Ku (protein)
| Ku complex family | |||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
 | |||||||||||||||||||||||||||||||||||||||||||||||||||
| Identifiers | |||||||||||||||||||||||||||||||||||||||||||||||||||
| Aliases | Ku70:Ku80 heterodimerKu70:Ku80Ku Autoantigen | ||||||||||||||||||||||||||||||||||||||||||||||||||
| External IDs | GeneCards: [1] | ||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| Wikidata | |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| X-ray repair cross-complementing 5 | |||||||
|---|---|---|---|---|---|---|---|
 | |||||||
| Identifiers | |||||||
| Symbol | XRCC5 | ||||||
| Alt. symbols | Ku80 | ||||||
| NCBI gene | 7520 | ||||||
| HGNC | 12833 | ||||||
| OMIM | 194364 | ||||||
| PDB | 1JEY | ||||||
| RefSeq | NM_021141 | ||||||
| UniProt | P13010 | ||||||
| Other data | |||||||
| Locus | Chr. 2 q35 | ||||||
| |||||||
| X-ray repair cross-complementing 6 | |||||||
|---|---|---|---|---|---|---|---|
| Identifiers | |||||||
| Symbol | XRCC6 | ||||||
| Alt. symbols | Ku70, G22P1 | ||||||
| NCBI gene | 2547 | ||||||
| HGNC | 4055 | ||||||
| OMIM | 152690 | ||||||
| PDB | 1JEY | ||||||
| RefSeq | NM_001469 | ||||||
| UniProt | P12956 | ||||||
| Other data | |||||||
| Locus | Chr. 22 q11-q13 | ||||||
| |||||||
| Ku70/Ku80 N-terminal alpha/beta domain | |||||||||
|---|---|---|---|---|---|---|---|---|---|
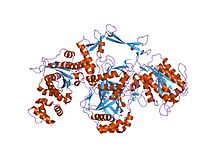 crystal structure of the ku heterodimer | |||||||||
| Identifiers | |||||||||
| Symbol | Ku_N | ||||||||
| Pfam | PF03731 | ||||||||
| Pfam clan | CL0128 | ||||||||
| InterPro | IPR005161 | ||||||||
| SCOP2 | 1jey / SCOPe / SUPFAM | ||||||||
| |||||||||
| Ku70/Ku80 beta-barrel domain | |||||||||
|---|---|---|---|---|---|---|---|---|---|
 crystal structure of the ku heterodimer bound to dna | |||||||||
| Identifiers | |||||||||
| Symbol | Ku | ||||||||
| Pfam | PF02735 | ||||||||
| InterPro | IPR006164 | ||||||||
| PROSITE | PDOC00252 | ||||||||
| SCOP2 | 1jey / SCOPe / SUPFAM | ||||||||
| |||||||||
| Ku70/Ku80 C-terminal arm | |||||||||
|---|---|---|---|---|---|---|---|---|---|
 crystal structure of the ku heterodimer bound to dna | |||||||||
| Identifiers | |||||||||
| Symbol | Ku_C | ||||||||
| Pfam | PF03730 | ||||||||
| InterPro | IPR005160 | ||||||||
| SCOP2 | 1jey / SCOPe / SUPFAM | ||||||||
| |||||||||
| Ku C terminal domain like | |||||||||
|---|---|---|---|---|---|---|---|---|---|
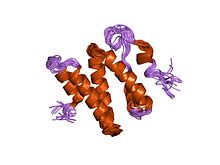 the 3d solution structure of the c-terminal region of ku86 | |||||||||
| Identifiers | |||||||||
| Symbol | Ku_PK_bind | ||||||||
| Pfam | PF08785 | ||||||||
| InterPro | IPR014893 | ||||||||
| SCOP2 | 1q2z / SCOPe / SUPFAM | ||||||||
| |||||||||
Ku is a dimeric protein complex that binds to DNA double-strand break ends and is required for the non-homologous end joining (NHEJ) pathway of DNA repair. Ku is evolutionarily conserved from bacteria to humans. The ancestral bacterial Ku is a homodimer (two copies of the same protein bound to each other).[2] Eukaryotic Ku is a heterodimer of two polypeptides, Ku70 (XRCC6) and Ku80 (XRCC5), so named because the molecular weight of the human Ku proteins is around 70 kDa and 80 kDa. The two Ku subunits form a basket-shaped structure that threads onto the DNA end.[1] Once bound, Ku can slide down the DNA strand, allowing more Ku molecules to thread onto the end. In higher eukaryotes, Ku forms a complex with the DNA-dependent protein kinase catalytic subunit (DNA-PKcs) to form the full DNA-dependent protein kinase, DNA-PK.[3] Ku is thought to function as a molecular scaffold to which other proteins involved in NHEJ can bind, orienting the double-strand break for ligation.
The Ku70 and Ku80 proteins consist of three structural domains. The N-terminal domain is an alpha/beta domain. This domain only makes a small contribution to the dimer interface. The domain comprises a six-stranded beta sheet of the Rossmann fold.[4] The central domain of Ku70 and Ku80 is a DNA-binding beta-barrel domain. Ku makes only a few contacts with the sugar-phosphate backbone, and none with the DNA bases, but it fits sterically to major and minor groove contours forming a ring that encircles duplex DNA, cradling two full turns of the DNA molecule. By forming a bridge between the broken DNA ends, Ku acts to structurally support and align the DNA ends, to protect them from degradation, and to prevent promiscuous binding to unbroken DNA. Ku effectively aligns the DNA, while still allowing access of polymerases, nucleases and ligases to the broken DNA ends to promote end joining.[5] The C-terminal arm is an alpha helical region which embraces the central beta-barrel domain of the opposite subunit.[1] In some cases a fourth domain is present at the C-terminus, which binds to DNA-dependent protein kinase catalytic subunit.[6]
Both subunits of Ku have been experimentally knocked out in mice. These mice exhibit chromosomal instability, indicating that NHEJ is important for genome maintenance.[7][8]
In many organisms, Ku has additional functions at telomeres in addition to its role in DNA repair.[9]
Abundance of Ku80 seems to be related to species longevity.[10]
Aging[edit]
Mutant mice defective in Ku70, or Ku80, or double mutant mice deficient in both Ku70 and Ku80 exhibit early aging.[11] The mean lifespans of the three mutant mouse strains were similar to each other, at about 37 weeks, compared to 108 weeks for the wild-type control. Six specific signs of aging were examined, and the three mutant mice were found to display the same aging signs as the control mice, but at a much earlier age. Cancer incidence was not increased in the mutant mice. These results suggest that Ku function is important for longevity assurance and that the NHEJ pathway of DNA repair (mediated by Ku) has a key role in repairing DNA double-strand breaks that would otherwise cause early aging.[12] (Also see DNA damage theory of aging.)
Plants[edit]
Ku70 and Ku80 have also been experimentally characterized in plants, where they appear to play a similar role to that in other eukaryotes. In rice, suppression of either protein has been shown to promote homologous recombination (HR)[13] This effect was exploited to improve gene targeting (GT) efficiency in Arabidopsis thaliana. In the study, the frequency of HR-based GT using a zinc-finger nuclease (ZFN) was increased up to sixteen times in ku70 mutants[14] This result has promising implications for genome editing across eukaryotes as DSB repair mechanisms are highly conserved. A substantial difference is that in plants, Ku is also involved in maintaining an alternate telomere morphology characterized by blunt-ends or short (≤ 3-nt) 3’ overhangs.[15] This function is independent of the role of Ku in DSB repair, as removing the ability of the Ku complex to translocate along DNA has been shown to preserve blunt-ended telomeres while impeding DNA repair.[16]
Bacteria and archaea[edit]
Bacteria usually have only one Ku gene (if they have one at all). Unusually, Mesorhizobium loti has two, mlr9624 and mlr9623.[17]
Archaea usually also only have one Ku gene (for the ~4% of species that have one at all). The evolutionary history is blurred by extensive horizontal gene transfer with bacteria.[18]
Bacterial and archaeal Ku proteins are unlike their eukaryotic counterparts in that they only have the central beta-barrel domain.
Name[edit]
The name 'Ku' is derived from the surname of the Japanese patient in which it was discovered.[19]
References[edit]
- ^ a b c PDB: 1JEY; Walker JR, Corpina RA, Goldberg J (August 2001). "Structure of the Ku heterodimer bound to DNA and its implications for double-strand break repair". Nature. 412 (6847): 607–14. Bibcode:2001Natur.412..607W. doi:10.1038/35088000. PMID 11493912. S2CID 4371575.
- ^ Doherty AJ, Jackson SP, Weller GR (July 2001). "Identification of bacterial homologues of the Ku DNA repair proteins". FEBS Lett. 500 (3): 186–8. doi:10.1016/S0014-5793(01)02589-3. PMID 11445083. S2CID 43588474.
- ^ Carter T, Vancurová I, Sun I, Lou W, DeLeon S (December 1990). "A DNA-activated protein kinase from HeLa cell nuclei". Mol. Cell. Biol. 10 (12): 6460–71. doi:10.1128/MCB.10.12.6460. PMC 362923. PMID 2247066.
- ^ Sugihara T, Wadhwa R, Kaul SC, Mitsui Y (April 1999). "A novel testis-specific metallothionein-like protein, tesmin, is an early marker of male germ cell differentiation". Genomics. 57 (1): 130–6. doi:10.1006/geno.1999.5756. PMID 10191092.
- ^ Aravind L, Koonin EV (August 2001). "Prokaryotic homologs of the eukaryotic DNA-end-binding protein Ku, novel domains in the Ku protein and prediction of a prokaryotic double-strand break repair system". Genome Res. 11 (8): 1365–74. doi:10.1101/gr.181001. PMC 311082. PMID 11483577.
- ^ Harris R, Esposito D, Sankar A, Maman JD, Hinks JA, Pearl LH, Driscoll PC (January 2004). "The 3D solution structure of the C-terminal region of Ku86 (Ku86CTR)". J. Mol. Biol. 335 (2): 573–82. doi:10.1016/j.jmb.2003.10.047. PMID 14672664.
- ^ Difilippantonio MJ, Zhu J, Chen HT, Meffre E, Nussenzweig MC, Max EE, Ried T, Nussenzweig A (March 2000). "DNA repair protein Ku80 suppresses chromosomal aberrations and malignant transformation". Nature. 404 (6777): 510–4. Bibcode:2000Natur.404..510D. doi:10.1038/35006670. PMC 4721590. PMID 10761921.
- ^ Ferguson DO, Sekiguchi JM, Chang S, Frank KM, Gao Y, DePinho RA, Alt FW (June 2000). "The nonhomologous end-joining pathway of DNA repair is required for genomic stability and the suppression of translocations". Proc. Natl. Acad. Sci. U.S.A. 97 (12): 6630–3. Bibcode:2000PNAS...97.6630F. doi:10.1073/pnas.110152897. PMC 18682. PMID 10823907.
- ^ Boulton SJ, Jackson SP (March 1998). "Components of the Ku-dependent non-homologous end-joining pathway are involved in telomeric length maintenance and telomeric silencing". EMBO J. 17 (6): 1819–28. doi:10.1093/emboj/17.6.1819. PMC 1170529. PMID 9501103.
- ^ Lorenzini A, Johnson FB, Oliver A, Tresini M, Smith JS, Hdeib M, Sell C, Cristofalo VJ, Stamato TD (Nov–Dec 2009). "Significant Correlation of Species Longevity with DNA Double Strand Break-Recognition but not with Telomere Length". Mech Ageing Dev. 130 (11–12): 784–92. doi:10.1016/j.mad.2009.10.004. PMC 2799038. PMID 19896964.
- ^ Li H, Vogel H, Holcomb VB, Gu Y, Hasty P (2007). "Deletion of Ku70, Ku80, or both causes early aging without substantially increased cancer". Mol. Cell. Biol. 27 (23): 8205–14. doi:10.1128/MCB.00785-07. PMC 2169178. PMID 17875923.
- ^ Bernstein H, Payne CM, Bernstein C, Garewal H, Dvorak K (2008). "Cancer and aging as consequences of un-repaired DNA damage". In: New Research on DNA Damages (Editors: Honoka Kimura and Aoi Suzuki) Nova Science Publishers, New York, Chapter 1, pp. 1-47. open access, but read only https://www.novapublishers.com/catalog/product_info.php?products_id=43247 Archived 2014-10-25 at the Wayback Machine ISBN 978-1604565812
- ^ Nishizawa-Yokoi A, Nonaka S, Saika H, Kwon YI, Osakabe K, Toki S (December 2012). "Suppression of Ku70/80 or Lig4 leads to decreased stable transformation and enhanced homologous recombination in rice". The New Phytologist. 196 (4): 1048–59. doi:10.1111/j.1469-8137.2012.04350.x. PMC 3532656. PMID 23050791.
- ^ Qi Y, Zhang Y, Zhang F, Baller JA, Cleland SC, Ryu Y, Starker CG, Voytas DF (March 2013). "Increasing frequencies of site-specific mutagenesis and gene targeting in Arabidopsis by manipulating DNA repair pathways". Genome Research. 23 (3): 547–54. doi:10.1101/gr.145557.112. PMC 3589543. PMID 23282329.
- ^ Kazda A, Zellinger B, Rössler M, Derboven E, Kusenda B, Riha K (August 2012). "Chromosome end protection by blunt-ended telomeres". Genes & Development. 26 (15): 1703–13. doi:10.1101/gad.194944.112. PMC 3418588. PMID 22810623.
- ^ Valuchova S, Fulnecek J, Prokop Z, Stolt-Bergner P, Janouskova E, Hofr C, Riha K (June 2017). "Protection of Arabidopsis Blunt-Ended Telomeres Is Mediated by a Physical Association with the Ku Heterodimer". The Plant Cell. 29 (6): 1533–1545. doi:10.1105/tpc.17.00064. PMC 5502450. PMID 28584163.
- ^ Pitcher RS, Brissett NC, Doherty AJ (2007). "Nonhomologous end-joining in bacteria: a microbial perspective". Annual Review of Microbiology. 61 (1). Annual Reviews: 259–82. doi:10.1146/annurev.micro.61.080706.093354. PMID 17506672.
- ^ Sharda M, Badrinarayanan A, Seshasayee AS (December 2020). "Evolutionary and Comparative Analysis of Bacterial Nonhomologous End Joining Repair". Genome Biology and Evolution. 12 (12): 2450–2466. doi:10.1093/gbe/evaa223. PMC 7719229. PMID 33078828.
- ^ Dynan WS, Yoo S (April 1998). "Interaction of Ku protein and DNA-dependent protein kinase catalytic subunit with nucleic acids". Nucleic Acids Research. 26 (7): 1551–9. doi:10.1093/nar/26.7.1551. PMC 147477. PMID 9512523.
