Polymerase chain reaction

The polymerase chain reaction (PCR) is a biochemistry and molecular biology technique[1] for exponentially amplifying DNA, via enzymatic replication, without using a living organism (such as E. coli or yeast). As PCR is an in vitro technique, it can be performed without restrictions on the form of DNA, and it can be extensively modified to perform a wide array of genetic manipulations.
Invented in 1983 by Kary Mullis, PCR is now a common technique used in medical and biological research labs for a variety of tasks, such as the detection of hereditary diseases, the identification of genetic fingerprints, the diagnosis of infectious diseases, the cloning of genes, paternity testing, and DNA computing.
PCR in practice
PCR is used to amplify specific regions of a DNA strand. This can be a single gene, just a part of a gene, or a non-coding sequence. Most PCR methods typically amplify DNA fragments of up to 10 kilo base pairs (kb), although some techniques allow for amplification of fragments up to 40 kb in size.[2]
PCR, as currently practiced, requires several basic components [3]. These components are:
- DNA template that contains the region of the DNA fragment to be amplified
- One or more primers, which are complementary to the DNA regions at the 5' and 3' ends of the DNA region that is to be amplified.
- a DNA polymerase (e.g. Taq polymerase or another DNA polymerase with a temperature optimum at around 70°C), used to synthesize a DNA copy of the region to be amplified
- Deoxynucleotide triphosphates, (dNTPs) from which the DNA polymerase builds the new DNA
- Buffer solution, which provides a suitable chemical environment for optimum activity and stability of the DNA polymerase
- Divalent cation, magnesium or manganese ions; generally Mg2+ is used, but Mn2+ can be utilized for PCR-mediated DNA mutagenesis, as higher Mn2+ concentration increases the error rate during DNA synthesis [4]
- Monovalent cation potassium ions
The PCR is carried out in small reaction tubes (0.2-0.5 ml volumes), containing a reaction volume typically of 15-100 μl, that are inserted into a thermal cycler. This is a machine that heats and cools the reaction tubes within it to the precise temperature required for each step of the reaction. Most thermal cyclers have heated lids to prevent condensation on the inside of the reaction tube caps. Alternatively, a layer of oil may be placed on the reaction mixture to prevent evaporation.
Procedure

The PCR usually consists of a series of 20 to 35 cycles. Most commonly, PCR is carried out in three steps (Fig. 2), often preceded by one temperature hold at the start and followed by one hold at the end.
- Prior to the first cycle, during an initialization step, the PCR reaction is often heated to a temperature of 94-96°C (or 98°C if extremely thermostable polymerases are used), and this temperature is then held for 1-9 minutes. This first hold is employed to ensure that most of the DNA template and primers are denatured, i.e., that the DNA is melted by disrupting the hydrogen bonds between complementary bases of the DNA strands. Also, some PCR polymerases require this step for activation (see hot-start PCR). Following this hold, cycling begins, with one step at 94-98°C for 20-30 seconds (denaturation step).
- The denaturation is followed by the annealing step. In this step the reaction temperature is lowered so that the primers can attach to the single-stranded DNA template. Brownian motion causes the primers to move around, and DNA-DNA hydrogen bonds are constantly formed and broken between primer and template. Stable bonds are only formed when the primer sequence exactly matches the template sequence, and to this short section of double-stranded DNA the polymerase attaches and begins DNA synthesis. The temperature at this step depends on the Tm of the primers (see above), and is usually between 50-64°C for 20-40 seconds.
- The annealing step is followed by an extension/elongation step during which the DNA polymerase synthesizes new DNA strands complementary to the DNA template strands. The temperature at this step depends on the DNA polymerase used. Taq polymerase has a temperature optimum of 70-74°C; thus, in most cases a temperature of 72°C is used. The hydrogen bonds between the extended primer and the DNA template are now strong enough to withstand forces breaking these attractions at the higher temperature. Primers that have annealed to DNA regions with mismatching bases dissociate from the template and are not extended. The DNA polymerase condenses the 5'-phosphate group of the dNTPs with the 3'-hydroxyl group at the end of the nascent (extending) DNA strand, i.e., the polymerase adds dNTP's that are complementary to the template in 5' to 3' direction, thus reading the template in 3' to 5' direction. The extension time depends both on the DNA polymerase used and on the length of the DNA fragment to be amplified. As a rule-of-thumb, at its optimum temperature, the DNA polymerase will polymerize a thousand bases in one minute. A final elongation step of 5-15 minutes (depending on the length of the DNA template) after the last cycle may be used to ensure that any remaining single-stranded DNA is fully extended. A final hold of 4-15°C for an indefinite time may be employed for short-term storage of the reaction, e.g., if reactions are run overnight.
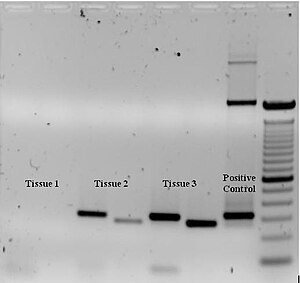
To check whether the PCR generated the anticipated DNA fragment (also sometimes referred to as amplimer), agarose gel electrophoresis is commonly employed for size separation of the PCR products. The size(s) of PCR products is thereby determined by comparison with a DNA ladder, which contains DNA fragments of known size, ran on the gel alongside the PCR products (see Fig. 3).
PCR optimization
In practice, PCR can fail for various reasons, in part due to its sensitivity to contamination causing amplification of spurious DNA products. Because of this, a number of techniques and procedures have been developed for optimizing PCR conditions. Contamination with extraneous DNA is addressed with lab protocols and procedures that separate pre-PCR reactions from potential DNA contaminants. This usually involves spatial separation of PCR-setup areas from areas for analysis or purification of PCR products, and thoroughly cleaning the work surface between reaction setups. Primer-design techniques are important in improving PCR product yield and in avoiding the formation of spurious products, and the usage of alternate buffer components or polymerase enzymes can help with amplification of long or otherwise problematic regions of DNA.
Practical modifications to the PCR technique
- Nested PCR - Nested PCR increases the specificity of DNA amplification, by reducing background due to non-specific amplification of DNA. Two sets of primers are being used in two successive PCR reactions. In the first reaction, one pair of primers is used to generate DNA products, which besides the intended target, may still consist of non-specifically amplified DNA fragments. The product(s) (sometimes after gel purification after electrophoresis of the PCR product) are then used in a second PCR reaction with a set of primers whose binding sites are completely or partially different from the primer pair used in the first reaction, but are completely within the DNA target fragment. Nested PCR is often more successful in specifically amplifying long DNA fragments than conventional PCR, but it requires more detailed knowledge of the target sequences.
- Intersequence specific (ISSR) PCR
- Ligation-mediated PCR
- Inverse PCR - Inverse PCR is a method used to allow PCR when only one internal sequence is known. This is especially useful in identifying flanking sequences to various genomic inserts. This involves a series of DNA digestions and self ligation, resulting in known sequences at either end of the unknown sequence.
- RT-PCR - RT-PCR (Reverse Transcription PCR) is a method used to amplify, isolate or identify a known sequence from a cellular or tissue RNA. The PCR reaction is preceded by a reaction using reverse transcriptase to convert RNA to cDNA. RT-PCR is widely used in expression profiling, to determine the expression of a gene or to identify the sequence of an RNA transcript, including transcription start and termination sites and, if the genomic DNA sequence of a gene is known, to map the location of exons and introns in the gene. The 5' end of a gene (corresponding to the transcription start site) is typically identified by a RT-PCR method, named RACE-PCR, short for Rapid Amplification of cDNA Ends.
- Assembly PCR - Assembly PCR is the completely artificial synthesis of long gene products by performing PCR on a pool of long oligonucleotides with short overlapping segments. The oligonucleotides alternate between sense and antisense directions, and the overlapping segments serve to order the PCR fragments so that they selectively produce their final product.
- Asymmetric PCR - Asymmetric PCR is used to preferentially amplify one strand of the original DNA more than the other. It finds use in some types of sequencing and hybridization probing where having only one of the two complementary stands is required. PCR is carried out as usual, but with a great excess of the primers for the chosen strand. Due to the slow (arithmetic) amplification later in the reaction after the limiting primer has been used up, extra cycles of PCR are required. A recent modification on this process, known as Linear-After-The-Exponential-PCR (LATE-PCR), uses a limiting primer with a higher melting temperature (Tm) than the excess primer to maintain reaction efficiency as the limiting primer concentration decreases mid-reaction.
- Quantitative PCR - Q-PCR (Quantitative PCR) is used to measure the quantity of a PCR product (preferably real-time). It is the method of choice to quantitatively measure starting amounts of DNA, cDNA or RNA. Q-PCR is commonly used to determine whether a DNA sequence is present in a sample and the number of its copies in the sample. The method with currently the highest level of accuracy is Quantitative real-time PCR. It is often confusingly known as RT-PCR (Real Time PCR) or RQ-PCR. QRT-PCR or RTQ-PCR are more appropriate contractions. RT-PCR commonly refers to reverse transcription PCR (see above), which is often used in conjunction with Q-PCR. QRT-PCR methods use fluorescent dyes, such as Sybr Green, or fluorophore-containing DNA probes, such as TaqMan, to measure the amount of amplified product in real time.
- Touchdown PCR - Touchdown PCR is a variant of PCR that aims to reduce nonspecific background by gradually lowering the annealing temperature as PCR cycling progresses. The annealing temperature at the initial cycles is usually a few degrees above the Tm of the primers used, while at the later cycles, it is a few degrees below the primer Tm. The higher temperatures give greater specificity for primer binding, and the lower temperatures permit more efficient amplification from the specific products formed during the initial cycles.
- Hot-start PCR is a technique that reduces non-specific amplification during the initial set up stages of the PCR. The technique may be performed manually by simply heating the reaction components briefly at the melting temperature (e.g., 95˚C) before adding the polymerase. Specialized enzyme systems have been developed that inhibit the polymerase's activity at ambient temperature, either by the binding of an antibody or by the presence of covalently bound inhibitors that only dissociate after a high-temperature activation step. Hot-start/cold-finish PCR is achieved with new hybrid polymerases that are inactive at ambient temperature and are instantly activated at elongation temperature.
- Colony PCR - Bacterial clones (E.coli) can be rapidly screened for correct DNA vector constructs. Selected bacterial colonies are picked with a sterile toothpick from an agarose plate and dabbed into the master mix or sterile water. Primers (and the master mix) are added, and the PCR is started with an extended time at 95˚C when standard polymerase is used or with a shortened denaturation step at 100˚C and special chimeric DNA polymerase.[5]
- Multiplex-PCR - The use of multiple, unique primer sets within a single PCR reaction to produce amplicons of varying sizes specific to different DNA sequences. By targeting multiple genes at once, additional information may be gained from a single test run that otherwise would require several times the reagents and more time to perform. Annealing temperatures for each of the primer sets must be optimized to work correctly within a single reaction, and amplicon sizes, i.e., their base pair length, should be different enough to form distinct bands when visualized by gel electrophoresis.
- Methylation Specific PCR - Methylation Specific PCR (MSP) is used to detect methylation of CpG islands in genomic DNA. DNA is first treated with sodium bisulfite, which converts unmethylated cytosine bases to uracil, which is recognized by PCR primers as thymine. Two PCR reactions are then carried out on the modified DNA, using primer sets identical except at any CpG islands within the primer sequences. At these points, one primer set recognizes DNA with cytosines to amplify methylated DNA, and one set recognizes DNA with uracil or thymine to amplify unmethylated DNA. MSP using qPCR can also be performed to obtain quantitative rather than qualitative information about methylation.
Recent developments in PCR techniques
- A more recent method which excludes a temperature cycle, but uses enzymes, is helicase-dependent amplification.
- TAIL-PCR, developed by Liu et al. in 1995, is the thermal asymmetric interlaced PCR.
- Meta-PCR, developed by Andrew Wallace, allows to optimize amplification and direct sequence analysis of complex genes. Details at National Genetic Reference Laboratory, Manchester, UK
- Multiplex Ligation-dependent Probe Amplification (MLPA) permits multiple targets to be amplified with only a single primer pair, thus avoiding the resolution limitations of multiplex PCR.
Uses of PCR
PCR can be used for a broad variety of experiments and analyses. Some examples are discussed below.
Genetic fingerprinting
Genetic fingerprinting is a forensic technique used to identify a person by comparing his or her DNA with the DNA in a given sample. An example is blood from a crime scene whose DNA is being genetically compared to DNA from a suspect. The sample may contain only a very small amount of DNA (obtained from a source such as blood, semen, saliva, hair, or other DNA-containing organic material). With the use of PCR, in theory, only a single DNA strand is needed, providing very high sensitivity to this technique, but increasing the risk of confounding results due to possible contamination with, and amplification of, DNA from extraneous sources. There are different PCR-based methods for fingerprinting, summarized in Genetic fingerprinting. The overall pattern of PCR-generated DNA fragments after gel electrophoresis and visualization by ethidium bromide staining (or hybridization with a DNA probe after Southern blotting), can be considered a DNA fingerprint analogous to the fingerprint pattern unique to each individual.
Paternity testing

Although these resulting 'fingerprints' are unique (except for identical twins), genetic relationships, for example, parent-child or siblings, can be determined from two or more genetic fingerprints, which can be used for paternity tests (Fig. 4). A variation of this technique can also be used to determine evolutionary relationships between organisms.
Detection of hereditary diseases
The detection of hereditary diseases in a given genome is a long and difficult process, which can be shortened significantly by using PCR. Each gene in question can easily be amplified through PCR by using the appropriate primers and then sequenced to detect mutations.
Viral diseases, too, can be detected using PCR through amplification of the viral DNA. This analysis is possible right after infection, which can be from several days to several months before actual symptoms occur. Such early diagnoses give physicians a significant lead in treatment.
Cloning genes
Cloning a gene, not to be confused with cloning a whole organism, describes the process of isolating a gene from one organism and then inserting it into another organism (now termed a genetically modified organism (GMO)). PCR is often used to amplify the gene, which can then be inserted into a vector (a vector is a piece of DNA which 'carries' the gene into the GMO) such as a plasmid (a circular DNA molecule) (Fig. 5). The DNA can then be transferred into an organism (the GMO) where the gene and its product can be studied more closely. Expressing a cloned gene (when a gene is expressed the gene product (usually protein or RNA) is produced by the GMO) can also be a way of mass-producing useful proteins, for example medicines or the enzymes in biological washing powders. The incorporation of an affinity tag on a recombinant protein will generate a fusion protein which can be more easily purified by affinity chromatography.

(1) Chromosomal DNA of organism A. (2) PCR. (3) Multiple copies of a single gene from organism A. (4) Insertion of the gene into a plasmid. (5) Plasmid with gene from organism A. (6) Insertion of the plasmid in organism B. (7) Multiplication or expression of the gene, originally from organism A, occurring in organism B.
Mutagenesis
Mutagenesis is a way of introducing changes to the sequence of nucleotides in the DNA. There are situations in which researchers are interested in mutated (changed) copies of a given DNA strand, for example, when trying to assess the function of a particular nucleotide sequence within a gene, intergenic regulatory sequence, or for in in-vitro protein evolution (also known as Directed evolution). Mutations can be introduced into DNA sequences in two fundamentally different ways in the PCR process. Site-directed mutagenesis allows the experimenter to introduce a mutation at a specific location in the DNA strand. Usually, the desired mutation is incorporated in the primers used in the PCR. Random mutagenesis, on the other hand, is based on the use of error-prone polymerases or addition of specific ions (such as Mn2+) to the PCR. In random mutagenesis, the location and nature of the mutations cannot be controlled. One application of random mutagenesis is to analyze structure-function relationships of a protein. By randomly altering a DNA sequence, one can compare the resulting protein with the original and determine the function of each part of the protein.
Analysis of ancient DNA
Using PCR, it becomes possible to analyze DNA that is thousands of years old. PCR techniques have been successfully used on animals, such as a forty-thousand-year-old mammoth, and also on human DNA, in applications ranging from the analysis of Egyptian mummies to the identification of a Russian Tsar,[6].
Genotyping of specific mutations
Through the use of allele-specific PCR, one can easily determine which allele of a mutation or polymorphism an individual has. Here, one of the two primers is common, and would anneal a short distance away from the mutation, while the other anneals right on the variation. The 3' end of the allele-specific primer is modified, to only anneal if it matches one of the alleles. If the mutation of interest is a T or C single nucleotide polymorphism (T/C SNP), one would use two reactions, one containing a primer ending in T, and the other ending in C. The common primer would be the same. Following PCR, these two sets of reactions would be run out on an agarose gel, and the band pattern will tell you if the individual is homozygous T, homozygous C, or heterozygous T/C. This methodology has several applications, such as amplifying certain haplotypes (when certain alleles at 2 or more SNPs occur together on the same chromosome Linkage Disequilibrium) or detection of recombinant chromosomes and the study of meiotic recombination.
Comparison of gene expression
Researchers have used traditional PCR as a way to estimate changes in the amount of a gene's expression. Ribonucleic acid (RNA) is the molecule into which DNA is transcribed prior to making a protein, and those strands of RNA that hold the instructions for protein sequence are known as messenger RNA (mRNA). Once RNA is isolated it can be reverse transcribed back into DNA (complementary DNA or cDNA), at which point DNA-based PCR can be applied to amplify the gene, a method called RT-PCR. The proportion of mRNA transcripts from a given gene in a sample determines the relative proportion of cDNA amplified by PCR from this sample. When cDNA products of the PCR are fractionated on an agarose gel (see Figure 3 above) a band, corresponding to a highly expressed gene, often has higher intensity, due to its containing greater amounts of amplified DNA. This qualitative approach may be suited for a quick analysis of gene expression in differently treated organisms or tissues to identify levels of expression of a gene of interest. A more quantitative RT-PCR method has been developed, called Real-time PCR.
History
Polymerase chain reaction was invented by Kary Mullis [7][8]. He was awarded the Nobel Prize in Chemistry in 1993 for his invention, only seven years after he and his colleagues at Cetus first reduced his proposal to practice.
At the time he thought up PCR in 1983, Mullis was working in Emeryville, California for Cetus Corporation, one of the first biotechnology companies. There, he was charged with making short chains of DNA for other scientists. Mullis has written that he conceived of PCR while cruising along the Pacific Coast Highway one night in his car[7]. He was playing in his mind with a new way of analyzing changes (mutations) in DNA when he realized that he had instead invented a method of amplifying any DNA region through repeated cycles of duplication driven by an enzyme called DNA polymerase.
In Scientific American, Mullis summarized the accomplishment: "Beginning with a single molecule of the genetic material DNA, the PCR can generate 100 billion similar molecules in an afternoon. The reaction is easy to execute. It requires no more than a test tube, a few simple reagents, and a source of heat."[9]
DNA polymerase occurs naturally in living organisms. In cells it functions to duplicate DNA when cells divide in mitosis and meiosis. Polymerase works by binding to a single DNA strand and creating the complementary strand. In the first of many original processes, the enzyme was used in vitro (in a controlled environment outside an organism). The double-stranded DNA was separated into two single strands by heating it to 94°C (201°F). At this temperature, however, the DNA polymerase used at the time were destroyed, so the enzyme had to be replenished after the heating stage of each cycle. The original procedure was very inefficient, since it required a great deal of time, large amounts of DNA polymerase, and continual attention throughout the process.
Later, this original PCR process was greatly improved by the use of DNA polymerase taken from thermophilic bacteria grown in geysers at a temperature of over 110°C (230°F). The DNA polymerase taken from these organisms is stable at high temperatures and, when used in PCR, does not break down when the mixture was heated to separate the DNA strands. Since there was no longer a need to add new DNA polymerase for each cycle, the process of copying a given DNA strand could be simplified and automated.
One of the first thermostable DNA polymerases was obtained from Thermus aquaticus and was called "Taq." Taq polymerase is widely used in current PCR practice. A disadvantage of Taq is that it sometimes makes mistakes when copying DNA, leading to mutations (errors) in the DNA sequence, since it lacks 3'→5' proofreading exonuclease activity. Polymerases such as Pwo or Pfu, obtained from Archaea, have proofreading mechanisms (mechanisms that check for errors) and can significantly reduce the number of mutations that occur in the copied DNA sequence. However these enzymes polymerise DNA at a much slower rate than Taq. Combinations of both Taq and Pfu are available nowadays that provide both high processivity (fast polymerisation) and high fidelity (accurate duplication of DNA).
PCR has been performed on DNA larger than 10 kilobases, but the average PCR is only several hundred to a few thousand bases of DNA. The problem with long PCR is that there is a balance between accuracy and processivity of the enzyme. Usually, the longer the fragment, the greater the probability of errors.
Patent wars
The PCR technique was patented by Cetus Corporation, where Mullis worked when he invented the technique in 1983. The Taq polymerase enzyme was also covered by patents. There have been several high-profile lawsuits related to the technique, including an unsuccessful lawsuit brought by DuPont. The pharmaceutical company Hoffmann-La Roche purchased the rights to the patents in 1992 and currently holds those that are still protected.
A related patent battle over the Taq polymerase enzyme is still ongoing in several jurisdictions around the world between Roche and Promega. Interestingly, it seems possible that the legal arguments will extend beyond the life of the original PCR and Taq polymerase patents, which expired on March 28, 2005.[10]
References
- ^ The history of PCR: Smithsonian Institution Archives, Institutional History Division. Retrieved 24 June 2006.
- ^ Cheng S, Fockler C, Barnes WM, Higuchi R (1994). "Effective amplification of long targets from cloned inserts and human genomic DNA". Proc Natl Acad Sci. 91: 5695–5699. PMID 8202550.
{{cite journal}}: CS1 maint: multiple names: authors list (link) - ^ Sambrook, Joseph (2001). Molecular Cloning: A Laboratory Manual (3rd ed. ed.). Cold Spring Harbor, N.Y.: Cold Spring Harbor Laboratory Press. ISBN 0-87969-576-5.
{{cite book}}:|edition=has extra text (help); Unknown parameter|coauthors=ignored (|author=suggested) (help) - ^ Pavlov AR, Pavlova NV, Kozyavkin SA, Slesarev AI (2004). "Recent developments in the optimization of thermostable DNA polymerases for efficient applications". Trends Biotechnol. 22: 253–260. PMID 15109812.
{{cite journal}}: CS1 maint: multiple names: authors list (link) - ^ Pavlov AR, Pavlova NV, Kozyavkin SA, Slesarev AI (2006). "Thermostable DNA Polymerases for a Wide Spectrum of Applications: Comparison of a Robust Hybrid TopoTaq to other enzymes". In Kieleczawa J (ed.). DNA Sequencing II: Optimizing Preparation and Cleanup. Jones and Bartlett. pp. pp. 241-257. ISBN 0-7637338-3-0.
{{cite book}}:|pages=has extra text (help); External link in|chapterurl=|chapterurl=ignored (|chapter-url=suggested) (help)CS1 maint: multiple names: authors list (link) - ^ http://photoscience.la.asu.edu/photosyn/courses/BIO_343/lecture/DNAtech.html
- ^ a b Mullis, Kary (1998). Dancing Naked in the Mind Field. New York: Pantheon Books. ISBN 0-679-44255-3.
- ^ Rabinow, Paul (1996). Making PCR: A Story of Biotechnology. Chicago: University of Chicago Press. ISBN 0-226-70146-8.
- ^ Mullis KB. The unusual origin of the polymerase chain reaction. Sci Am 1990;262(4):56-61, 64-5.
- ^ Advice on How to Survive the Taq Wars ¶2: GEN Genetic Engineering News Biobusiness Channel: Article. May 1 2006 (Vol. 26, No. 9).
External links
- primer3 Open-source PCR-primer design software
- Primer3Plus Easy and powerful primer design, based on primer3
- US Patent for PCR
- PCR Animation (a Flash document)
- PCR - Polymerase Chain Reaction Articles, news, bioinformatics, and protocols for PCR.
- PCR Interactive Animation
- Online simulation of PCR processes against sequenced prokaryotes.
- Shockwave Animation of PCR by Dolan DNA Learning Center.
- The PCR Jump Station Information and links on the polymerase chain reaction
- PCR Narrated flash animation
- PCR at Home - Amateur Scientist article in the July 2000 issue of Scientific American
- PCR Animation - PCR and Real-time PCR principles and comparison
- PCR in real life
