S phase
This article needs additional citations for verification. (December 2010) |

S phase (Synthesis phase) is the phase of the cell cycle in which DNA is replicated, occurring between G1 phase and G2 phase.[1] Since accurate duplication of the genome is critical to successful cell division, the processes that occur during S-phase are tightly regulated and widely conserved.
Regulation
[edit]Entry into S-phase is controlled by the G1 restriction point (R), which commits cells to the remainder of the cell-cycle if there is adequate nutrients and growth signaling.[2] This transition is essentially irreversible; after passing the restriction point, the cell will progress through S-phase even if environmental conditions become unfavorable.[2]
Accordingly, entry into S-phase is controlled by molecular pathways that facilitate a rapid, unidirectional shift in cell state. In yeast, for instance, cell growth induces accumulation of Cln3 cyclin, which complexes with the cyclin dependent kinase CDK2.[3] The Cln3-CDK2 complex promotes transcription of S-phase genes by inactivating the transcriptional repressor Whi5.[3] Since upregulation of S-phase genes drive further suppression of Whi5, this pathway creates a positive feedback loop that fully commits cells to S-phase gene expression.[3]
A remarkably similar regulatory scheme exists in mammalian cells.[3] Mitogenic signals received throughout G1-phase cause gradual accumulation of cyclin D, which complexes with CDK4/6.[3] Active cyclin D-CDK4/6 complex induces release of E2F transcription factor, which in turn initiates expression of S-phase genes.[3] Several E2F target genes promote further release of E2F, creating a positive feedback loop similar to the one found in yeast.[3]
DNA replication
[edit]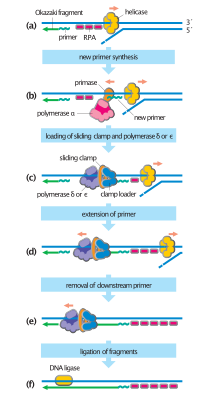
Throughout M phase and G1 phase, cells assemble inactive pre-replication complexes (pre-RC) on replication origins distributed throughout the genome.[4] During S-phase, the cell converts pre-RCs into active replication forks to initiate DNA replication.[4] This process depends on the kinase activity of Cdc7 and various S-phase CDKs, both of which are upregulated upon S-phase entry.[4]
Activation of the pre-RC is a closely regulated and highly sequential process. After Cdc7 and S-phase CDKs phosphorylate their respective substrates, a second set of replicative factors associate with the pre-RC.[4] Stable association encourages MCM helicase to unwind a small stretch of parental DNA into two strands of ssDNA, which in turn recruits replication protein A (RPA), an ssDNA binding protein.[4] RPA recruitment primes the replication fork for loading of replicative DNA polymerases and PCNA sliding clamps.[4] Loading of these factors completes the active replication fork and initiates synthesis of new DNA.
Complete replication fork assembly and activation only occurs on a small subset of replication origins. All eukaryotes possess many more replication origins than strictly needed during one cycle of DNA replication.[5] Redundant origins may increase the flexibility of DNA replication, allowing cells to control the rate of DNA synthesis and respond to replication stress.[5]
Histone synthesis
[edit]Since new DNA must be packaged into nucleosomes to function properly, synthesis of canonical (non-variant) histone proteins occurs alongside DNA replication. During early S-phase, the cyclin E-Cdk2 complex phosphorylates NPAT, a nuclear coactivator of histone transcription.[6] NPAT is activated by phosphorylation and recruits the Tip60 chromatin remodeling complex to the promoters of histone genes.[6] Tip60 activity removes inhibitory chromatin structures and drives a three to ten-fold increase in transcription rate.[1][6]
In addition to increasing transcription of histone genes, S-phase entry also regulates histone production at the RNA level. Instead of polyadenylated tails, canonical histone transcripts possess a conserved 3` stem loop motif that selective binds to Stem Loop Binding Protein (SLBP).[7] SLBP binding is required for efficient processing, export, and translation of histone mRNAs, allowing it to function as a highly sensitive biochemical "switch".[7] During S-phase, accumulation of SLBP acts together with NPAT to drastically increase the efficiency of histone production.[7] However, once S-phase ends, both SLBP and bound RNA are rapidly degraded.[8] This immediately halts histone production and prevents a toxic buildup of free histones.[9]
Nucleosome replication
[edit]
Free histones produced by the cell during S-phase are rapidly incorporated into new nucleosomes. This process is closely tied to the replication fork, occurring immediately in “front” and “behind” the replication complex. Translocation of MCM helicase along the leading strand disrupts parental nucleosome octamers, resulting in the release of H3-H4 and H2A-H2B subunits.[10] Reassembly of nucleosomes behind the replication fork is mediated by chromatin assembly factors (CAFs) that are loosely associated with replication proteins.[4][11] Though not fully understood, the reassembly does not appear to utilize the semi-conservative scheme seen in DNA replication.[11] Labeling experiments indicate that nucleosome duplication is predominantly conservative.[11][10] The paternal H3-H4 core nucleosome remains completely segregated from newly synthesized H3-H4, resulting in the formation of nucleosomes that either contain exclusively old H3-H4 or exclusively new H3-H4.[10][11] “Old” and “new” histones are assigned to each daughter strand semi-randomly, resulting in equal division of regulatory modifications.[10]
Reestablishment of chromatin domains
[edit]Immediately after division, each daughter chromatid only possesses half the epigenetic modifications present in the paternal chromatid.[10] The cell must use this partial set of instructions to re-establish functional chromatin domains before entering mitosis.
For large genomic regions, inheritance of old H3-H4 nucleosomes is sufficient for accurate re-establishment of chromatin domains.[10] Polycomb Repressive Complex 2 (PRC2) and several other histone-modifying complexes can "copy" modifications present on old histones onto new histones.[10] This process amplifies epigenetic marks and counters the dilutive effect of nucleosome duplication.[10]
However, for small domains approaching the size of individual genes, old nucleosomes are spread too thinly for accurate propagation of histone modifications.[10] In these regions, chromatin structure is probably controlled by incorporation of histone variants during nucleosome reassembly.[10] The close correlation seen between H3.3/H2A.Z and transcriptionally active regions lends support to this proposed mechanism.[10] Unfortunately, a causal relationship has yet to be proven.[10]
DNA damage checkpoints
[edit]During S-phase, the cell continuously scrutinizes its genome for abnormalities. Detection of DNA damage induces activation of three canonical S-phase "checkpoint pathways" that delay or arrest further cell cycle progression:[12]
- The Replication Checkpoint detects stalled replication forks by integrating signals from RPA, ATR Interacting Protein (ATRIP), and RAD17.[12] Upon activation, the replication checkpoint upregulates nucleotide biosynthesis and blocks replication initiation from unfired origins.[12] Both of these processes contribute to rescue of stalled forks by increasing the availability of dNTPs.[12]
- The S-M Checkpoint blocks mitosis until the entire genome has been successfully duplicated.[12] This pathway induces arrest by inhibiting the Cyclin-B-CDK1 complex, which gradually accumulates throughout the cell cycle to promote mitotic entry.[12]
- The intra-S Phase Checkpoint detects Double Strand Breaks (DSBs) through activation of ATR and ATM kinases.[12] In addition to facilitating DNA repair, active ATR and ATM stalls cell cycle progression by promoting degradation of CDC25A, a phosphatase that removes inhibitory phosphate residues from CDKs.[12] Homologous recombination, an accurate process for repairing DNA double-strand breaks, is most active in S phase, declines in G2/M and is nearly absent in G1 phase.[13]
In addition to these canonical checkpoints, recent evidence suggests that abnormalities in histone supply and nucleosome assembly can also alter S-phase progression.[14] Depletion of free histones in Drosophila cells dramatically prolongs S-phase and causes permanent arrest in G2-phase.[14] This unique arrest phenotype is not associated with activation of canonical DNA damage pathways, indicating that nucleosome assembly and histone supply may be scrutinized by a novel S-phase checkpoint.[14]
See also
[edit]- S phase index (SPI)
- S-fraction or S-phase fraction (oncology/pathology prognosis)
- Restriction point
References
[edit]- ^ a b David M (2007). The cell cycle : principles of control. Oxford University Press. ISBN 978-0199206100. OCLC 813540567.
- ^ a b Pardee AB, Blagosklonny MV (2013). The Restriction Point of the Cell Cycle. Landes Bioscience.
- ^ a b c d e f g Bertoli C, Skotheim JM, de Bruin RA (August 2013). "Control of cell cycle transcription during G1 and S phases". Nature Reviews. Molecular Cell Biology. 14 (8): 518–28. doi:10.1038/nrm3629. PMC 4569015. PMID 23877564.
- ^ a b c d e f g Takeda DY, Dutta A (April 2005). "DNA replication and progression through S phase". Oncogene. 24 (17): 2827–43. doi:10.1038/sj.onc.1208616. PMID 15838518.
- ^ a b Leonard AC, Méchali M (October 2013). "DNA replication origins". Cold Spring Harbor Perspectives in Biology. 5 (10): a010116. doi:10.1101/cshperspect.a010116. PMC 3783049. PMID 23838439.
- ^ a b c DeRan M, Pulvino M, Greene E, Su C, Zhao J (January 2008). "Transcriptional activation of histone genes requires NPAT-dependent recruitment of TRRAP-Tip60 complex to histone promoters during the G1/S phase transition". Molecular and Cellular Biology. 28 (1): 435–47. doi:10.1128/MCB.00607-07. PMC 2223310. PMID 17967892.
- ^ a b c Marzluff WF, Koreski KP (October 2017). "Birth and Death of Histone mRNAs". Trends in Genetics. 33 (10): 745–759. doi:10.1016/j.tig.2017.07.014. PMC 5645032. PMID 28867047.
- ^ Whitfield ML, Zheng LX, Baldwin A, Ohta T, Hurt MM, Marzluff WF (June 2000). "Stem-loop binding protein, the protein that binds the 3' end of histone mRNA, is cell cycle regulated by both translational and posttranslational mechanisms". Molecular and Cellular Biology. 20 (12): 4188–98. doi:10.1128/MCB.20.12.4188-4198.2000. PMC 85788. PMID 10825184.
- ^ Ma Y, Kanakousaki K, Buttitta L (2015). "How the cell cycle impacts chromatin architecture and influences cell fate". Frontiers in Genetics. 6: 19. doi:10.3389/fgene.2015.00019. PMC 4315090. PMID 25691891.
- ^ a b c d e f g h i j k l Ramachandran S, Henikoff S (August 2015). "Replicating Nucleosomes". Science Advances. 1 (7): e1500587. Bibcode:2015SciA....1E0587R. doi:10.1126/sciadv.1500587. PMC 4530793. PMID 26269799.
- ^ a b c d Annunziato AT (April 2005). "Split decision: what happens to nucleosomes during DNA replication?". The Journal of Biological Chemistry. 280 (13): 12065–8. doi:10.1074/jbc.R400039200. PMID 15664979.
- ^ a b c d e f g h Bartek J, Lukas C, Lukas J (October 2004). "Checking on DNA damage in S phase". Nature Reviews. Molecular Cell Biology. 5 (10): 792–804. doi:10.1038/nrm1493. PMID 15459660. S2CID 33560392.
- ^ Mao Z, Bozzella M, Seluanov A, Gorbunova V (September 2008). "DNA repair by nonhomologous end joining and homologous recombination during cell cycle in human cells". Cell Cycle. 7 (18): 2902–6. doi:10.4161/cc.7.18.6679. PMC 2754209. PMID 18769152.
- ^ a b c Günesdogan U, Jäckle H, Herzig A (September 2014). "Histone supply regulates S phase timing and cell cycle progression". eLife. 3: e02443. doi:10.7554/eLife.02443. PMC 4157229. PMID 25205668.
