Aspartic protease
| Eukaryotic aspartyl protease | |||||||||
|---|---|---|---|---|---|---|---|---|---|
 Structure of the dimeric aspartic protease HIV protease in white and grey, with peptide substrate in black and active site aspartate side chains in red. (PDB: 1KJF) | |||||||||
| Identifiers | |||||||||
| Symbol | Asp | ||||||||
| Pfam | PF00026 | ||||||||
| InterPro | IPR001461 | ||||||||
| PROSITE | PDOC00128 | ||||||||
| SCOP2 | 1mpp / SCOPe / SUPFAM | ||||||||
| OPM superfamily | 100 | ||||||||
| OPM protein | 1lyb | ||||||||
| Membranome | 315 | ||||||||
| |||||||||
Aspartic proteases (also "aspartyl proteases", "aspartic endopeptidases") are a catalytic type of protease enzymes that use an activated water molecule bound to one or more aspartate residues for catalysis of their peptide substrates. In general, they have two highly conserved aspartates in the active site and are optimally active at acidic pH. Nearly all known aspartyl proteases are inhibited by pepstatin.[1]
Aspartic endopeptidases EC 3.4.23. of vertebrate, fungal and retroviral origin have been characterised.[2] More recently, aspartic endopeptidases associated with the processing of bacterial type 4 prepilin[3] and archaean preflagellin have been described.[4][5]
Eukaryotic aspartic proteases include pepsins, cathepsins, and renins. They have a two-domain structure, arising from ancestral duplication. Retroviral and retrotransposon proteases (retroviral aspartyl proteases) are much smaller and appear to be homologous to a single domain of the eukaryotic aspartyl proteases. Each domain contributes a catalytic Asp residue, with an extended active site cleft localized between the two lobes of the molecule. One lobe has probably evolved from the other through a gene duplication event in the distant past. In modern-day enzymes, although the three-dimensional structures are very similar, the amino acid sequences are more divergent, except for the catalytic site motif, which is very conserved. The presence and position of disulfide bridges are other conserved features of aspartic peptidases.
Catalytic mechanism
[edit]
Aspartyl proteases are a highly specific family of proteases – they tend to cleave dipeptide bonds that have hydrophobic residues as well as a beta-methylene group. Unlike serine or cysteine proteases these proteases do not form a covalent intermediate during cleavage. Proteolysis therefore occurs in a single step.
While a number of different mechanisms for aspartyl proteases have been proposed, the most widely accepted is a general acid-base mechanism involving coordination of a water molecule between the two highly conserved aspartate residues.[6][7] One aspartate activates the water by abstracting a proton, enabling the water to perform a nucleophilic attack on the carbonyl carbon of the substrate scissile bond, generating a tetrahedral oxyanion intermediate stabilized by hydrogen-bonding with the second aspartic acid. Rearrangement of this intermediate leads to protonation of the scissile amide which results in the splitting of the substrate peptide into two product peptides.
Inhibition
[edit]Pepstatin is an inhibitor of aspartate proteases.[1]
Classification
[edit]Five superfamilies (clans) of aspartic proteases are known, each representing an independent evolution of the same active site and mechanisms. Each superfamily contains several families with similar sequences. The MEROPS classification systematic names these clans alphabetically.
- Clan AA (e.g. Pepsin family)
- Clan AC (e.g. Signal peptidase II family)
- Clan AD (e.g. Presenilin family)
- Clan AE (e.g. GPR endopeptidase family)
- Clan AF (e.g. Omptin family)
Propeptide
[edit]| A1_Propeptide | |||||||||
|---|---|---|---|---|---|---|---|---|---|
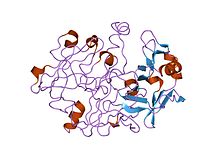 crystal and molecular structures of human progastricsin at 1.62 angstroms resolution | |||||||||
| Identifiers | |||||||||
| Symbol | A1_Propeptide | ||||||||
| Pfam | PF07966 | ||||||||
| InterPro | IPR012848 | ||||||||
| |||||||||
Many eukaryotic aspartic endopeptidases (MEROPS peptidase family A1) are synthesised with signal and propeptides. The animal pepsin-like endopeptidase propeptides form a distinct family of propeptides, which contain a conserved motif approximately 30 residues long. In pepsinogen A, the first 11 residues of the mature pepsin sequence are displaced by residues of the propeptide. The propeptide contains two helices that block the active site cleft, in particular the conserved Asp11 residue, in pepsin, hydrogen bonds to a conserved Arg residue in the propeptide. This hydrogen bond stabilises the propeptide conformation and is probably responsible for triggering the conversion of pepsinogen to pepsin under acidic conditions.[8][9]
Examples
[edit]Human
[edit]- BACE1, BACE2
- Cathepsin D
- Cathepsin E
- Chymosin (or "rennin")
- Napsin-A
- Nepenthesin
- Pepsin
- Presenilin
- Renin
Human proteins containing this domain
[edit]BACE1; BACE2; CTSD; CTSE; NAPSA; PGA5; PGC; REN;
Other organisms
[edit]- HIV-1 protease – a major drug-target for treatment of HIV
- Plasmepsin – a group of aspartyl proteases found in the Malaria-causing parasite Plasmodium
See also
[edit]References
[edit]- ^ a b Fusek M, Mares M, Vetvicka V (2013-01-01). "Chapter 8 - Cathepsin D". In Rawlings ND, Salvesen G (eds.). Handbook of Proteolytic Enzymes (Third ed.). Academic Press. pp. 54–63. doi:10.1016/b978-0-12-382219-2.00008-9. ISBN 978-0-12-382219-2.
- ^ Szecsi PB (1992). "The aspartic proteases". Scandinavian Journal of Clinical and Laboratory Investigation. Supplementum. 210: 5–22. doi:10.3109/00365519209104650. PMID 1455179.
- ^ LaPointe CF, Taylor RK (January 2000). "The type 4 prepilin peptidases comprise a novel family of aspartic acid proteases". The Journal of Biological Chemistry. 275 (2): 1502–10. doi:10.1074/jbc.275.2.1502. PMID 10625704.
- ^ Ng SY, Chaban B, Jarrell KF (2006). "Archaeal flagella, bacterial flagella and type IV pili: a comparison of genes and posttranslational modifications". Journal of Molecular Microbiology and Biotechnology. 11 (3–5): 167–91. doi:10.1159/000094053. PMID 16983194. S2CID 30386932.
- ^ Bardy SL, Jarrell KF (November 2003). "Cleavage of preflagellins by an aspartic acid signal peptidase is essential for flagellation in the archaeon Methanococcus voltae". Molecular Microbiology. 50 (4): 1339–47. doi:10.1046/j.1365-2958.2003.03758.x. PMID 14622420. S2CID 11913649.
- ^ a b Suguna K, Padlan EA, Smith CW, Carlson WD, Davies DR (October 1987). "Binding of a reduced peptide inhibitor to the aspartic proteinase from Rhizopus chinensis: implications for a mechanism of action". Proceedings of the National Academy of Sciences of the United States of America. 84 (20): 7009–13. Bibcode:1987PNAS...84.7009S. doi:10.1073/pnas.84.20.7009. PMC 299218. PMID 3313384.
- ^ Brik A, Wong CH (January 2003). "HIV-1 protease: mechanism and drug discovery". Organic & Biomolecular Chemistry. 1 (1): 5–14. doi:10.1039/b208248a. PMID 12929379.
- ^ Hartsuck JA, Koelsch G, Remington SJ (May 1992). "The high-resolution crystal structure of porcine pepsinogen". Proteins. 13 (1): 1–25. doi:10.1002/prot.340130102. PMID 1594574. S2CID 43462673.
- ^ Sielecki AR, Fujinaga M, Read RJ, James MN (June 1991). "Refined structure of porcine pepsinogen at 1.8 A resolution". Journal of Molecular Biology. 219 (4): 671–92. doi:10.1016/0022-2836(91)90664-R. PMID 2056534.
External links
[edit]- The MEROPS online database for peptidases and their inhibitors: Aspartic Peptidases
- Aspartic+Endopeptidases at the U.S. National Library of Medicine Medical Subject Headings (MeSH)
- MEROPS family A1
