Histone H2A
Histone H2A is one of the five main histone proteins involved in the structure of chromatin in eukaryotic cells.
The other histone proteins are: H1, H2B, H3 and H4.

Background
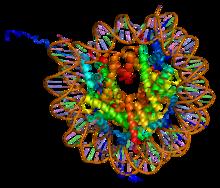
[edit]Histones are proteins that package DNA into nucleosomes.[1] Histones are responsible for maintaining the shape and structure of a nucleosome. One chromatin molecule is composed of at least one of each core histones per 100 base pairs of DNA.[2] There are five families of histones known to date; these histones are termed H1/H5, H2A, H2B, H3, and H4.[3] H2A is considered a core histone, along with H2B, H3 and H4. Core formation first occurs through the interaction of two H2A molecules.[3] Then, H2A forms a dimer with H2B; the core molecule is complete when H3-H4 also attaches to form a tetramer.
Sequence variants
[edit]Histone H2A is composed of non-allelic variants.[4] The term "Histone H2A" is intentionally non-specific and refers to a variety of closely related proteins that vary often by only a few amino acids. Apart from the canonical form, notable variants include H2A.1, H2A.2, H2A.X, and H2A.Z. H2A variants can be explored using "HistoneDB with Variants" database
Changes in variant composition occur in differentiating cells. This was observed in differentiating neurons during synthesis and turnover; changes in variant composition were seen among the H2A.1 histone. The only variant that remained constant in the neural differentiation was variant H2A.Z.[4] H2A.Z is a variant that exchanges with conventional H2A core protein; this variant is important for gene silencing.[5]
Physically, there are small changes on the surface area of the nucleosome that make the histone differ from H2A. Recent research suggests that H2AZ is incorporated into the nucleosome using a Swr1, a Swi2/Snf2- related adenosine triphosphatase.[6]
Another H2A variant that has been identified is H2AX. This variant has a C-terminal extension that is utilized for DNA repair. The method of repair this variant employs is non-homologous end joining. Direct DNA damage can induce changes to the sequence variants. Experiments performed with ionizing radiation linked γ- phosphorylation of H2AX to DNA double-strand break.[7] A large amount of chromatin is involved with each DNA double-strand break; a response to DNA damage is the formation of γ- H2AX.
Lastly, MacroH2A variant is a variant that is similar to H2A; it is encoded by the H2AFY gene. This variant differs from H2A because of the addition of a fold domain in its C-terminal tail. MacroH2A is expressed in the inactive X chromosome in females.[8]
Structure
[edit]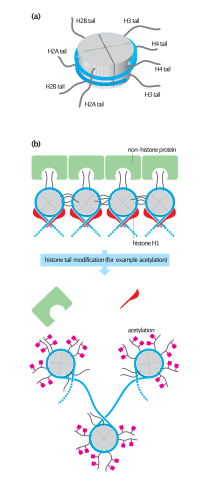
H2A consists of a main globular domain, an N-terminal tail and a C-terminal tail.[9] Both tails are the location of post-translational modification. Thus far, researchers have not identified any secondary structures that arise in the tails. H2A utilizes a protein fold known as the ‘histone fold’. The histone fold is a three-helix core domain that is connected by two loops. This connection forms a ‘handshake arrangement.’ Most notably, this is termed the helix-turn-helix motif, which allows for dimerization with H2B. The ‘histone fold’ is conserved among H2A at the structural level; however the genetic sequence that encodes for this structure differs between variants.[10]
The structure of macroH2A variant was exposed through X-ray crystallography. The conserved domain contains a DNA binding structure and a peptidase fold.[11] The function of this conserved domain remains unknown. Research suggests that this conserved domain may function as an anchor site for Xist DNA or it may also function as a modifying enzyme.
Function
[edit]
DNA Folding: H2A is important for packaging DNA into chromatin. Since H2A packages DNA molecules into chromatin, the packaging process will affect gene expression. H2A has been correlated with DNA modification and epigenetics. H2A plays a major role in determining the overall structure of chromatin. Inadvertently, H2A has been found to regulate gene expression.[10]
DNA modification by H2A occurs in the cell nucleus. Proteins responsible for nuclear import of H2A protein are karyopherin and importin.[12] Recent studies also show that nucleosome assembly protein 1 is also used to transport of H2A into the nucleus so it can wrap DNA. Other functions of H2A have been seen in the histone variant H2A.Z. This variant is associated with gene activation, silencing and suppression of antisense RNA. In addition, when H2A.Z was studied in human and yeast cells, it was used to promote RNA polymerase II recruitment.[13]
Antimicrobial peptide: Histones are conserved eukaryotic cationic proteins present in the cells and are involved in the antimicrobial activities. In vertebrates and invertebrates, Histone H2A variant is reported to be involved in host immune response by acting as antimicrobial peptides (AMPs). H2A are α-helical molecule, amphipathic protein with hydrophobic and hydrophilic residues on opposing sides that enhances the antimicrobial activity of H2A.[14]
DNA damage response
[edit]Site specific ubiquitination of histone H2A has a role in the recruitment of DNA repair proteins to DNA double strand breaks which then may be repaired by either homologous recombination or non-homologous end joining.[15] In the DNA damage response, it is thought that ubiquitination of H2A by the BRCA1/BARD1 heterodimer promotes homologous recombination, and that ubiquitination of H2A by RNF168 protein promotes non-homologous end joining.[15]
Genetics
[edit]H2A is coded by many genes in the human genome, including: H2AFB1, H2AFB2, H2AFB3, H2AFJ, H2AFV, H2AFX, H2AFY, H2AFY2, and H2AFZ. Genetic patterns among the different H2A molecules are mostly conserved among variants. The variability in gene expression exists among the regulatory machinery that manages H2A expression. Researchers studied eukaryotic evolutionary lineages of histone proteins and found diversification among the regulatory genes. The greatest differences were observed in core histone gene cis-regulatory sequence motifs and associated protein factors. Variability in gene sequence was seen in bacterial, fungi, plant, and mammalian genes.[10]
One variant of H2A protein is H2ABbd (Barr body deficient) variant. This variant is composed of a different genetic sequence compared to H2A. The variant functions with transcriptionally active domains.[10] Other variations associated with H2ABbd are located within its C-terminus. H2ABbd has a shorter C-terminal domain compared to the large C-terminal found on H2A. The two C terminals are about 48% identical. H2ABbd functions with active chromosomes. Thus far, it is missing from Xi chromosomes in fibroblast cells. Lastly, it found to be associated with acetylated H4.[16]
Different functions of H2A.Z compared to H2A are correlated with genetic differences between H2A and the variant. Resistance to nucleosomes occurs in H2A.Z by binding to H1 factor. H2A.Z gene is an essential gene in yeast and it is denoted as Htz1. Comparatively, vertebrates have two H2A.Z genes.[10] These genes, H2A.Z1 and H2A.Z2 encode for proteins that differ from H2A.Z by three residues. At first researchers figured that these genes were redundant; however, when a mutant H2A.Z1 was created, it resulted in lethality during mammalian tests.[16] Therefore, H2A.Z1 is an essential gene. On the other hand, researchers have not identified the function of H2A.Z2 variant. It is known that it is transcribed in mammals and this gene expression is conserved among mammalian species. This conservation suggests that the gene is functional.[16] When studying H2A.Z in plants species, the protein different among residues from species to species. These differences contribute to differences in cell-cycle regulation.[16] This phenomenon was only observed in plants.
Phylogenetic trees were created to show the divergence of variants from their ancestors. The divergence of variant, H2A.X, from H2A occurred at multiple origins in a phylogenetic tree. Acquisition of the phosphorylation motif was consistent with the many origins of H2A that arose from an ancestral H2A.X. Finally, the presence of H2A.X and absence of H2A in fungi leads researchers to believe that H2A.X was the original ancestor of the histone protein H2A [10]
Modification of H2A
[edit]H2A modification is under current research. However, modification of H2A does occur. Serine phosphorylation sites have been identified on H2A. Threonine O-GlcNAc has also been identified on H2A. Large differences exist between the modified residues of H2A variants. For example, H2ABbd lacks modified residues that exist in H2A.[16] The differences in modification change the function of H2ABbd compared to H2A. As previously mentioned, variant H2AX was found to function in DNA repair. This function is dependent upon the phosphorylation of H2AX C-terminal.[7] Once H2AX becomes phosphorylated, it can function in DNA repair. The H2A.X variant differs from H2A through modification. The C-terminal of H2A.X contains an additional motif compared to H2A. The motif that is added is Ser-Gln-(Glu/Asp)- (hydrophobic residue).[16] The motif becomes heavily phosphorylated at the serine residue; if this phosphorylation occurs the variant becomes γH2A.X. Phosphorylation occurs due to dsDNA breaks.[16] Modification on histone proteins can sometimes result in a change in function. Different H2A variants were exploited to have different functions, genetic sequences, and modifications.
See also
[edit]References
[edit]- ^ Youngson RM (2006). Collins dictionary of human biology. Glasgow [Scotland]: Collins. ISBN 978-0-00-722134-9.
- ^ Khorasanizadeh S (January 2004). "The nucleosome: from genomic organization to genomic regulation". Cell. 116 (2): 259–72. doi:10.1016/s0092-8674(04)00044-3. PMID 14744436. S2CID 15504162.
- ^ a b Cox MM, Lehninger AL, Nelson DL (2005). Lehninger principles of biochemistry (4th ed.). New York: W.H. Freeman. ISBN 978-0-7167-4339-2.
- ^ a b Bosch A, Suau P (November 1995). "Changes in core histone variant composition in differentiating neurons: the roles of differential turnover and synthesis rates". European Journal of Cell Biology. 68 (3): 220–5. PMID 8603674.
- ^ Suto RK, Clarkson MJ, Tremethick DJ, Luger K (December 2000). "Crystal structure of a nucleosome core particle containing the variant histone H2A.Z". Nature Structural Biology. 7 (12): 1121–4. doi:10.1038/81971. PMID 11101893. S2CID 5966635.
- ^ Mizuguchi G, Shen X, Landry J, Wu WH, Sen S, Wu C (January 2004). "ATP-driven exchange of histone H2AZ variant catalyzed by SWR1 chromatin remodeling complex". Science. 303 (5656): 343–8. Bibcode:2004Sci...303..343M. doi:10.1126/science.1090701. PMID 14645854. S2CID 9881829.
- ^ a b Jakob B, Splinter J, Conrad S, Voss KO, Zink D, Durante M, Löbrich M, Taucher-Scholz G (August 2011). "DNA double-strand breaks in heterochromatin elicit fast repair protein recruitment, histone H2AX phosphorylation and relocation to euchromatin". Nucleic Acids Research. 39 (15): 6489–99. doi:10.1093/nar/gkr230. PMC 3159438. PMID 21511815.
- ^ Costanzi C, Pehrson JR (June 1998). "Histone macroH2A1 is concentrated in the inactive X chromosome of female mammals". Nature. 393 (6685): 599–601. Bibcode:1998Natur.393..599C. doi:10.1038/31275. PMID 9634239. S2CID 205001095.
- ^ Ghoneim M, Fuchs HA, Musselman CA (2021). "Histone Tail Conformations: A Fuzzy Affair with DNA". Trends Biochem Sci. 46 (7): 564–578. doi:10.1016/j.tibs.2020.12.012. PMC 8195839. PMID 33551235.
{{cite journal}}: CS1 maint: multiple names: authors list (link) - ^ a b c d e f Mariño-Ramírez L, Jordan IK, Landsman D (2006). "Multiple independent evolutionary solutions to core histone gene regulation". Genome Biology. 7 (12): R122. doi:10.1186/gb-2006-7-12-r122. PMC 1794435. PMID 17184543.
- ^ Allen MD, Buckle AM, Cordell SC, Löwe J, Bycroft M (July 2003). "The crystal structure of AF1521 a protein from Archaeoglobus fulgidus with homology to the non-histone domain of macroH2A". Journal of Molecular Biology. 330 (3): 503–11. doi:10.1016/s0022-2836(03)00473-x. PMID 12842467.
- ^ Mosammaparast N, Ewart CS, Pemberton LF (December 2002). "A role for nucleosome assembly protein 1 in the nuclear transport of histones H2A and H2B". The EMBO Journal. 21 (23): 6527–38. doi:10.1093/emboj/cdf647. PMC 136951. PMID 12456659.
- ^ Mariño-Ramírez L, Levine KM, Morales M, Zhang S, Moreland RT, Baxevanis AD, Landsman D (2011). "The Histone Database: an integrated resource for histones and histone fold-containing proteins". Database. 2011: bar048. doi:10.1093/database/bar048. PMC 3199919. PMID 22025671.
- ^ Arockiaraj J, Gnanam AJ, Kumaresan V, Palanisamy R, Bhatt P, Thirumalai MK, Roy A, Pasupuleti M, Kasi M (November 2013). "An unconventional antimicrobial protein histone from freshwater prawn Macrobrachium rosenbergii: analysis of immune properties". Fish & Shellfish Immunology. 35 (5): 1511–22. doi:10.1016/j.fsi.2013.08.018. PMID 23994279.
- ^ a b Uckelmann M, Sixma TK. Histone ubiquitination in the DNA damage response. DNA Repair (Amst). 2017 Aug;56:92-101. doi: 10.1016/j.dnarep.2017.06.011. Epub 2017 Jun 9. PMID 28624371
- ^ a b c d e f g Talbert PB, Henikoff S (April 2010). "Histone variants--ancient wrap artists of the epigenome". Nature Reviews. Molecular Cell Biology. 11 (4): 264–75. doi:10.1038/nrm2861. PMID 20197778. S2CID 10934412.
External links
[edit]- Nextbio Archived 2020-05-30 at the Wayback Machine
