Werner syndrome helicase
Werner syndrome ATP-dependent helicase, also known as DNA helicase, RecQ-like type 3, is an enzyme that in humans is encoded by the WRN gene. WRN is a member of the RecQ Helicase family.[5] Helicase enzymes generally unwind and separate double-stranded DNA. These activities are necessary before DNA can be copied in preparation for cell division (DNA replication). Helicase enzymes are also critical for making a blueprint of a gene for protein production, a process called transcription. Further evidence suggests that Werner protein plays a critical role in repairing DNA. Overall, this protein helps maintain the structure and integrity of a person's DNA.
The WRN gene is located on the short (p) arm of chromosome 8 between positions 12 and 11.2, from base pair 31,010,319 to base pair 31,150,818.
Structure and function[edit]
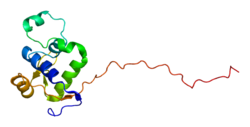
WRN is a member of the RecQ Helicase family. It is the only RecQ Helicase that contains 3' to 5' exonuclease activity. These exonuclease activities include degradation of recessed 3' ends and initiation of DNA degradation from a gap in dsDNA. WRN is important in repair of double strand breaks by homologous recombination[6][7] or non-homologous end joining,[8] repair of single nucleotide damages by base excision repair,[9][10][5] and is effective in replication arrest recovery.[11] WRN may also be important in telomere maintenance and replication, especially the replication of the G-rich sequences.[12]
WRN is an oligomer that can act as a monomer when unwinding DNA, but as a dimer in solution or a tetramer when complexed with DNA, and has also been observed in hexameric forms. The diffusion of WRN has been measured to 1.62 in nucleoplasm and 0.12 at nucleoli.[13] Orthologs of WRN have been found in a number of other organisms, including Drosophila, Xenopus, and C. elegans. WRN is important to genome stability, and cells with mutations to WRN are more susceptible to DNA damage and DNA breaks.[14]
The amino terminus of WRN is involved in both helicase and nuclease activities, while the carboxyl-terminus interacts with p53, an important tumor suppressor.[15] WRN may function as an exonuclease in DNA repair, recombination, or replication, as well as resolution of DNA secondary structures. It is involved in branch migration at Holliday junctions, and it interacts with other DNA replication intermediates.[11] mRNA that codes for WRN has been identified in most human tissues.[15]
Post-translational modification[edit]
Phosphorylation of WRN at serine/threonine inhibits helicase and exonuclease activities which are important to post-replication DNA repair. De-phosphorylation at these sites enhances the catalytic activities of WRN. Phosphorylation may affect other post-translational modifications, including sumoylation and acetylation.[12]
Methylation of WRN causes the gene to turn off. This suppresses the production of the WRN protein and its functions in DNA repair.[16]
Clinical significance[edit]
Werner syndrome is caused by mutations in the WRN gene.[15] More than 20 mutations in the WRN gene are known to cause Werner syndrome. Many of these mutations result in an abnormally shortened Werner protein. Evidence suggests that the altered protein is not transported into the cell nucleus, where it normally interacts with DNA.[17] This shortened protein may also be broken down too quickly, leading to a loss of Werner protein in the cell. Without normal Werner protein in the nucleus, cells cannot perform the tasks of DNA replication, repair, and transcription.[18] Researchers are still determining how these mutations cause the appearance of premature aging seen in Werner syndrome.
Roles in DNA repair pathways[edit]
Homologous recombinational repair[edit]
WRN is active in homologous recombination. Cells defective in the WRN gene have a 23-fold reduction in spontaneous mitotic recombination, with especial deficiency in conversion-type events.[19] WRN defective cells, when exposed to x-rays, have more chromosome breaks and micronuclei than cells with wild-type WRN.[20] Cells defective in the WRN gene are not more sensitive than wild-type cells to gamma-irradiation, UV light, 4 – 6 cyclobutane pyrimidines, or mitomycin C, but are sensitive to type I and type II topoisomerase inhibitors.[21] These findings suggested that the WRN protein takes part in homologous recombinational repair and in the processing of stalled replication forks.[22]
Non-homologous end joining[edit]
WRN has an important role in non-homologous end joining (NHEJ) DNA repair. As shown by Shamanna et al.,[8] WRN is recruited to double-strand breaks (DSBs) and participates in NHEJ with its enzymatic and non-enzymatic functions. At DSBs, in association with Ku (protein), it promotes standard or canonical NHEJ (c-NHEJ), repairing double-strand breaks in DNA with its enzymatic functions and with a fair degree of accuracy. WRN inhibits an alternative form of NHEJ, called alt-NHEJ or microhomology-mediated end joining (MMEJ). MMEJ is an inaccurate mode of repair for double-strand breaks.
Base excision repair[edit]
WRN has a role in base excision repair (BER) of DNA. As shown by Das et al.,[9] WRN associates with NEIL1 in the early damage-sensing step of BER. WRN stimulates NEIL1 in excision of oxidative lesions. NEIL1 is a DNA glycosylase that initiates the first step in BER by cleaving bases damaged by reactive oxygen species (ROS) and introducing a DNA strand break via NEIL1's associated lyase activity.[23] NEIL1 recognizes (targets) and removes certain ROS-damaged bases and then incises the abasic site via β,δ elimination, leaving 3′ and 5′ phosphate ends. NEIL1 recognizes oxidized pyrimidines, formamidopyrimidines, thymine residues oxidized at the methyl group, and both stereoisomers of thymine glycol.[24]
WRN also participates in BER through its interaction with Polλ.[10] WRN binds to the catalytic domain of Polλ and specifically stimulates DNA gap filling by Polλ over 8-oxo-G followed by strand displacement synthesis. This allows WRN to promote long-patch DNA repair synthesis by Polλ during MUTYH-initiated repair of 8-oxo-G:A mispairs.
Replication arrest recovery[edit]
WRN is also involved in replication arrest recovery. If WRN is defective, replication arrest results in accumulation of DSBs and enhanced chromosome fragmentation.[25] As shown by Pichierri et al.,[25] WRN interacts with the RAD9-RAD1-HUS1 (9.1.1) complex, one of the central factors of the replication checkpoint. This interaction is mediated by the binding of the RAD1 subunit to the N-terminal region of WRN and is instrumental for WRN relocalization to nuclear foci and its phosphorylation in response to replication arrest. (In the absence of DNA damage or replication fork stalling, WRN protein remains localized to the nucleoli.[26]) The interaction of WRN with the 9.1.1 complex results in prevention of DSB formation at stalled replication forks.[25]
Role in apoptosis[edit]
The p53 protein and WRN helicase engage in direct protein-protein interaction.[27] Increased cellular WRN levels elicit increased cellular p53 levels and also potentiate p53-mediated apoptosis.[27] This finding suggests that WRN helicase participates in the activation of p53 in response to certain types of DNA damage.[27] p53-mediated apoptosis is attenuated in cells from patients with Werner syndrome.[28]
Both Repair of DNA damages and apoptosis are enzymatic processes necessary for maintaining integrity of the genome in humans. Cells with insufficient DNA repair tend to accumulate DNA damages, and when such cells are also defective in apoptosis they tend to survive even though excessive DNA damages are present.[29] Replication of DNA in such deficient cells tends to lead to mutations and such mutations may cause cancer. Thus Werner syndrome helicase appears to have two roles related to the prevention of cancer, where the first role is to promote repair of specific types of damage and the second role is to induce apoptosis if the level of such DNA damage is beyond the cell’s repair capability[29]
WRN deficiencies in cancer[edit]
Cells expressing limiting amounts of WRN have elevated mutation frequencies compared with wildtype cells.[30] Increased mutation may give rise to cancer. Patients with Werner Syndrome, with homozygous mutations in the WRN gene, have an increased incidence of cancers, including soft tissue sarcomas, osteosarcoma, thyroid cancer and melanoma.[31]
Mutations in WRN are rare in the general population. The rate of heterozygous loss of-function mutation in WRN is approximately one per million. In a Japanese population the rate is 6 per 1,000, which is higher, but still infrequent.[32]
Mutational defects in the WRN gene are relatively rare in cancer cells compared to the frequency of epigenetic alterations in WRN that reduce WRN expression and could contribute to carcinogenesis. The situation is similar to other DNA repair genes whose expression is reduced in cancers due to mainly epigenetic alterations rather than mutations (see Frequencies of epimutations in DNA repair genes).[citation needed]
The table shows results of analysis of 630 human primary tumors for WRN CpG island hypermethylation.[33] This hypermethylation caused reduced protein expression of WRN, a common event in tumorigenesis.[33]
| Cancer | Frequency of reduction in cancer[33] |
|---|---|
| Colorectal cancer | 37.9% |
| Non-small cell lung cancer | 37.5% |
| Gastric cancer | 25% |
| Prostate cancer | 20% |
| Breast cancer | 17.2% |
| Thyroid cancer | 12.5% |
| Non-Hodgkin lymphoma | 23.7% |
| Acute myeloblastic leukemia | 4.8% |
| Chondrosarcomas | 33.3% |
| Osteosarcomas | 11.1% |
Interactions[edit]
Werner syndrome ATP-dependent helicase has been shown to interact with:
References[edit]
- ^ a b c GRCh38: Ensembl release 89: ENSG00000165392 – Ensembl, May 2017
- ^ a b c GRCm38: Ensembl release 89: ENSMUSG00000031583 – Ensembl, May 2017
- ^ "Human PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ "Mouse PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ a b Monnat RJ (October 2010). "Human RECQ helicases: roles in DNA metabolism, mutagenesis and cancer biology". Semin. Cancer Biol. 20 (5): 329–39. doi:10.1016/j.semcancer.2010.10.002. PMC 3040982. PMID 20934517.
- ^ Saintigny Y, Makienko K, Swanson C, Emond MJ, Monnat RJ (2002). "Homologous recombination resolution defect in werner syndrome". Mol. Cell. Biol. 22 (20): 6971–8. doi:10.1128/mcb.22.20.6971-6978.2002. PMC 139822. PMID 12242278.
- ^ Sturzenegger A, Burdova K, Kanagaraj R, Levikova M, Pinto C, Cejka P, Janscak P (2014). "DNA2 cooperates with the WRN and BLM RecQ helicases to mediate long-range DNA end resection in human cells". J. Biol. Chem. 289 (39): 27314–26. doi:10.1074/jbc.M114.578823. PMC 4175362. PMID 25122754.
- ^ a b Shamanna RA, Lu H, de Freitas JK, Tian J, Croteau DL, Bohr VA (2016). "WRN regulates pathway choice between classical and alternative non-homologous end joining". Nat Commun. 7: 13785. Bibcode:2016NatCo...713785S. doi:10.1038/ncomms13785. PMC 5150655. PMID 27922005.
- ^ a b Das A, Boldogh I, Lee JW, Harrigan JA, Hegde ML, Piotrowski J, de Souza Pinto N, Ramos W, Greenberg MM, Hazra TK, Mitra S, Bohr VA (2007). "The human Werner syndrome protein stimulates repair of oxidative DNA base damage by the DNA glycosylase NEIL1". J. Biol. Chem. 282 (36): 26591–602. doi:10.1074/jbc.M703343200. PMID 17611195.
- ^ a b Kanagaraj R, Parasuraman P, Mihaljevic B, van Loon B, Burdova K, König C, Furrer A, Bohr VA, Hübscher U, Janscak P (2012). "Involvement of Werner syndrome protein in MUTYH-mediated repair of oxidative DNA damage". Nucleic Acids Res. 40 (17): 8449–59. doi:10.1093/nar/gks648. PMC 3458577. PMID 22753033.
- ^ a b Pichierri P, Ammazzalorso F, Bignami M, Franchitto A (2011). "The Werner syndrome protein: linking the replication checkpoint response to genome stability". Aging. 3 (3): 311–8. doi:10.18632/aging.100293. PMC 3091524. PMID 21389352.
- ^ a b Ding SL, Shen CY (2008). "Model of human aging: recent findings on Werner's and Hutchinson–Gilford progeria syndromes". Clin Interv Aging. 3 (3): 431–44. doi:10.2147/CIA.S1957. PMC 2682376. PMID 18982914.
- ^ Bendtsen KM, Jensen MB, May A, Rasmussen LJ, Trusina A, Bohr VA, Jensen MH (2014). "Dynamics of the DNA repair proteins WRN and BLM in the nucleoplasm and nucleoli". European Biophysics Journal. 43 (10–11): 509–16. doi:10.1007/s00249-014-0981-x. PMC 5576897. PMID 25119658.
- ^ Rossi ML, Ghosh AK, Bohr VA (2010). "Roles of Werner syndrome protein in protection of genome integrity". DNA Repair (Amst.). 9 (3): 331–44. doi:10.1016/j.dnarep.2009.12.011. PMC 2827637. PMID 20075015.
- ^ a b c Oshima J (2000). "The Werner syndrome protein: an update". BioEssays. 22 (10): 894–901. doi:10.1002/1521-1878(200010)22:10<894::AID-BIES4>3.0.CO;2-B. PMID 10984715. S2CID 36746466.
- ^ "WRN". US National Library of Medicine. Retrieved 18 March 2014.
- ^ Huang S, Lee L, Hanson NB, Lenaerts C, Hoehn H, Poot M, Rubin CD, Chen DF, Yang CC, Juch H, Dorn T, Spiegel R, Oral EA, Abid M, Battisti C, Lucci-Cordisco E, Neri G, Steed EH, Kidd A, Isley W, Showalter D, Vittone JL, Konstantinow A, Ring J, Meyer P, Wenger SL, von Herbay A, Wollina U, Schuelke M, Huizenga CR, Leistritz DF, Martin GM, Mian IS, Oshima J (2006). "The spectrum of WRN mutations in Werner syndrome patients". Hum. Mutat. 27 (6): 558–67. doi:10.1002/humu.20337. PMC 1868417. PMID 16673358.
- ^ Lebel M (2001). "Werner syndrome: genetic and molecular basis of a premature aging disorder". Cell. Mol. Life Sci. 58 (7): 857–67. doi:10.1007/s00018-001-8398-y. PMID 11497235. S2CID 24801894.
- ^ Prince PR, Emond MJ, Monnat RJ (2001). "Loss of Werner syndrome protein function promotes aberrant mitotic recombination". Genes Dev. 15 (8): 933–8. doi:10.1101/gad.877001. PMC 312674. PMID 11316787.
- ^ Weirich-Schwaiger H, Weirich HG, Gruber B, Schweiger M, Hirsch-Kauffmann M (1994). "Correlation between senescence and DNA repair in cells from young and old individuals and in premature aging syndromes". Mutat. Res. 316 (1): 37–48. doi:10.1016/0921-8734(94)90006-x. PMID 7507567.
- ^ Lebel M, Leder P (1998). "A deletion within the murine Werner syndrome helicase induces sensitivity to inhibitors of topoisomerase and loss of cellular proliferative capacity". Proc. Natl. Acad. Sci. U.S.A. 95 (22): 13097–102. Bibcode:1998PNAS...9513097L. doi:10.1073/pnas.95.22.13097. PMC 23722. PMID 9789047.
- ^ Sakamoto S, Nishikawa K, Heo SJ, Goto M, Furuichi Y, Shimamoto A (2001). "Werner helicase relocates into nuclear foci in response to DNA damaging agents and co-localizes with RPA and Rad51". Genes Cells. 6 (5): 421–30. doi:10.1046/j.1365-2443.2001.00433.x. PMID 11380620. S2CID 26078155.
- ^ Jacobs AC, Calkins MJ, Jadhav A, Dorjsuren D, Maloney D, Simeonov A, Jaruga P, Dizdaroglu M, McCullough AK, Lloyd RS (2013). "Inhibition of DNA glycosylases via small molecule purine analogs". PLOS ONE. 8 (12): e81667. Bibcode:2013PLoSO...881667J. doi:10.1371/journal.pone.0081667. PMC 3857224. PMID 24349107.
- ^ Nemec AA, Wallace SS, Sweasy JB (Oct 2010). "Variant base excision repair proteins: contributors to genomic instability". Seminars in Cancer Biology. 20 (5): 320–8. doi:10.1016/j.semcancer.2010.10.010. PMC 3254599. PMID 20955798.
- ^ a b c Pichierri P, Nicolai S, Cignolo L, Bignami M, Franchitto A (2012). "The RAD9-RAD1-HUS1 (9.1.1) complex interacts with WRN and is crucial to regulate its response to replication fork stalling". Oncogene. 31 (23): 2809–23. doi:10.1038/onc.2011.468. PMC 3272477. PMID 22002307.
- ^ Constantinou A, Tarsounas M, Karow JK, Brosh RM, Bohr VA, Hickson ID, West SC (2000). "Werner's syndrome protein (WRN) migrates Holliday junctions and co-localizes with RPA upon replication arrest". EMBO Rep. 1 (1): 80–4. doi:10.1093/embo-reports/kvd004. PMC 1083680. PMID 11256630.
- ^ a b c Blander G, Zalle N, Leal JF, Bar-Or RL, Yu CE, Oren M. The Werner syndrome protein contributes to induction of p53 by DNA damage. FASEB J. 2000 Nov;14(14):2138-40. doi: 10.1096/fj.00-0171fje. PMID: 11023999
- ^ Spillare EA, Wang XW, von Kobbe C, Bohr VA, Hickson ID, Harris CC. Redundancy of DNA helicases in p53-mediated apoptosis. Oncogene. 2006 Mar 30;25(14):2119-23. doi: 10.1038/sj.onc.1209242. PMID: 16288211; PMCID: PMC1420682
- ^ a b Bernstein C, Bernstein H, Payne CM, Garewal H. DNA repair/pro-apoptotic dual-role proteins in five major DNA repair pathways: fail-safe protection against carcinogenesis. Mutat Res. 2002 Jun;511(2):145-78. doi: 10.1016/s1383-5742(02)00009-1. PMID: 12052432
- ^ Kamath-Loeb AS, Shen JC, Schmitt MW, Loeb LA (2012). "The Werner syndrome exonuclease facilitates DNA degradation and high fidelity DNA polymerization by human DNA polymerase δ". J. Biol. Chem. 287 (15): 12480–90. doi:10.1074/jbc.M111.332577. PMC 3320997. PMID 22351772.
- ^ Goto M, Miller RW, Ishikawa Y, Sugano H (1996). "Excess of rare cancers in Werner syndrome (adult progeria)". Cancer Epidemiol. Biomarkers Prev. 5 (4): 239–46. PMID 8722214.
- ^ Chun SG, Shaeffer DS, Bryant-Greenwood PK (2011). "The Werner's Syndrome RecQ helicase/exonuclease at the nexus of cancer and aging". Hawaii Med J. 70 (3): 52–5. PMC 3071901. PMID 21365542.
- ^ a b c Agrelo R, Cheng WH, Setien F, Ropero S, Espada J, Fraga MF, Herranz M, Paz MF, Sanchez-Cespedes M, Artiga MJ, Guerrero D, Castells A, von Kobbe C, Bohr VA, Esteller M (2006). "Epigenetic inactivation of the premature aging Werner syndrome gene in human cancer". Proc. Natl. Acad. Sci. U.S.A. 103 (23): 8822–7. Bibcode:2006PNAS..103.8822A. doi:10.1073/pnas.0600645103. PMC 1466544. PMID 16723399.
- ^ von Kobbe C, Karmakar P, Dawut L, Opresko P, Zeng X, Brosh RM, Hickson ID, Bohr VA (June 2002). "Colocalization, physical, and functional interaction between Werner and Bloom syndrome proteins". J. Biol. Chem. 277 (24): 22035–44. doi:10.1074/jbc.M200914200. PMID 11919194.
- ^ Kim ST, Lim DS, Canman CE, Kastan MB (Dec 1999). "Substrate specificities and identification of putative substrates of ATM kinase family members". J. Biol. Chem. 274 (53): 37538–43. doi:10.1074/jbc.274.53.37538. PMID 10608806.
- ^ Karmakar P, Piotrowski J, Brosh RM, Sommers JA, Miller SP, Cheng WH, Snowden CM, Ramsden DA, Bohr VA (May 2002). "Werner protein is a target of DNA-dependent protein kinase in vivo and in vitro, and its catalytic activities are regulated by phosphorylation". J. Biol. Chem. 277 (21): 18291–302. doi:10.1074/jbc.M111523200. PMID 11889123.
- ^ Sharma S, Sommers JA, Wu L, Bohr VA, Hickson ID, Brosh RM (March 2004). "Stimulation of flap endonuclease-1 by the Bloom's syndrome protein". J. Biol. Chem. 279 (11): 9847–56. doi:10.1074/jbc.M309898200. PMID 14688284.
- ^ Brosh RM, von Kobbe C, Sommers JA, Karmakar P, Opresko PL, Piotrowski J, Dianova I, Dianov GL, Bohr VA (October 2001). "Werner syndrome protein interacts with human flap endonuclease 1 and stimulates its cleavage activity". EMBO J. 20 (20): 5791–801. doi:10.1093/emboj/20.20.5791. PMC 125684. PMID 11598021.
- ^ a b Karmakar P, Snowden CM, Ramsden DA, Bohr VA (August 2002). "Ku heterodimer binds to both ends of the Werner protein and functional interaction occurs at the Werner N-terminus". Nucleic Acids Res. 30 (16): 3583–91. doi:10.1093/nar/gkf482. PMC 134248. PMID 12177300.
- ^ a b Li B, Comai L (September 2000). "Functional interaction between Ku and the werner syndrome protein in DNA end processing". J. Biol. Chem. 275 (37): 28349–52. doi:10.1074/jbc.C000289200. PMID 10880505.
- ^ Yang Q, Zhang R, Wang XW, Spillare EA, Linke SP, Subramanian D, Griffith JD, Li JL, Hickson ID, Shen JC, Loeb LA, Mazur SJ, Appella E, Brosh RM, Karmakar P, Bohr VA, Harris CC (August 2002). "The processing of Holliday junctions by BLM and WRN helicases is regulated by p53". J. Biol. Chem. 277 (35): 31980–7. doi:10.1074/jbc.M204111200. hdl:10026.1/10341. PMID 12080066.
- ^ Brosh RM, Karmakar P, Sommers JA, Yang Q, Wang XW, Spillare EA, Harris CC, Bohr VA (September 2001). "p53 Modulates the exonuclease activity of Werner syndrome protein". J. Biol. Chem. 276 (37): 35093–102. doi:10.1074/jbc.M103332200. PMID 11427532.
- ^ Rodríguez-López AM, Jackson DA, Nehlin JO, Iborra F, Warren AV, Cox LS (February 2003). "Characterisation of the interaction between WRN, the helicase/exonuclease defective in progeroid Werner's syndrome, and an essential replication factor, PCNA". Mech. Ageing Dev. 124 (2): 167–74. doi:10.1016/S0047-6374(02)00131-8. PMID 12633936. S2CID 37287691.
- ^ Huang S, Beresten S, Li B, Oshima J, Ellis NA, Campisi J (June 2000). "Characterization of the human and mouse WRN 3'-->5' exonuclease". Nucleic Acids Res. 28 (12): 2396–405. doi:10.1093/nar/28.12.2396. PMC 102739. PMID 10871373.
- ^ Opresko PL, von Kobbe C, Laine JP, Harrigan J, Hickson ID, Bohr VA (October 2002). "Telomere-binding protein TRF2 binds to and stimulates the Werner and Bloom syndrome helicases". J. Biol. Chem. 277 (43): 41110–9. doi:10.1074/jbc.M205396200. PMID 12181313.
- ^ Branzei D, Hayashi T, Suzuki H, Masuko T, Onoda F, Heo SJ, Ikeda H, Shimamoto A, Furuichi Y, Seki M, Enomoto T (June 2001). "A novel protein interacts with the Werner's syndrome gene product physically and functionally". J. Biol. Chem. 276 (23): 20364–9. doi:10.1074/jbc.C100035200. PMID 11301316.
Further reading[edit]
- Comai L, Li B (2004). "The Werner syndrome protein at the crossroads of DNA repair and apoptosis". Mech Ageing Dev. 125 (8): 521–8. doi:10.1016/j.mad.2004.06.004. PMID 15336909. S2CID 30529954.
- Lee JW, Harrigan J, Opresko PL, Bohr VA (2005). "Pathways and functions of the Werner syndrome protein". Mech Ageing Dev. 126 (1): 79–86. doi:10.1016/j.mad.2004.09.011. PMID 15610765. S2CID 39834357.
- Monnat RJ Jr; Saintigny Y (2004). "Werner syndrome protein--unwinding function to explain disease" (PDF). Sci Aging Knowledge Environ. 2004 (13): re3. doi:10.1126/sageke.2004.13.re3. PMID 15056797. S2CID 15789751.
- Ozgenc A, Loeb LA (2005). "Current advances in unraveling the function of the Werner syndrome protein". Mutat Res. 577 (1–2): 237–51. doi:10.1016/j.mrfmmm.2005.03.020. PMID 15946710.
- Swanson C, Saintigny Y, Emond MJ, Monnat RJ Jr (2004). "The Werner syndrome protein has separable recombination and survival functions" (PDF). DNA Repair (Amst). 3 (5): 475–82. doi:10.1016/j.dnarep.2004.01.002. PMID 15084309. S2CID 21780379.
- Moser MJ, Oshima J, Monnat RJ (1999). "WRN mutations in Werner syndrome". Hum. Mutat. 13 (4): 271–9. doi:10.1002/(SICI)1098-1004(1999)13:4<271::AID-HUMU2>3.0.CO;2-Q. PMID 10220139. S2CID 35814236.
- Kastan MB, Lim DS (2001). "The many substrates and functions of ATM". Nat. Rev. Mol. Cell Biol. 1 (3): 179–86. doi:10.1038/35043058. PMID 11252893. S2CID 10691352.
External links[edit]
- Oshima J, Martin GM, Hisama FM (February 2012). Werner Syndrome. University of Washington, Seattle. PMID 20301687. NBK1514. In Adam MP, Everman DB, Mirzaa GM, Pagon RA, Wallace SE, Bean LJH, Gripp KW, Amemiya A (1993). Pagon RA, Bird TD, Dolan CR, et al. (eds.). GeneReviews [Internet]. Seattle WA: University of Washington, Seattle. PMID 20301295.
- GeneCard
- Werner Syndrome Mutational Database Archived 2012-07-21 at the Wayback Machine