editing of large animal genomes". Adv Genet. Advances in Genetics. 80: 37–97. doi:10.1016/B978-0-12-404742-6.00002-8. ISBN 9780124047426. PMC 3683964. PMID 23084873.
{{cite journal}}: Cite has empty unknown parameter:|month=(help) - ^ Puchta, H.; Fauser, F. (2013). "Gene targeting in plants: 25 years later". Int. J. Dev. Biol. 57 (6–7–8): 629–637. doi:10.1387/ijdb.130194hp.
{{cite journal}}: Cite has empty unknown parameter:|month=(help) - ^ Ran FA, Hsu PD, Wright J, Agarwala V, Scott DA, Zhang F (2013). "Genome engineering using the CRISPR-Cas9 system". Nat Protoc. 8 (11): 2281–308. doi:10.1038/nprot.2013.143. PMC 3969860. PMID 24157548.
- ^ Kittleson, Joshua (2012). "Successes and failures in modular genetic engineering". Current Opinion in Chemical Biology. doi:10.1016/j.cbpa.2012.06.009.
- ^ Berg, P.; Mertz, J. E. (2010). "Personal Reflections on the Origins and Emergence of Recombinant DNA Technology". Genetics. 184 (1): 9–17. doi:10.1534/genetics.109.112144. PMC 2815933. PMID 20061565.
- ^ Austin, Christopher P.; Battey, James F.; Bradley, Allan; Bucan, Maja; Capecchi, Mario; Collins, Francis S.; Dove, William F.; Duyk, Geoffrey; Dymecki, Susan (September 2004). "The Knockout Mouse Project". Nature Genetics. 36 (9): 921–924. doi:10.1038/ng0904-921. ISSN 1061-4036. PMC 2716027. PMID 15340423.
- ^ "A review of current large-scale mouse knockout efforts – Guan – 2010 – genesis – Wiley Online Library". doi.wiley.com.
- ^ Deng C (2007). "In celebration of Dr. Mario R. Capecchi's Nobel Prize". International Journal of Biological Sciences. 3 (7): 417–419. doi:10.7150/ijbs.3.417. PMID 17998949.
Further reading
- Watson, James D.; Baker, Tania A.; Bell, Stephen P.; Gann, Alexander; Levine, Michael; Losick, Richard (2013). Molecular Biology of the Gene (7th ed.). Benjamin Cummings. ISBN 978-0-321-90537-6.
{{cite book}}: Unknown parameter|name-list-format=ignored (|name-list-style=suggested) (help) - Dawkins, Richard (1990). The Selfish Gene. Oxford University Press. ISBN 0-19-286092-5.
{{cite book}}: Unknown parameter|name-list-format=ignored (|name-list-style=suggested) (help) Google Book Search; first published 1976. - Ridley, Matt (1999). Genome: The Autobiography of a Species in 23 Chapters. Fourth Estate. ISBN 0-00-763573-7.
{{cite book}}: Unknown parameter|name-list-format=ignored (|name-list-style=suggested) (help) - Brown, T (2002). Genomes (2nd ed.). New York: Wiley-Liss. ISBN 0-471-25046-5.
External links
- Comparative Toxicogenomics Database
- DNA From The Beginning – a primer on genes and DNA
- Genes And DNA – Introduction to genes and DNA aimed at non-biologist
- Entrez Gene – a searchable database of genes
- IDconverter – converts gene IDs between public databases
- iHOP – Information Hyperlinked over Proteins
- TranscriptomeBrowser – Gene expression profile analysis
- The Protein Naming Utility, a database to identify and correct deficient gene names
- Genes – an Open Access journal
- IMPC (International Mouse Phenotyping Consortium) – Encyclopedia of mammalian gene function
- Global Genes Project – Leading non-profit organization supporting people living with genetic diseases
- ENCODE threads Explorer Characterization of intergenic regions and gene definition. Nature