Cofactor (biochemistry): Difference between revisions
TimVickers (talk | contribs) →External links: add |
TimVickers (talk | contribs) |
||
| Line 60: | Line 60: | ||
| '''Cofactor''' || '''Vitamin''' || '''Additional component''' || '''Chemical group(s) transferred''' || '''Distribution''' |
| '''Cofactor''' || '''Vitamin''' || '''Additional component''' || '''Chemical group(s) transferred''' || '''Distribution''' |
||
|- |
|- |
||
|[[Thiamine diphosphate]] <ref |
|[[Thiamine diphosphate]] <ref>{{cite journal |author=Frank RA, Leeper FJ, Luisi BF |title=Structure, mechanism and catalytic duality of thiamine-dependent enzymes |journal=Cell. Mol. Life Sci. |volume=64 |issue=7-8 |pages=892–905 |year=2007 |pmid=17429582 |doi=10.1007/s00018-007-6423-5}}</ref>|| [[Thiamine]] (B<sub>1</sub>) || None ||2-carbon groups, α cleavage || [[Bacteria]], [[archaea]] and [[eukaryote]]s |
||
|- |
|- |
||
| [[Nicotinamide adenine dinucleotide|NAD<sup>+</sup>]] and [[Nicotinamide adenine dinucleotide phosphate|NADP<sup>+</sup>]] <ref name=Pollak/> || [[Niacin]] (B<sub>3</sub>) || ADP || [[Electron]]s || [[Bacteria]], [[archaea]] and [[eukaryote]]s |
| [[Nicotinamide adenine dinucleotide|NAD<sup>+</sup>]] and [[Nicotinamide adenine dinucleotide phosphate|NADP<sup>+</sup>]] <ref name=Pollak/> || [[Niacin]] (B<sub>3</sub>) || ADP || [[Electron]]s || [[Bacteria]], [[archaea]] and [[eukaryote]]s |
||
Revision as of 01:07, 11 March 2009
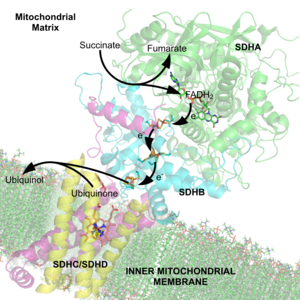
A cofactor is a non-protein chemical compound that is bound (either tightly or loosely) to an enzyme and is required for catalysis.[1] They can be considered "helper molecules/ions" that assist in biochemical transformations. Certain substances such as water and various abundant ions may be bound tightly by enzymes, but are not considered to be cofactors since they are ubiquitous and rarely limiting. Some sources limit the use of the term "cofactor" to inorganic substances.[2][3] An inactive enzyme, without the cofactor is called an apoenzyme, while the complete enzyme with cofactor is the holoenzyme.[4]
Some enzymes or enzyme complexes require several cofactors. A good example is the multienzyme complex pyruvate dehydrogenase.[5] This enzyme complex at the junction of glycolysis and the citric acid cycle requires five organic cofactors and one metal ion : loosely bound thiamine diphosphate (ThDP), covalently bound lipoamide and flavin adenine dinucleotide (FAD), and the cosubstrates nicotinamide adenine dinucleotide (NAD+) and coenzyme A (CoA) and a metal ion (Mg2+).
Organic cofactors are often vitamins or are made from vitamins. Many contain the nucleotide adenosine monophosphate (AMP) as part of their structures, such as ATP, coenzyme A, FAD and NAD+. This common structure may reflect a common evolutionary origin as part of ribozymes in an ancient RNA world. It has been suggested that the AMP part of the molecule can be considered a kind a "handle" by which the enzyme can "grasp" the coenzyme to switch it between different catalytic centers.[6]
Classification
Cofactors can be divided into two broad groups: organic cofactors, such as flavin or heme, and inorganic cofactors: such as the metal ions Mg2+, Cu+, Mn2+ or iron-sulfur clusters.
Organic cofactors are sometimes further divided into coenzymes and prosthetic groups. The term coenzyme refers specifically to enzymes and as such to the functional properties of a protein. On the other hand "prosthetic group", emphasizes the nature of the binding of a cofactor to a protein (tight or covalent) and thus refers to a structural property. Different sources give slightly different definitions of coenzymes, cofactors and prosthetic groups. Some consider tightly-bound organic molecules as prosthetic groups and not as coenzymes, while others define all non-protein organic molecules needed for enzyme activity as coenzymes, and classify those that are tightly bound as coenzyme prosthetic groups. Unsurprisingly, these terms are often used loosely.
A 1979 letter in Trends in Biochemical Sciences noted the confusion in the literature and the essentially arbitrary distinction made between prosthetic groups and coenzymes and proposed the following scheme. Here, cofactors were defined as an additional substance apart from protein and substrate that is required for enzyme activity and a prosthetic group as a substance that undergoes its whole catalytic cycle attached to a single enzyme molecule. However, the author could not arrive at a single all-encompassing definition of a "coenzyme" and proposed that this term be dropped from use in the literature.[7]
Inorganic
Metal ions
Metal ions are common cofactors. The study of these cofactors falls under the area of bioinorganic chemistry. In nutrition, the list of essential trace elements reflects their role as cofactors. In humans this list commonly includes iron, manganese, cobalt, copper, zinc, selenium, and molybdenum.[8] Although chromium deficiency causes impaired glucose tolerance, no human enzyme that uses this metal as a cofactor has been identified.[9][10] Iodine is also an essential trace element, but this element is used as part of the structure of thyroid hormones rather than as an enzyme cofactor.[11] Calcium is another special case, in that it is required as a component of the human diet, and it is needed for the full activity of many enzymes: such as nitric oxide synthase, protein phosphatases or adenylate kinase, but calcium activates these enzymes in allosteric regulation, often binding to these enzymes in a complex with calmodulin.[12] Calcium is therefore a cell signaling molecule, and not usually considered as a cofactor of the enzymes it regulates.[13]
Other organisms require additional metals as enzyme cofactors, such as vanadium in the nitrogenase of the nitrogen-fixing bacteria of the genus Azotobacter,[14] tungsten in the aldehyde ferredoxin oxidoreductase of the thermophilic archaean Pyrococcus furiosus,[15] and even cadmium in the carbonic anhydrase from the marine diatom Thalassiosira weissflogii.[16][17]
In many cases, the cofactor includes both an inorganic and organic component. One diverse set of examples are the haem proteins, which consists of a porphyrin ring coordinated to iron.

Iron-sulfur clusters
Iron-sulfur clusters are complexes of iron and sulfur atoms held within proteins by cysteinyl residues. They play both structural and functional roles, including electron transfer, redox sensing, and as structural modules.
Organic
Organic cofactors are small organic molecules (typically a molecular mass less than 1000 Da) that can be either loosely or tightly bound to the enzyme and directly participate in the reaction.[18][4][19][20] In the latter case, when it is difficult to remove without denaturing the enzyme, it can be called a prosthetic group. It is important to emphasize that there is no sharp division between loosely and tightly bound cofactors.[4] Indeed, many, such as NAD+ can be tightly bound in some enzymes, while it is loosely bound in others.[4] Another example is thiamine diphosphate (ThDP) is tightly bound in transketolase or pyruvate decarboxylase, while it is less tightly bound in pyruvate dehydrogenase. Other coenzymes, flavin adenine dinucleotide (FAD), biotin or lipoamide for instance, are covalently bound. Tightly-bound cofactors are generally regenerated during the same reaction cycle, while loosely-bound cofactors can be regenerated in a subsequent reaction catalyzed by a different enzyme. In the latter case, the cofactor can also be considered a substrate or cosubstrate.
Vitamins can serve as precursors to many organic cofactors (e.g. vitamins B1, B2, B6, B12, niacin, folic acid) or as coenzymes themselves (e.g. vitamin C). However, vitamins do have other functions in the body.[21] Many organic cofactors also contain a nucleotide: such as the electron carriers NAD and FAD, or coenzyme A, which carries acyl groups. Most of these cofactors are found in a huge variety of species, and some are universal to all forms of life. An exception to this wide distribution is a group of unique cofactors that evolved in methanogens, which are restricted to this group of archaea.[22]
Vitamins and derivatives
Non-vitamins
| Coenzyme | Chemical group(s) transferred | Distribution |
| Adenosine triphosphate [30] | Phosphate group | Bacteria, archaea and eukaryotes |
| S-Adenosyl methionine [31] | Methyl group | Bacteria, archaea and eukaryotes |
| 3'-Phosphoadenosine-5'-phosphosulfate [32] | Sulfate group | Bacteria, archaea and eukaryotes |
| Coenzyme Q [33] | Electrons | Bacteria, archaea and eukaryotes |
| Tetrahydrobiopterin [34] | Oxygen atom and electrons | Bacteria, archaea and eukaryotes |
| Cytidine triphosphate [35] | Diacylglycerols and lipid head groups | Bacteria, archaea and eukaryotes |
| Nucleotide sugars [36] | Monosaccharides | Bacteria, archaea and eukaryotes |
| Glutathione [37][38] | Electrons | Some bacteria and most eukaryotes |
| Coenzyme M [39][40] | Methyl group | Methanogens |
| Coenzyme B [41] | Electrons | Methanogens |
| Methanofuran [42] | Formyl group | Methanogens |
| Tetrahydromethanopterin [43] | Methyl group | Methanogens |
Cofactors as metabolic intermediates

Metabolism involves a vast array of chemical reactions, but most fall under a few basic types of reactions that involve the transfer of functional groups.[44] This common chemistry allows cells to use a small set of metabolic intermediates to carry chemical groups between different reactions.[45] These group-transfer intermediates are the loosely-bound organic cofactors, often called coenzymes.
Each class of group-transfer reaction is carried out by a particular cofactor, which is the substrate for a set of enzymes that produce it, and a set of enzymes that consume it. An example of this are the dehydrogenases that use nicotinamide adenine dinucleotide (NAD+) as a cofactor. Here, hundreds of separate types of enzymes remove electrons from their substrates and reduce NAD+ to NADH. This reduced cofactor is then a substrate for any of the reductases in the cell that require electrons to reduce their substrates.[24]
These cofactors are therefore continuously recycled as part of metabolism. As an example, the total quantity of ATP in the human body is about 0.1 mole. This ATP is constantly being broken down into ADP, and then converted back into ATP. Thus, at any given time, the total amount of ATP + ADP remains fairly constant. The energy used by human cells requires the hydrolysis of 100 to 150 moles of ATP daily which is around 50 to 75 kg. Typically, a human will use up their body weight of ATP over the course of the day.[46] This means that each ATP molecule is recycled 1000 to 1500 times daily.
Evolution
Organic cofactors, such as ATP and NADH, are present in all known forms of life and form a core part of metabolism. Such universal conservation indicates that these molecules evolved very early in the development of living things.[47] At least some of the current set of cofactors may therefore have been present in the last universal ancestor, which lived about 4 billion years ago.[48][49]
Organic cofactors may have been present even earlier in the history of life on Earth.[50] Interestingly, the nucleotide adenosine is present in cofactors that catalyse many basic metabolic reactions such as methyl, acyl, and phosphoryl group transfer, as well as redox reactions. This ubiquitous chemical scaffold has therefore been proposed to be a remnant of the RNA world, with early ribozymes evolving to bind a restricted set of nucleotides and related compounds.[51][52] Adenosine-based cofactors are thought to have acted as interchangeable adaptors that allowed enzymes and ribozymes to bind new cofactors through small modifications in existing adenosine-binding domains, which had originally evolved to bind a different cofactor.[6] This process of adapting a pre-evolved structure for a novel use is referred to as exaptation.
History
The first organic cofactor to be discovered was NAD+, which was identified by Arthur Harden and William Youndin 1906.[53] They noticed that adding boiled and filtered yeast extract greatly accelerated alcoholic fermentation in unboiled yeast extracts. They called the unidentified factor responsible for this effect a coferment. Through a long and difficult purification from yeast extracts, this heat-stable factor was identified as a nucleotide sugar phosphate by Hans von Euler-Chelpin.[54] Other cofactors were identified throughout the early 20th century, with ATP being isolated in 1929 by Karl Lohmann,[55] and coenzyme A being discovered in 1945 by Fritz Albert Lipmann.[56]
The functions of these molecules were at first mysterious, but in 1936, Otto Heinrich Warburg identified the function of NAD+ in hydride transfer.[57] This discovery was followed in the early 1940s by the work of Herman Kalckar, who established the link between the oxidation of sugars and the generation of ATP.[58] This confirmed the central role of ATP in energy transfer that had been proposed by Fritz Albert Lipmann in 1941.[59] Later, in 1949, Morris Friedkin and Albert L. Lehninger proved that NAD+ linked metabolic pathways such as the citric acid cycle and the synthesis of ATP.[60]
Non-enzymatic cofactors
The term is used in other areas of biology to refer more broadly to non-protein (or even protein) molecules that either activate, inhibit or are required for the protein to function. For example, ligands such as hormones that bind to and activate receptor proteins are termed cofactors or coactivators, while molecules that inhibit receptor proteins are termed corepressors.
See also
References
- ^ de Bolster, M.W.G. (1997). "Glossary of Terms Used in Bioinorganic Chemistry: Cofactors". International Union of Pure and Applied Chemistry. Retrieved 2007-10-30.
- ^ "coenzymes and cofactors". Retrieved 2007-11-17.
- ^ "Enzyme Cofactors". Retrieved 2007-11-17.
- ^ a b c d e f g h i Metzler DE (2001) Biochemistry. The chemical reactions of living cells, 2nd edition, Harcourt, San Diego.
- ^ Thiamine. Catalytic mechanisms in normal and disease states (2004) (Eds. Jordan F., Patel M.S.), pp 588, Marcel Dekker Inc, New York, Basel.
- ^ a b Denessiouk KA, Rantanen VV, Johnson MS (2001). "Adenine recognition: a motif present in ATP-, CoA-, NAD-, NADP-, and FAD-dependent proteins". Proteins. 44 (3): 282–91. doi:10.1002/prot.1093. PMID 11455601.
{{cite journal}}: CS1 maint: multiple names: authors list (link) - ^ Bryce CFA (1979). "SAM - semantics and misunderstandings". Trends Biochem. Sci. 4: N62.
{{cite journal}}: Unknown parameter|month=ignored (help) - ^ Aggett PJ (1985). "Physiology and metabolism of essential trace elements: an outline". Clin Endocrinol Metab. 14 (3): 513–43. doi:10.1016/S0300-595X(85)80005-0. PMID 3905079.
- ^ Stearns DM (2000). "Is chromium a trace essential metal?". Biofactors. 11 (3): 149–62. PMID 10875302.
- ^ Vincent JB (2000). "The biochemistry of chromium". J. Nutr. 130 (4): 715–8. PMID 10736319.
- ^ Cavalieri RR (1997). "Iodine metabolism and thyroid physiology: current concepts". Thyroid. 7 (2): 177–81. PMID 9133680.
- ^ Clapham DE (2007). "Calcium signaling". Cell. 131 (6): 1047–58. doi:10.1016/j.cell.2007.11.028. PMID 18083096.
- ^ Niki I, Yokokura H, Sudo T, Kato M, Hidaka H (1996). "Ca2+ signaling and intracellular Ca2+ binding proteins". J. Biochem. 120 (4): 685–98. PMID 8947828.
{{cite journal}}: CS1 maint: multiple names: authors list (link) - ^ Eady RR (1988). "The vanadium-containing nitrogenase of Azotobacter". Biofactors. 1 (2): 111–6. PMID 3076437.
- ^ Chan MK, Mukund S, Kletzin A, Adams MW, Rees DC (1995). "Structure of a hyperthermophilic tungstopterin enzyme, aldehyde ferredoxin oxidoreductase". Science. 267 (5203): 1463–9. doi:10.1126/science.7878465. PMID 7878465.
{{cite journal}}: CS1 maint: multiple names: authors list (link) - ^ Lane TW, Morel FM (2000). "A biological function for cadmium in marine diatoms". Proc. Natl. Acad. Sci. U.S.A. 97 (9): 4627–31. doi:10.1073/pnas.090091397. PMID 10781068.
- ^ Lane TW, Saito MA, George GN, Pickering IJ, Prince RC, Morel FM (2005). "Biochemistry: a cadmium enzyme from a marine diatom". Nature. 435 (7038): 42. doi:10.1038/435042a. PMID 15875011.
{{cite journal}}: CS1 maint: multiple names: authors list (link) - ^ Palmer T (1981) Understanding enzymes. Ellis Horwood Limited, Chichester, UK
- ^ Nelson DL and Cox M.M (2000) Lehninger, Principles of Biochemistry, 3rd edition, Worth Publishers, New York
- ^ Campbell MK and Farrell SO (2009) Biochemistry, 6th edition, Thomson Brooks/Cole, Belmont, California
- ^ Bolander FF (2006). "Vitamins: not just for enzymes". Curr Opin Investig Drugs. 7 (10): 912–5. PMID 17086936.
- ^ Rouvière PE, Wolfe RS (1988). "Novel biochemistry of methanogenesis". J. Biol. Chem. 263 (17): 7913–6. PMID 3131330.
- ^ Frank RA, Leeper FJ, Luisi BF (2007). "Structure, mechanism and catalytic duality of thiamine-dependent enzymes". Cell. Mol. Life Sci. 64 (7–8): 892–905. doi:10.1007/s00018-007-6423-5. PMID 17429582.
{{cite journal}}: CS1 maint: multiple names: authors list (link) - ^ a b Pollak N, Dölle C, Ziegler M (2007). "The power to reduce: pyridine nucleotides--small molecules with a multitude of functions". Biochem. J. 402 (2): 205–18. doi:10.1042/BJ20061638. PMID 17295611.
{{cite journal}}: CS1 maint: multiple names: authors list (link) - ^ Leonardi R, Zhang YM, Rock CO, Jackowski S (2005). "Coenzyme A: back in action". Prog. Lipid Res. 44 (2–3): 125–53. doi:10.1016/j.plipres.2005.04.001. PMID 15893380.
{{cite journal}}: CS1 maint: multiple names: authors list (link) - ^ Donnelly JG (2001). "Folic acid". Crit Rev Clin Lab Sci. 38 (3): 183–223. doi:10.1080/20014091084209. PMID 11451208.
- ^ Søballe B, Poole RK (1999). "Microbial ubiquinones: multiple roles in respiration, gene regulation and oxidative stress management" (PDF). Microbiology (Reading, Engl.). 145 ( Pt 8): 1817–30. PMID 10463148.
- ^ Linster CL, Van Schaftingen E (2007). "Vitamin C. Biosynthesis, recycling and degradation in mammals". FEBS J. 274 (1): 1–22. doi:10.1111/j.1742-4658.2006.05607.x. PMID 17222174.
- ^ Mack M, Grill S (2006). "Riboflavin analogs and inhibitors of riboflavin biosynthesis". Appl. Microbiol. Biotechnol. 71 (3): 265–75. doi:10.1007/s00253-006-0421-7. PMID 16607521.
- ^ Bugg, Tim (1997). An introduction to enzyme and coenzyme chemistry. Oxford: Blackwell Science. p. 95. ISBN 0-86542-793-3.
- ^ Chiang P, Gordon R, Tal J, Zeng G, Doctor B, Pardhasaradhi K, McCann P (1996). "S-Adenosylmethionine and methylation". FASEB J. 10 (4): 471–80. PMID 8647346.
{{cite journal}}: CS1 maint: multiple names: authors list (link) - ^ Negishi M, Pedersen LG, Petrotchenko E; et al. (2001). "Structure and function of sulfotransferases". Arch. Biochem. Biophys. 390 (2): 149–57. doi:10.1006/abbi.2001.2368. PMID 11396917.
{{cite journal}}: Explicit use of et al. in:|author=(help)CS1 maint: multiple names: authors list (link) - ^ Crane FL (2001). "Biochemical functions of coenzyme Q10". Journal of the American College of Nutrition. 20 (6): 591–8. PMID 11771674.
- ^ Thony B, Auerbach G, Blau N (2000). "Tetrahydrobiopterin biosynthesis, regeneration and functions". Biochem J. 347 Pt 1: 1–16. doi:10.1042/0264-6021:3470001. PMID 10727395.
{{cite journal}}: CS1 maint: multiple names: authors list (link) - ^ Buchanan (2000). Biochemistry & molecular biology of plants (1st ed. ed.). American society of plant physiology. ISBN 0-943088-39-9.
{{cite book}}:|edition=has extra text (help); Unknown parameter|coauthors=ignored (|author=suggested) (help) - ^ Ginsburg V (1978). "Comparative biochemistry of nucleotide-linked sugars". Prog. Clin. Biol. Res. 23: 595–600. PMID 351635.
- ^ Grill D, Tausz T, De Kok LJ (2001). Significance of glutathione in plant adaptation to the environment. Springer. ISBN 1402001789.
{{cite book}}: CS1 maint: multiple names: authors list (link) - ^ Meister A, Anderson ME (1983). "Glutathione". Annu. Rev. Biochem. 52: 711–60. doi:10.1146/annurev.bi.52.070183.003431. PMID 6137189.
- ^ Taylor CD, Wolfe RS (1974). "Structure and methylation of coenzyme M(HSCH2CH2SO3)". J. Biol. Chem. 249 (15): 4879–85. PMID 4367810.
- ^ Balch WE, Wolfe RS (1979). "Specificity and biological distribution of coenzyme M (2-mercaptoethanesulfonic acid)". J. Bacteriol. 137 (1): 256–63. PMID 104960.
- ^ Noll KM, Rinehart KL, Tanner RS, Wolfe RS (1986). "Structure of component B (7-mercaptoheptanoylthreonine phosphate) of the methylcoenzyme M methylreductase system of Methanobacterium thermoautotrophicum". Proc. Natl. Acad. Sci. U.S.A. 83 (12): 4238–42. doi:10.1073/pnas.83.12.4238. PMID 3086878.
{{cite journal}}: CS1 maint: multiple names: authors list (link) - ^ Vorholt JA, Thauer RK (1997). "The active species of 'CO2' utilized by formylmethanofuran dehydrogenase from methanogenic Archaea". Eur. J. Biochem. 248 (3): 919–24. doi:10.1111/j.1432-1033.1997.00919.x. PMID 9342247.
- ^ DiMarco AA, Bobik TA, Wolfe RS (1990). "Unusual coenzymes of methanogenesis". Annu. Rev. Biochem. 59: 355–94. doi:10.1146/annurev.bi.59.070190.002035. PMID 2115763.
{{cite journal}}: CS1 maint: multiple names: authors list (link) - ^ Mitchell P (1979). "The Ninth Sir Hans Krebs Lecture. Compartmentation and communication in living systems. Ligand conduction: a general catalytic principle in chemical, osmotic and chemiosmotic reaction systems". Eur J Biochem. 95 (1): 1–20. doi:10.1111/j.1432-1033.1979.tb12934.x. PMID 378655.
- ^ Wimmer M, Rose I. "Mechanisms of enzyme-catalyzed group transfer reactions". Annu Rev Biochem. 47: 1031–78. doi:10.1146/annurev.bi.47.070178.005123. PMID 354490.
- ^ Di Carlo, S. E. and Coliins, H. L. (2001) "Estimating ATP resynthesis during a marathon run: a method to introduce metabolism" Advan. Physiol. Edu. 25: 70-71.
- ^ Chen X, Li N, Ellington AD (2007). "Ribozyme catalysis of metabolism in the RNA world". Chem. Biodivers. 4 (4): 633–55. doi:10.1002/cbdv.200790055. PMID 17443876.
{{cite journal}}: CS1 maint: multiple names: authors list (link) - ^ Koch A (1998). "How did bacteria come to be?". Adv Microb Physiol. 40: 353–99. PMID 9889982.
- ^ Ouzounis C, Kyrpides N (1996). "The emergence of major cellular processes in evolution". FEBS Lett. 390 (2): 119–23. doi:10.1016/0014-5793(96)00631-X. PMID 8706840.
- ^ White HB (1976). "Coenzymes as fossils of an earlier metabolic state". J. Mol. Evol. 7 (2): 101–4. doi:10.1007/BF01732468. PMID 1263263.
- ^ Saran D, Frank J, Burke DH (2003). "The tyranny of adenosine recognition among RNA aptamers to coenzyme A". BMC Evol. Biol. 3: 26. doi:10.1186/1471-2148-3-26. PMID 14687414.
{{cite journal}}: CS1 maint: multiple names: authors list (link) CS1 maint: unflagged free DOI (link) - ^ Jadhav VR, Yarus M (2002). "Coenzymes as coribozymes". Biochimie. 84 (9): 877–88. doi:10.1016/S0300-9084(02)01404-9. PMID 12458080.
- ^ Harden A, Young WJ. "The Alcoholic Ferment of Yeast-Juice" Proceedings of the Royal Society of London. Series B, Containing Papers of a Biological Character Vol. 78, No. 526 (Oct., 1906), pp. 369-375
- ^ "Fermentation of sugars and fermentative enzymes: Nobel Lecture, May 23, 1930" (PDF). Nobel Foundation. Retrieved 2007-09-30.
- ^ Lohmann, K. (1929) Über die Pyrophosphatfraktion im Muskel. Naturwissenschaften 17, 624–625.
- ^ Lipmann F (1945). "Acetylation of sulfanilamide by liver homogenates and extracts". J. Biol. Chem. 160 (1): 173–190.
- ^ Warburg O, Christian W. (1936). "Pyridin, the hydrogen-transferring component of the fermentation enzymes (pyridine nucleotide)". Biochemische Zeitschrift. 287: 291.
- ^ Kalckar HM (1974). "Origins of the concept oxidative phosphorylation". Mol. Cell. Biochem. 5 (1–2): 55–63. doi:10.1007/BF01874172. PMID 4279328.
- ^ Lipmann F, (1941). "Metabolic generation and utilization of phosphate bond energy". Adv Enzymol. 1: 99–162.
{{cite journal}}: CS1 maint: extra punctuation (link) - ^ Friedkin M, Lehninger AL. (1949). "Esterification of inorganic phosphate coupled to electron transport between dihydrodiphosphopyridine nucleotide and oxygen". J. Biol. Chem. 178 (2): 611–23. PMID 18116985.
Further reading
- Bugg, Tim (1997). An introduction to enzyme and coenzyme chemistry. Oxford: Blackwell Science. ISBN 0-86542-793-3.
External links
- Template:EMedicineDictionary
- Enzyme+cofactors at the U.S. National Library of Medicine Medical Subject Headings (MeSH)
