Defective interfering particle
Defective interfering particles (DIPs), also known as defective interfering viruses, are spontaneously generated virus mutants in which a critical portion of the particle's genome has been lost due to defective replication or non-homologous recombination.[2][3] The mechanism of their formation is presumed to be as a result of template-switching during replication of the viral genome, although non-replicative mechanisms involving direct ligation of genomic RNA fragments have also been proposed.[4][5] DIPs are derived from and associated with their parent virus, and particles are classed as DIPs if they are rendered non-infectious due to at least one essential gene of the virus being lost or severely damaged as a result of the defection.[4] A DIP can usually still penetrate host cells, but requires another fully functional virus particle (the 'helper' virus) to co-infect a cell with it, in order to provide the lost factors.[6][7]
DIPs were first observed as early as the 1950s by Von Magnus and Schlesinger, both working with influenza viruses.[8] However, direct evidence for DIPs was only found in the 1960s by Hackett who noticed presence of ‘stumpy’ particles of vesicular stomatitis virus in electron micrographs[9] and the formalization of DIPs terminology was in 1970 by Huang and Baltimore.[10] DIPs can occur within nearly every class of both DNA and RNA viruses both in clinical and laboratory settings including poliovirus, SARS coronavirus, measles, alphaviruses, respiratory syncytial virus and influenza virus.[11][12][13][14][15][16][17][18]
Defection
[edit]DIPs are a naturally occurring phenomenon that can be recreated under experimental conditions in the lab and can also be synthesized for experimental use. They are spontaneously produced by error-prone viral replication, something particularly prevalent in RNA viruses over DNA viruses due to the enzyme used (replicase, or RNA-dependent RNA polymerase.)[4][19] DI genomes typically retain the termini sequences needed for recognition by viral polymerases, and sequences for packaging of their genome into new particles, but little else.[20][21] The size of the genomic deletion event can vary greatly, with one such example in a DIP derived from rabies virus exhibiting a 6.1 kb deletion.[22] In another example, the size of several DI-DNA plant virus genomes varied from one tenth of the size of the original genome to one half.[23]
Interference
[edit]The particles are considered interfering when they affect the function of the parent virus through competitive inhibition[4] during coinfection. In other words, defective and non-defective viruses replicate simultaneously, but when defective particles increase, the amount of replicated non-defective virus is decreased. The extent of interference depends on the type and size of defection in the genome; large deletions of genomic data allow rapid replication of the defective genome.[20] In SARS-CoV-2, synthetic DIPs made by removing 90% of the genome replicate three times faster than the virus.[24] During the coinfection of a host cell, a critical ratio will eventually be reached in which more viral factors are being used to produce the non-infectious DIPs than infectious particles.[20] Defective particles and defective genomes have also been demonstrated to stimulate the host innate immune responses and their presence during a viral infection correlates with the strength of the antiviral response.[11] However, in some viruses such as SARS-CoV-2, the effect of competitive inhibition by interfering particles reduces viral-mediated innate immune responses and inflammation producing a therapeutic effect.[25]
This interfering nature is becoming more and more important for research on virus therapies.[26][27] It is thought that because of their specificity, DIPs will be targeted to sites of infection. In one example, scientists have used DIPs to create "protecting viruses", which attenuated the pathogenicity of an influenza A infection in mice, through inducing an interferon response, to a point that it was no longer lethal.[28] For SARS-CoV-2, the first synthetic DIPs were made in 2020 [24] and the interference effect was used to generate therapeutic interfering particles (TIPs) that reduced pathogenesis and protected hamsters from serious disease.[29]
Pathogenesis
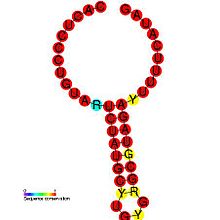
[edit]DIPs have been shown to play a role in pathogenesis of certain viruses. One study demonstrates the relationship between a pathogen and its defective variant, showing how regulation of DI production allowed the virus to attenuate its own infectious replication, decreasing viral load and thus enhance its parasitic efficiency by preventing the host from dying too fast.[30] This also provides the virus with more time to spread and infect new hosts. DIP generation is regulated within viruses: the Coronavirus SL-III cis-acting replication element (shown in the image) is a higher-order genomic structure implicated in the mediation of DIP production in bovine coronavirus, with apparent homologs detected in other coronavirus groups.[1] A more in-depth introduction can be found in Alice Huang and David Baltimore's work from 1970.[10]
Types of defective RNA genomes
[edit]- Deletions defections are when a fragment of the template is skipped. Examples of this type of defection can be found in tomato spotted wilt virus and Flock House virus.[31][21]
- Snapbacks defections are when replicase transcribes part of one strand then uses this new strand as a template. The result of this can produce a hairpin. Snapback defections have been observed in vesicular stomatitis virus.[32]
- Panhandle defections are when the polymerase carries a partially made strand and then switches back to transcribe the 5' end, forming the panhandle shape. Panhandle defections are found in influenza viruses.[33]
- Compound defections are when both a deletion and snapback defection happens together.
- Mosaic or complex DI genome, in which the various regions may come from the same helper virus genome but in the wrong order; from different helper genome segments, or could include segments of host RNA. Duplications may also occur.[3]
Research
[edit]Research has been conducted by virologists to learn more about the interference in infection of host cells and how DI genomes could potentially work as immunostimulatory antiviral agents.[3] Another branch of research has pursued the concept of engineering DIPs into antiviral therapeutic interfering particles (TIPs),[34] a purely theoretical concept until recently.[35] A 2014 article describes the pre-clinical work to test the immunostimulatory effectiveness of a DIP against influenza viruses by inducing the innate antiviral immune responses (i.e., interferon).[36] Subsequent work tested the pre-clinical efficacy of TIPs against HIV[37] and SARS-CoV-2.[24][25] DI-RNAs have also been found to aid in the infection of fungi via viruses of the family Partitiviridae for the first time, which makes room for more interdisciplinary work.[19]
Several tools as ViReMa[38] and DI-tector[39] have been developed to help to detect defective viral genomes in next-generation sequencing data and high-throughput approaches, such as random-deletion library sequencing (RanDeL-Seq),[40] allow rational mapping of the viral genetic elements that are required for DI-particle propagation.
References
[edit]- ^ a b Raman S, Bouma P, Williams GD, Brian DA (June 2003). "Stem-loop III in the 5' untranslated region is a cis-acting element in bovine coronavirus defective interfering RNA replication". Journal of Virology. 77 (12): 6720–6730. doi:10.1128/JVI.77.12.6720-6730.2003. PMC 156170. PMID 12767992.
- ^ White KA, Morris TJ (January 1994). "Nonhomologous RNA recombination in tombusviruses: generation and evolution of defective interfering RNAs by stepwise deletions". Journal of Virology. 68 (1): 14–24. doi:10.1128/JVI.68.1.14-24.1994. PMC 236259. PMID 8254723.
- ^ a b c d Pathak KB, Nagy PD (December 2009). "Defective Interfering RNAs: Foes of Viruses and Friends of Virologists". Viruses. 1 (3): 895–919. doi:10.3390/v1030895. PMC 3185524. PMID 21994575.
- ^ Gmyl AP, Belousov EV, Maslova SV, Khitrina EV, Chetverin AB, Agol VI (November 1999). "Nonreplicative RNA recombination in poliovirus". Journal of Virology. 73 (11): 8958–8965. doi:10.1128/JVI.73.11.8958-8965.1999. PMC 112927. PMID 10516001.
- ^ Makino S, Shieh CK, Soe LH, Baker SC, Lai MM (October 1988). "Primary structure and translation of a defective interfering RNA of murine coronavirus". Virology. 166 (2): 550–560. doi:10.1016/0042-6822(88)90526-0. PMC 7131284. PMID 2845661.
- ^ Palmer SR, Soulsby L, Torgerson P, Brown DW, eds. (2011). Oxford Textbook of Zoonoses: Biology, Clinical Practice, and Public Health Control. OUP Oxford. pp. 399–400. doi:10.1093/med/9780198570028.001.0001. ISBN 978-0-19-857002-8.
- ^ Gard S, Von Magnus P, Svedmyr A, Birch-Andersen A (October 1952). "Studies on the sedimentation of influenza virus". Archiv für die Gesamte Virusforschung. 4 (5): 591–611. doi:10.1007/BF01242026. PMID 14953289. S2CID 21838623.
- ^ Hackett AJ (September 1964). "A possible morphologic basis for the autointerference phenomenon in vesicular stomatitis virus". Virology. 24 (1): 51–59. doi:10.1016/0042-6822(64)90147-3. PMID 14208902.
- ^ a b Huang AS, Baltimore D (April 1970). "Defective viral particles and viral disease processes". Nature. 226 (5243): 325–327. Bibcode:1970Natur.226..325H. doi:10.1038/226325a0. PMID 5439728. S2CID 4184206.
- ^ a b Sun Y, Jain D, Koziol-White CJ, Genoyer E, Gilbert M, Tapia K, et al. (September 2015). "Immunostimulatory Defective Viral Genomes from Respiratory Syncytial Virus Promote a Strong Innate Antiviral Response during Infection in Mice and Humans". PLOS Pathogens. 11 (9): e1005122. doi:10.1371/journal.ppat.1005122. PMC 4559413. PMID 26336095.
- ^ Dimmock NJ, Dove BK, Scott PD, Meng B, Taylor I, Cheung L, et al. (2012). "Cloned defective interfering influenza virus protects ferrets from pandemic 2009 influenza A virus and allows protective immunity to be established". PLOS ONE. 7 (12): e49394. Bibcode:2012PLoSO...749394D. doi:10.1371/journal.pone.0049394. PMC 3521014. PMID 23251341.
- ^ Saira K, Lin X, DePasse JV, Halpin R, Twaddle A, Stockwell T, et al. (July 2013). "Sequence analysis of in vivo defective interfering-like RNA of influenza A H1N1 pandemic virus". Journal of Virology. 87 (14): 8064–8074. doi:10.1128/JVI.00240-13. PMC 3700204. PMID 23678180.
- ^ Petterson E, Guo TC, Evensen Ø, Mikalsen AB (November 2016). "Experimental piscine alphavirus RNA recombination in vivo yields both viable virus and defective viral RNA". Scientific Reports. 6: 36317. Bibcode:2016NatSR...636317P. doi:10.1038/srep36317. PMC 5090867. PMID 27805034.
- ^ Cattaneo R, Schmid A, Eschle D, Baczko K, ter Meulen V, Billeter MA (October 1988). "Biased hypermutation and other genetic changes in defective measles viruses in human brain infections". Cell. 55 (2): 255–265. doi:10.1016/0092-8674(88)90048-7. PMC 7126660. PMID 3167982.
- ^ Makino S, Yokomori K, Lai MM (December 1990). "Analysis of efficiently packaged defective interfering RNAs of murine coronavirus: localization of a possible RNA-packaging signal". Journal of Virology. 64 (12): 6045–6053. doi:10.1128/JVI.64.12.6045-6053.1990. PMC 248778. PMID 2243386.
- ^ Lundquist RE, Sullivan M, Maizel JV (November 1979). "Characterization of a new isolate of poliovirus defective interfering particles". Cell. 18 (3): 759–769. doi:10.1016/0092-8674(79)90129-6. PMID 229964. S2CID 35964939.
- ^ Stauffer Thompson KA, Rempala GA, Yin J (April 2009). "Multiple-hit inhibition of infection by defective interfering particles". The Journal of General Virology. 90 (Pt 4): 888–899. doi:10.1099/vir.0.005249-0. PMC 2889439. PMID 19264636.
- ^ a b Chiba S, Lin YH, Kondo H, Kanematsu S, Suzuki N (February 2013). "Effects of defective interfering RNA on symptom induction by, and replication of, a novel partitivirus from a phytopathogenic fungus, Rosellinia necatrix". Journal of Virology. 87 (4): 2330–2341. doi:10.1128/JVI.02835-12. PMC 3571465. PMID 23236074.
- ^ a b c Dimmock NJ, Easton AJ, Leppard KN, eds. (2015). "Innate and intrinsic immunity". Introduction to Modern Virology. John Wiley & Sons. pp. 201–217. ISBN 978-1-119-09453-1.
- ^ a b Resende R, de Haan P, van de Vossen E, de Avila AC, Goldbach R, Peters D (October 1992). "Defective interfering L RNA segments of tomato spotted wilt virus retain both virus genome termini and have extensive internal deletions". The Journal of General Virology. 73 (10): 2509–2516. doi:10.1099/0022-1317-73-10-2509. PMID 1402797.
- ^ Conzelmann KK, Cox JH, Thiel HJ (October 1991). "An L (polymerase)-deficient rabies virus defective interfering particle RNA is replicated and transcribed by heterologous helper virus L proteins". Virology. 184 (2): 655–663. doi:10.1016/0042-6822(91)90435-e. PMID 1887588.
- ^ Patil BL, Dasgupta I (2006). "Defective Interfering Dnas of Plant Viruses". Critical Reviews in Plant Sciences. 25 (1): 47–64. Bibcode:2006CRvPS..25...47P. doi:10.1080/07352680500391295. S2CID 85790514.
- ^ a b c Yao S, Narayanan A, Majowicz S, Jose J, Archetti M (2020-11-23). "A Synthetic Defective Interfering SARS-CoV-2": 2020.11.22.393587. doi:10.1101/2020.11.22.393587.
{{cite journal}}: Cite journal requires|journal=(help) - ^ a b Chaturvedi S, Vasen G, Pablo M, Chen X, Beutler N, Kumar A, et al. (December 2021). "Identification of a therapeutic interfering particle-A single-dose SARS-CoV-2 antiviral intervention with a high barrier to resistance". Cell. 184 (25): 6022–6036.e18. doi:10.1016/j.cell.2021.11.004. PMC 8577993. PMID 34838159.
- ^ Weinberger LS, Schaffer DV, Arkin AP (September 2003). "Theoretical design of a gene therapy to prevent AIDS but not human immunodeficiency virus type 1 infection". Journal of Virology. 77 (18): 10028–10036. doi:10.1128/jvi.77.18.10028-10036.2003. PMC 224590. PMID 12941913.
- ^ Thompson KA, Yin J (September 2010). "Population dynamics of an RNA virus and its defective interfering particles in passage cultures". Virology Journal. 7: 257. doi:10.1186/1743-422X-7-257. PMC 2955718. PMID 20920247.
- ^ Easton AJ, Scott PD, Edworthy NL, Meng B, Marriott AC, Dimmock NJ (March 2011). "A novel broad-spectrum treatment for respiratory virus infections: influenza-based defective interfering virus provides protection against pneumovirus infection in vivo" (PDF). Vaccine. 29 (15): 2777–2784. doi:10.1016/j.vaccine.2011.01.102. PMID 21320545.
- ^ Villanueva MT (December 2021). "Interfering viral-like particles inhibit SARS-CoV-2 replication". Nature Reviews. Drug Discovery. 21 (1): d41573–021–00205-5. doi:10.1038/d41573-021-00205-5. PMID 34873320. S2CID 244935707.
- ^ Lukhovitskaya NI, Thaduri S, Garushyants SK, Torrance L, Savenkov EI (June 2013). "Deciphering the mechanism of defective interfering RNA (DI RNA) biogenesis reveals that a viral protein and the DI RNA act antagonistically in virus infection". Journal of Virology. 87 (11): 6091–6103. doi:10.1128/JVI.03322-12. PMC 3648117. PMID 23514891.
- ^ Jaworski E, Routh A (May 2017). "Parallel ClickSeq and Nanopore sequencing elucidates the rapid evolution of defective-interfering RNAs in Flock House virus". PLOS Pathogens. 13 (5): e1006365. doi:10.1371/journal.ppat.1006365. PMC 5435362. PMID 28475646.
- ^ Schubert M, Lazzarini RA (February 1981). "Structure and origin of a snapback defective interfering particle RNA of vesicular stomatitis virus". Journal of Virology. 37 (2): 661–672. doi:10.1128/JVI.37.2.661-672.1981. PMC 171054. PMID 6261012.
- ^ Fodor E, Pritlove DC, Brownlee GG (June 1994). "The influenza virus panhandle is involved in the initiation of transcription". Journal of Virology. 68 (6): 4092–4096. doi:10.1128/JVI.68.6.4092-4096.1994. PMC 236924. PMID 8189550.
- ^ Metzger VT, Lloyd-Smith JO, Weinberger LS (March 2011). "Autonomous targeting of infectious superspreaders using engineered transmissible therapies". PLOS Computational Biology. 7 (3): e1002015. Bibcode:2011PLSCB...7E2015M. doi:10.1371/journal.pcbi.1002015. PMC 3060167. PMID 21483468.
- ^ Gladstone Institutes. "A New Class of Antiviral Therapy Could Treat COVID-19". www.prnewswire.com (Press release). Retrieved 2021-12-28.
- ^ Dimmock NJ, Easton AJ (May 2014). "Defective interfering influenza virus RNAs: time to reevaluate their clinical potential as broad-spectrum antivirals?". Journal of Virology. 88 (10): 5217–5227. doi:10.1128/JVI.03193-13. PMC 4019098. PMID 24574404.
- ^ Tanner EJ, Jung SY, Glazier J, Thompson C, Zhou Y, Martin B, Son HI, Riley JL, Weinberger LS (2019-10-30). "Discovery and Engineering of a Therapeutic Interfering Particle (TIP): a combination self-renewing antiviral" (PDF). doi:10.1101/820456. S2CID 208600143.
{{cite journal}}: Cite journal requires|journal=(help) - ^ Routh A, Johnson JE (January 2014). "Discovery of functional genomic motifs in viruses with ViReMa-a Virus Recombination Mapper-for analysis of next-generation sequencing data". Nucleic Acids Research. 42 (2): e11. doi:10.1093/nar/gkt916. PMC 3902915. PMID 24137010.
- ^ Beauclair G, Mura M, Combredet C, Tangy F, Jouvenet N, Komarova AV (October 2018). "DI-tector: defective interfering viral genomes' detector for next-generation sequencing data". RNA. 24 (10): 1285–1296. doi:10.1261/rna.066910.118. PMC 6140465. PMID 30012569.
- ^ Notton T, Glazier JJ, Saykally VR, Thompson CE, Weinberger LS (January 2021). "RanDeL-Seq: a High-Throughput Method to Map Viral cis- and trans-Acting Elements". mBio. 12 (1): e01724–20. doi:10.1128/mBio.01724-20. PMC 7845639. PMID 33468683.
