Telomere
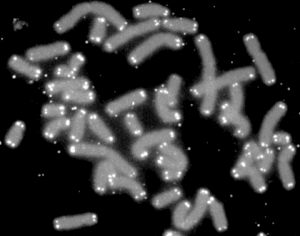
A telomere (/ˈtɛləmɪər/ or /ˈtiləmɪər/) is a region of repetitive nucleotide sequences at each end of a chromosome, which protects the end of the chromosome from deterioration or from fusion with neighboring chromosomes. Its name is derived from the Greek nouns telos (τέλος) "end" and merοs (μέρος, root: μερ-) "part". For vertebrates, the sequence of nucleotides in telomeres is 5′-TTAGGG-3′,[1] with the complementary DNA strand being 3′-AATCCC-5′, with a single-stranded TTAGGG overhang.[2] This sequence of TTAGGG is repeated approximately 2,500 times in humans.[3] In humans, average telomere length declines from about 11 kilobases at birth[4] to fewer than 4 kilobases in old age,[5] with the average rate of decline being greater in men than in women.[6]
During chromosome replication, the enzymes that duplicate DNA cannot continue their duplication all the way to the end of a chromosome, so in each duplication the end of the chromosome is shortened (this is because the synthesis of Okazaki fragments requires RNA primers attaching ahead on the lagging strand). The telomeres are disposable buffers at the ends of chromosomes which are truncated during cell division; their presence protects the genes before them on the chromosome from being truncated instead. The telomeres themselves are protected by a complex of shelterin proteins, as well as by the RNA that telomeric DNA encodes (TERRA).
Over time, due to each cell division, the telomere ends become shorter.[7] They are replenished by an enzyme, telomerase reverse transcriptase.[8]
Discovery
In the early 1970s, Russian theorist Alexei Olovnikov first recognized that chromosomes could not completely replicate their ends. Building on this, and to accommodate Leonard Hayflick's idea of limited somatic cell division, Olovnikov suggested that DNA sequences are lost every time a cell replicates until the loss reaches a critical level, at which point cell division ends.[9]
In 1975–1977, Elizabeth Blackburn, working as a postdoctoral fellow at Yale University with Joseph G. Gall, discovered the unusual nature of telomeres, with their simple repeated DNA sequences composing chromosome ends.[10] Blackburn, Carol Greider, and Jack Szostak were awarded the 2009 Nobel Prize in Physiology or Medicine for the discovery of how chromosomes are protected by telomeres and the enzyme telomerase.[11]
In 1983, Barbara McClintock, an American cytogeneticist and the first woman to receive an unshared Nobel Prize in Physiology or Medicine, received the Nobel Prize for observing that the chromosomes lacking end parts became "sticky" and hypothesized the existence of a special structure at the chromosome tip that would maintain chromosome stability.[12]
Nature and function
Structure, function and evolutionary biology

Telomeres are repetitive nucleotide sequences located at the termini of linear chromosomes of most eukaryotic organisms. For vertebrates, the sequence of nucleotides in telomeres is TTAGGG.[13] Most prokaryotes, having circular chromosomes rather than linear, do not have telomeres.[14] Telomeres compensate for incomplete semi-conservative DNA replication at chromosomal ends.[15] A protein complex known as shelterin serves to protect the ends of telomeres from being recognised as double-strand breaks by inhibiting homologous recombination (HR) and non-homologous end joining (NHEJ).[16][17][18]
In most prokaryotes, chromosomes are circular and thus do not have ends to suffer premature replication termination. A small fraction of bacterial chromosomes (such as those in Streptomyces, Agrobacterium, and Borrelia) are linear and possess telomeres, which are very different from those of the eukaryotic chromosomes in structure and functions. The known structures of bacterial telomeres take the form of proteins bound to the ends of linear chromosomes, or hairpin loops of single-stranded DNA at the ends of the linear chromosomes.[19]
While replicating DNA, the eukaryotic DNA replication enzymes (the DNA polymerase protein complex) cannot replicate the sequences present at the ends of the chromosomes (or more precisely the chromatid fibres). Hence, these sequences and the information they carry may get lost. This is the reason telomeres are so important in context of successful cell division: They "cap" the end-sequences and themselves get lost in the process of DNA replication. But the cell has an enzyme called telomerase, which carries out the task of adding repetitive nucleotide sequences to the ends of the DNA. Telomerase "replenishes" the telomere "cap." In most multicellular eukaryotic organisms, telomerase is active only in germ cells, some types of stem cells such as embryonic stem cells, and certain white blood cells. Telomerase can be reactivated and telomeres reset back to an embryonic state by somatic cell nuclear transfer.[20] The steady shortening of telomeres with each replication in somatic (body) cells may have a role in senescence and in the prevention of cancer.[21][22] This is because the telomeres act as a sort of time-delay "fuse", eventually running out after a certain number of cell divisions and resulting in the eventual loss of vital genetic information from the cell's chromosome with future divisions.[23]
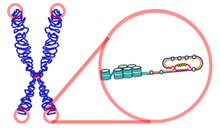
Telomere length varies greatly between species, from approximately 300 base pairs in yeast[24] to many kilobases in humans, and usually is composed of arrays of guanine-rich, six- to eight-base-pair-long repeats. Eukaryotic telomeres normally terminate with 3′ single-stranded-DNA overhang, which is essential for telomere maintenance and capping. Multiple proteins binding single- and double-stranded telomere DNA have been identified.[25] These function in both telomere maintenance and capping. Telomeres form large loop structures called telomere loops, or T-loops. Here, the single-stranded DNA curls around in a long circle, stabilized by telomere-binding proteins.[26] At the very end of the T-loop, the single-stranded telomere DNA is held onto a region of double-stranded DNA by the telomere strand disrupting the double-helical DNA, and base pairing to one of the two strands. This triple-stranded structure is called a displacement loop or D-loop.[27]
Role in the cell cycle
Telomere shortening in humans can induce replicative senescence, which blocks cell division. This mechanism appears to prevent genomic instability and development of cancer in human aged cells by limiting the number of cell divisions. However, shortened telomeres impair immune function that might also increase cancer susceptibility.[28] If telomeres become too short, they have the potential to unfold from their presumed closed structure. The cell may detect this uncapping as DNA damage and then either stop growing, enter cellular old age (senescence), or begin programmed cell self-destruction (apoptosis) depending on the cell's genetic background (p53 status). Uncapped telomeres also result in chromosomal fusions. Since this damage cannot be repaired in normal somatic cells, the cell may even go into apoptosis. Many aging-related diseases are linked to shortened telomeres. Organs deteriorate as more and more of their cells die off or enter cellular senescence.

Shelterin
At the very distal end of the telomere is a 300 base pair single-stranded portion, which forms the T-loop. This loop is analogous to a knot, which stabilizes the telomere, preventing the telomere ends from being recognized as break points by the DNA repair machinery. Should non-homologous end joining occur at the telomeric ends, chromosomal fusion will result. The T-loop is held together by several proteins, the most notable ones being TRF1, TRF2, POT1, RAP1, and TIN2, collectively referred to as the shelterin complex. In humans, the shelterin complex consists of six proteins identified as TRF1, TRF2, TIN2, POT1, TPP1, and RAP1.[16]
Shortening
Telomeres shorten in part because of the end replication problem that is exhibited during DNA replication in eukaryotes only. Because DNA replication does not begin at either end of the DNA strand, but starts in the center, and considering that all known DNA polymerases read the template strand in the 3' to 5' direction, one finds a leading and a lagging strand on the DNA molecule being replicated.
On the leading strand, DNA polymerase can make a complementary DNA strand without any difficulty because it reads the template strand from 3' to 5'. However, there is a problem going in the other direction on the lagging strand. To counter this, short sequences of RNA acting as primers attach to the lagging strand a short distance ahead of where the initiation site was. The DNA polymerase can start replication at that point and go to the end of the initiation site. This causes the formation of Okazaki fragments. More RNA primers attach further on the DNA strand and DNA polymerase comes along and continues to make a new DNA strand.

Eventually, the last RNA primer attaches, and DNA polymerase, RNA nuclease, and DNA ligase come along to convert the RNA (of the primers) to DNA and to seal the gaps in between the Okazaki fragments. But, in order to change RNA to DNA, there must be another DNA strand in front of the RNA primer. This happens at all the sites of the lagging strand, but it does not happen at the end where the last RNA primer is attached. Ultimately, that RNA is destroyed by enzymes that degrade any RNA left on the DNA. Thus, a section of the telomere is lost during each cycle of replication at the 5' end of the lagging strand's daughter.
However, test-tube studies have shown that telomeres are highly susceptible to oxidative stress. There is evidence that oxidative stress-mediated DNA damage is an important determinant of telomere shortening.[29] Telomere shortening due to free radicals explains the difference between the estimated loss per division because of the end-replication problem (c. 20 bp) and actual telomere shortening rates (50–100 bp), and has a greater absolute impact on telomere length than shortening caused by the end-replication problem. Population-based studies have also indicated an interaction between anti-oxidant intake and telomere length. In the Long Island Breast Cancer Study Project (LIBCSP), authors found a moderate increase in breast cancer risk among women with the shortest telomeres and lower dietary intake of beta carotene, vitamin C or E.[30] These results [31] suggest that cancer risk due to telomere shortening may interact with other mechanisms of DNA damage, specifically oxidative stress.
Telomere shortening is associated with aging, mortality and aging-related diseases. Normal aging is associated with telomere shortening in both humans and mice, and studies on genetically modified animal models suggest causal links between telomere erosion and aging.[32] However, it is not known whether short telomeres are just a sign of cellular age or if they themselves actually contribute to the aging process.[33]
The age of a father plays a role in the length of a child’s telomeres, which has evolutionary implications. Although leukocyte telomeres shorten with age, sperm telomeres lengthen with age. Shorter telomeres are theorized to impose lower energy costs (due to less replication) but also have immune system-related and other aging- and disease-related costs, so the effect of paternal age on telomere length might be an adaptation to increase the chances that the child will be fit for the environment they’re born into.[34][35]
Potential effect of psychological stress
Meta-analyses found that increased perceived psychological stress was associated with a small decrease in telomere length—but that these associations attenuate to no significant association when accounting for publication bias. The literature concerning telomeres as integrative biomarkers of exposure to stress and adversity is dominated by cross-sectional and correlational studies, which makes causal interpretation problematic.[31][36] A 2020 review argued that the relationship between psychosocial stress and telomere length appears strongest for stress experienced in utero or early life.[37]
Lengthening

The phenomenon of limited cellular division was first observed by Leonard Hayflick, and is now referred to as the Hayflick limit.[38][39] Significant discoveries were subsequently made by a group of scientists organized at Geron Corporation by Geron's founder Michael D. West, that tied telomere shortening with the Hayflick limit.[40] The cloning of the catalytic component of telomerase enabled experiments to test whether the expression of telomerase at levels sufficient to prevent telomere shortening was capable of immortalizing human cells. Telomerase was demonstrated in a 1998 publication in Science to be capable of extending cell lifespan, and now is well-recognized as capable of immortalizing human somatic cells.[41]
It is becoming apparent that reversing shortening of telomeres through temporary activation of telomerase may be a potent means to slow aging. The reason that this would extend human life is because it would extend the Hayflick limit. Three routes have been proposed to reverse telomere shortening: drugs, gene therapy, or metabolic suppression, so-called torpor/hibernation. So far these ideas have not been proven in humans, but it has been demonstrated that telomere shortening is reversed in hibernation and aging is slowed (Turbill, et al. 2012 & 2013) and that hibernation prolongs life-span (Lyman et al. 1981). It has also been demonstrated that telomere extension has successfully reversed some signs of aging in laboratory mice [42][43] and the nematode worm species Caenorhabditis elegans.[44] It has been hypothesized that longer telomeres and especially telomerase activation might cause increased cancer (e.g. Weinstein and Ciszek, 2002[45]). However, longer telomeres might also protect against cancer, because short telomeres are associated with cancer. It has also been suggested that longer telomeres might cause increased energy consumption.[28]
Techniques to extend telomeres could be useful for tissue engineering, because they might permit healthy, noncancerous mammalian cells to be cultured in amounts large enough to be engineering materials for biomedical repairs.
Two studies on long-lived seabirds demonstrate that the role of telomeres is far from being understood. In 2003, scientists observed that the telomeres of Leach's storm-petrel (Oceanodroma leucorhoa) seem to lengthen with chronological age, the first observed instance of such behaviour of telomeres.[46] In 2006, Juola et al.[47] reported that in another unrelated, long-lived seabird species, the great frigatebird (Fregata minor), telomere length did decrease until at least c. 40 years of age (i.e. probably over the entire lifespan), but the speed of decrease slowed down massively with increasing ages, and that rates of telomere length decrease varied strongly between individual birds. They concluded that in this species (and probably in frigatebirds and their relatives in general), telomere length could not be used to determine a bird's age sufficiently well. Thus, it seems that there is much more variation in the behavior of telomere length than initially believed.
Furthermore, Gomes et al. found, in a study of the comparative biology of mammalian telomeres, that telomere length of different mammalian species correlates inversely, rather than directly, with lifespan, and they concluded that the contribution of telomere length to lifespan remains controversial.[48] Harris et al. found little evidence that, in humans, telomere length is a significant biomarker of normal aging with respect to important cognitive and physical abilities.[49] Gilley and Blackburn tested whether cellular senescence in paramecium is caused by telomere shortening, and found that telomeres were not shortened during senescence.[50]
Sequences
Known, up-to-date telomere nucleotide sequences are listed in Telomerase Database website.
| Group | Organism | Telomeric repeat (5' to 3' toward the end) |
|---|---|---|
| Vertebrates | Human, mouse, Xenopus | TTAGGG |
| Filamentous fungi | Neurospora crassa | TTAGGG |
| Slime moulds | Physarum, Didymium | TTAGGG |
| Dictyostelium | AG(1-8) | |
| Kinetoplastid protozoa | Trypanosoma, Crithidia | TTAGGG |
| Ciliate protozoa | Tetrahymena, Glaucoma | TTGGGG |
| Paramecium | TTGGG(T/G) | |
| Oxytricha, Stylonychia, Euplotes | TTTTGGGG | |
| Apicomplexan protozoa | Plasmodium | TTAGGG(T/C) |
| Higher plants | Arabidopsis thaliana | TTTAGGG |
| Cestrum elegans | TTTTTTAGGG[51] | |
| Allium | CTCGGTTATGGG[52] | |
| Zostera marina | TTAGGG[53] | |
| Green algae | Chlamydomonas | TTTTAGGG |
| Insects | Bombyx mori | TTAGG |
| Roundworms | Ascaris lumbricoides | TTAGGC |
| Fission yeasts | Schizosaccharomyces pombe | TTAC(A)(C)G(1-8) |
| Budding yeasts | Saccharomyces cerevisiae | TGTGGGTGTGGTG (from RNA template) or G(2-3)(TG)(1-6)T (consensus) |
| Saccharomyces castellii | TCTGGGTG | |
| Candida glabrata | GGGGTCTGGGTGCTG | |
| Candida albicans | GGTGTACGGATGTCTAACTTCTT | |
| Candida tropicalis | GGTGTA[C/A]GGATGTCACGATCATT | |
| Candida maltosa | GGTGTACGGATGCAGACTCGCTT | |
| Candida guillermondii | GGTGTAC | |
| Candida pseudotropicalis | GGTGTACGGATTTGATTAGTTATGT | |
| Kluyveromyces lactis | GGTGTACGGATTTGATTAGGTATGT |
Research on disease risk
This section needs more reliable medical references for verification or relies too heavily on primary sources. (March 2018) |  |
Telomeres are critical for maintaining genomic integrity and may be factors for age-related diseases.[54] Laboratory studies show that telomere dysfunction or shortening is commonly acquired due process of cellular aging and tumor development.[54][55] Short telomeres can lead to genomic instability, chromosome loss and the formation of non-reciprocal translocations; and telomeres in tumor cells and their precursor lesions are significantly shorter than surrounding normal tissue.[56][57]
Observational studies have found shortened telomeres in many types of experimental cancers.[58] In addition, people with cancer have been found to possess shorter leukocyte telomeres than healthy controls.[59] Recent meta-analyses suggest 1.4 to 3.0 fold increased risk of cancer for those with the shortest vs. longest telomeres.[60][61] However, the increase in risk varies by age, sex, tumor type, and differences in lifestyle factors.[58]
Measurement
Several techniques are currently employed to assess average telomere length in eukaryotic cells. One method is the Terminal Restriction Fragment (TRF) southern blot.[62][63] A Real-Time PCR assay for telomere length involves determining the Telomere-to-Single Copy Gene (T/S)ratio, which is demonstrated to be proportional to the average telomere length in a cell.[64].
Tools have also been developed to estimate the length of telomere from whole genome sequencing (WGS) experiments. Amongst these are TelSeq[65], telomerecat[66] and telomereHunter[67]. Length estimation from WGS typically works by differentiating telomere sequencing reads and then inferring the length of telomere that produced that number of reads. These methods have been shown to correlate with preexisting methods of estimation such as PCR and TRF. Flow-FISH is used to quantify the length of telomeres in human white blood cells. A semi-automated method for measuring the average length of telomeres with Flow FISH was published in Nature Protocols in 2006.[68]
While multiple companies offer telomere length measurement services, the utility of these measurements for widespread clinical or personal use has been questioned.[69][70] Nobel Prize winner Elizabeth Blackburn, who was co-founder of one company, promoted the clinical utility of telomere length measures.[71]
Ectothermic telomeres
Most research on telomere length and regulation, and its relationship to cancer and ageing, has been performed on mammals, especially humans, which have little or no somatic telomerase production. Ectotherms are significantly more likely than endotherms to have variation in somatic telomerase expression. For instance, in many fish, telomerase occurs throughout the body (and associated with this, telomere length is roughly the same across all its tissue). Studies on ectotherms, and other non-mammalian organisms, show that there’s no single universal model of telomere erosion; rather, there is wide variation in relevant dynamics across Metazoa, and even within smaller taxonomic groups these patterns appear diverse. Due to the different reproductive timelines of some ectotherms, selection on disease is relevant for a much larger fraction of these creatures’ lives than it is for mammals, so early- and late-life telomere length, and their possible links to cancer, seem especially important in these species from a life history theory point of view.[72]
See also
- Biological clock
- Epigenetic clock
- Centromere
- DNA damage theory of aging
- Immortality
- Maximum life span
- Rejuvenation (aging)
- Senescence, biological aging
- Tankyrase
- Telomere-binding protein
References
- ^ Biochemistry, Lippincott's Illustrated Reviews, 6th edition, Richard Harvey, 2014, page 407.
- ^ Witzany G (2008). "The viral origins of telomeres, telomerases and their important role in eukaryogenesis and genome maintenance". Biosemiotics. 1 (2): 191–206. doi:10.1007/s12304-008-9018-0. S2CID 207415262.
- ^ Sadava, D., Hillis, D., Heller, C., & Berenbaum, M. (2011). Life: The science of biology (9th ed.), Sunderland, MA: Sinauer Associates Inc.[page needed]
- ^ Okuda K, Bardeguez A, Gardner JP, Rodriguez P, Ganesh V, Kimura M, et al. (September 2002). "Telomere length in the newborn" (PDF). Pediatric Research. 52 (3): 377–81. doi:10.1203/00006450-200209000-00012. PMID 12193671. S2CID 4004959.
- ^ Arai Y, Martin-Ruiz CM, Takayama M, Abe Y, Takebayashi T, Koyasu S, et al. (October 2015). "Inflammation, But Not Telomere Length, Predicts Successful Ageing at Extreme Old Age: A Longitudinal Study of Semi-supercentenarians". EBioMedicine. 2 (10): 1549–58. doi:10.1016/j.ebiom.2015.07.029. PMC 4634197. PMID 26629551.
- ^ Dalgård C, Benetos A, Verhulst S, Labat C, Kark JD, Christensen K, et al. (October 2015). "Leukocyte telomere length dynamics in women and men: menopause vs age effects". International Journal of Epidemiology. 44 (5): 1688–95. doi:10.1093/ije/dyv165. PMC 4681111. PMID 26385867.
- ^ Passarge, Eberhard. Color atlas of genetics, 2007.
- ^ "TERT gene". Genetics Home Reference. Retrieved 2018-11-11.
- ^ Mender I, Shay JW (November 2015). "Telomerase Repeated Amplification Protocol (TRAP)". Bio-Protocol. 5 (22): e1657. doi:10.21769/bioprotoc.1657. PMC 4863463. PMID 27182535.
- ^ Blackburn EH, Gall JG (March 1978). "A tandemly repeated sequence at the termini of the extrachromosomal ribosomal RNA genes in Tetrahymena". Journal of Molecular Biology. 120 (1): 33–53. doi:10.1016/0022-2836(78)90294-2. PMID 642006.
- ^ "Elizabeth H. Blackburn, Carol W. Greider, Jack W. Szostak: The Nobel Prize in Physiology or Medicine 2009". Nobel Foundation. 2009-10-05. Retrieved 2012-06-12.
- ^ "Barbara McClintock: The Nobel Prize in Physiology or Medicine 1983". Nobel Foundation. 1983. Retrieved 10 March 2018.
- ^ Meyne J, Ratliff RL, Moyzis RK (September 1989). "Conservation of the human telomere sequence (TTAGGG)n among vertebrates". Proceedings of the National Academy of Sciences of the United States of America. 86 (18): 7049–53. Bibcode:1989PNAS...86.7049M. doi:10.1073/pnas.86.18.7049. PMC 297991. PMID 2780561.
- ^ Nelson, David L.; Lehninger, Albert L.; Cox, Michael M. (2008). Lehninger principles of biochemistry (5th ed.). New York: W.H. Freeman. ISBN 9780716771081. OCLC 191854286.
{{cite book}}: Unknown parameter|name-list-format=ignored (|name-list-style=suggested) (help) - ^ Webb CJ, Wu Y, Zakian VA (June 2013). "DNA repair at telomeres: keeping the ends intact". Cold Spring Harbor Perspectives in Biology. 5 (6): a012666. doi:10.1101/cshperspect.a012666. PMC 3660827. PMID 23732473.
- ^ a b Martínez P, Blasco MA (October 2010). "Role of shelterin in cancer and aging". Aging Cell. 9 (5): 653–66. doi:10.1111/j.1474-9726.2010.00596.x. PMID 20569239.
- ^ Evans SK, Lundblad V (October 2000). "Positive and negative regulation of telomerase access to the telomere". Journal of Cell Science. 113 Pt 19: 3357–64. PMID 10984427.
- ^ Ferreira MG, Miller KM, Cooper JP (January 2004). "Indecent exposure: when telomeres become uncapped". Molecular Cell. 13 (1): 7–18. doi:10.1016/S1097-2765(03)00531-8. PMID 14731390.
- ^ Maloy, Stanley (July 12, 2002). "Bacterial Chromosome Structure". Retrieved 2008-06-22.
{{cite web}}: Unknown parameter|name-list-format=ignored (|name-list-style=suggested) (help) - ^ Lanza RP, Cibelli JB, Blackwell C, Cristofalo VJ, Francis MK, Baerlocher GM, et al. (April 2000). "Extension of cell life-span and telomere length in animals cloned from senescent somatic cells". Science. 288 (5466): 665–9. Bibcode:2000Sci...288..665L. doi:10.1126/science.288.5466.665. PMID 10784448. S2CID 37387314.
- ^ Shay JW, Wright WE (May 2005). "Senescence and immortalization: role of telomeres and telomerase". Carcinogenesis. 26 (5): 867–74. doi:10.1093/carcin/bgh296. PMID 15471900.
- ^ Wai LK (July 2004). "Telomeres, telomerase, and tumorigenesis--a review". MedGenMed. 6 (3): 19. PMC 1435592. PMID 15520642.
- ^ Greider CW (August 1990). "Telomeres, telomerase and senescence". BioEssays. 12 (8): 363–9. doi:10.1002/bies.950120803. PMID 2241933.
- ^ Shampay J, Szostak JW, Blackburn EH (1984). "DNA sequences of telomeres maintained in yeast". Nature. 310 (5973): 154–7. Bibcode:1984Natur.310..154S. doi:10.1038/310154a0. PMID 6330571. S2CID 4360698.
- ^ Williams TL, Levy DL, Maki-Yonekura S, Yonekura K, Blackburn EH (November 2010). "Characterization of the yeast telomere nucleoprotein core: Rap1 binds independently to each recognition site". The Journal of Biological Chemistry. 285 (46): 35814–24. doi:10.1074/jbc.M110.170167. PMC 2975205. PMID 20826803.
{{cite journal}}: CS1 maint: unflagged free DOI (link) - ^ Griffith JD, Comeau L, Rosenfield S, Stansel RM, Bianchi A, Moss H, de Lange T (May 1999). "Mammalian telomeres end in a large duplex loop". Cell. 97 (4): 503–14. doi:10.1016/S0092-8674(00)80760-6. PMID 10338214. S2CID 721901.
- ^ Burge S, Parkinson GN, Hazel P, Todd AK, Neidle S (2006). "Quadruplex DNA: sequence, topology and structure". Nucleic Acids Research. 34 (19): 5402–15. doi:10.1093/nar/gkl655. PMC 1636468. PMID 17012276.
- ^ a b Eisenberg DT (2011). "An evolutionary review of human telomere biology: the thrifty telomere hypothesis and notes on potential adaptive paternal effects". American Journal of Human Biology. 23 (2): 149–67. doi:10.1002/ajhb.21127. PMID 21319244. S2CID 5540894.
- ^ Richter T, von Zglinicki T (November 2007). "A continuous correlation between oxidative stress and telomere shortening in fibroblasts". Experimental Gerontology. 42 (11): 1039–42. doi:10.1016/j.exger.2007.08.005. PMID 17869047. S2CID 23961767.
- ^ Shen J, Gammon MD, Terry MB, Wang Q, Bradshaw P, Teitelbaum SL, et al. (April 2009). "Telomere length, oxidative damage, antioxidants and breast cancer risk". International Journal of Cancer. 124 (7): 1637–43. doi:10.1002/ijc.24105. PMC 2727686. PMID 19089916.
- ^ a b Mathur MB, Epel E, Kind S, Desai M, Parks CG, Sandler DP, Khazeni N (May 2016). "Perceived stress and telomere length: A systematic review, meta-analysis, and methodologic considerations for advancing the field". Brain, Behavior, and Immunity. 54: 158–169. doi:10.1016/j.bbi.2016.02.002. PMC 5590630. PMID 26853993.
- ^ López-Otín C, Blasco MA, Partridge L, Serrano M, Kroemer G (June 2013). "The hallmarks of aging". Cell. 153 (6): 1194–217. doi:10.1016/j.cell.2013.05.039. PMC 3836174. PMID 23746838.
- ^ Alberts B, Johnson A, Lewis J, Morgan D, Raff M, Roberts K, Walter P (2017). Molecular Biology of the Cell (Sixth ed.). Garland Science. pp. 1230–1233. ISBN 978-1-315-73536-8.
- ^ Eisenberg DT (17 December 2010). "An evolutionary review of human telomere biology: the thrifty telomere hypothesis and notes on potential adaptive paternal effects". American Journal of Human Biology. 23 (2): 149–67. doi:10.1002/ajhb.21127. PMID 21319244. S2CID 5540894.
- ^ Aston KI, Hunt SC, Susser E, Kimura M, Factor-Litvak P, Carrell D, Aviv A (November 2012). "Divergence of sperm and leukocyte age-dependent telomere dynamics: implications for male-driven evolution of telomere length in humans". Molecular Human Reproduction. 18 (11): 517–22. doi:10.1093/molehr/gas028. PMC 3480822. PMID 22782639.
- ^ Pepper GV, Bateson M, Nettle D (August 2018). "Telomeres as integrative markers of exposure to stress and adversity: a systematic review and meta-analysis". Royal Society Open Science. 5 (8): 180744. Bibcode:2018RSOS....580744P. doi:10.1098/rsos.180744. PMC 6124068. PMID 30225068.
- ^ Rentscher, Kelly E.; Carroll, Judith E.; Mitchell, Colter (2020). "Psychosocial Stressors and Telomere Length: A Current Review of the Science". Annual Review of Public Health. 41: 223–245. doi:10.1146/annurev-publhealth-040119-094239. PMID 31900099.
- ^ Hayflick L, Moorhead PS (December 1961). "The serial cultivation of human diploid cell strains". Experimental Cell Research. 25 (3): 585–621. doi:10.1016/0014-4827(61)90192-6. PMID 13905658.
- ^ Hayflick L (March 1965). "The limited in vitro lifetime of human diploid cell strains". Experimental Cell Research. 37 (3): 614–36. doi:10.1016/0014-4827(65)90211-9. PMID 14315085.
- ^ Feng J, Funk WD, Wang SS, Weinrich SL, Avilion AA, Chiu CP, et al. (September 1995). "The RNA component of human telomerase". Science. 269 (5228): 1236–41. Bibcode:1995Sci...269.1236F. doi:10.1126/science.7544491. PMID 7544491.
- ^ Bodnar AG, Ouellette M, Frolkis M, Holt SE, Chiu CP, Morin GB, et al. (January 1998). "Extension of life-span by introduction of telomerase into normal human cells". Science. 279 (5349): 349–52. Bibcode:1998Sci...279..349B. doi:10.1126/science.279.5349.349. PMID 9454332. S2CID 35667874.
- ^ Sample, Ian (November 28, 2010). "Harvard scientists reverse the ageing process in mice – now for humans". The Guardian. London.
{{cite news}}: Unknown parameter|name-list-format=ignored (|name-list-style=suggested) (help) - ^ Jaskelioff M, Muller FL, Paik JH, Thomas E, Jiang S, Adams AC, et al. (January 2011). "Telomerase reactivation reverses tissue degeneration in aged telomerase-deficient mice". Nature. 469 (7328): 102–6. Bibcode:2011Natur.469..102J. doi:10.1038/nature09603. PMC 3057569. PMID 21113150.
- ^ Joeng KS, Song EJ, Lee KJ, Lee J (June 2004). "Long lifespan in worms with long telomeric DNA". Nature Genetics. 36 (6): 607–11. doi:10.1038/ng1356. PMID 15122256.
- ^ Weinstein BS, Ciszek D (May 2002). "The reserve-capacity hypothesis: evolutionary origins and modern implications of the trade-off between tumor-suppression and tissue-repair". Experimental Gerontology. 37 (5): 615–27. doi:10.1016/S0531-5565(02)00012-8. PMID 11909679. S2CID 12912742.
- ^ Nakagawa S, Gemmell NJ, Burke T (September 2004). "Measuring vertebrate telomeres: applications and limitations" (PDF). Molecular Ecology. 13 (9): 2523–33. doi:10.1111/j.1365-294X.2004.02291.x. PMID 15315667.
- ^ Juola FA, Haussmann MF, Dearborn DC, Vleck CM (July 2006). "Telomere shortening in a long-lived marine bird: cross-sectional analysis and test of an aging tool". The Auk. 123 (3): 775–83. doi:10.1642/0004-8038(2006)123[775:TSIALM]2.0.CO;2. JSTOR 4090554.
- ^ Gomes NM, Ryder OA, Houck ML, Charter SJ, Walker W, Forsyth NR, et al. (October 2011). "Comparative biology of mammalian telomeres: hypotheses on ancestral states and the roles of telomeres in longevity determination". Aging Cell. 10 (5): 761–8. doi:10.1111/j.1474-9726.2011.00718.x. PMC 3387546. PMID 21518243.
- ^ Harris SE, Martin-Ruiz C, von Zglinicki T, Starr JM, Deary IJ (July 2012). "Telomere length and aging biomarkers in 70-year-olds: the Lothian Birth Cohort 1936". Neurobiology of Aging. 33 (7): 1486.e3–8. doi:10.1016/j.neurobiolaging.2010.11.013. PMID 21194798. S2CID 10309423.
- ^ Gilley D, Blackburn EH (March 1994). "Lack of telomere shortening during senescence in Paramecium". Proceedings of the National Academy of Sciences of the United States of America. 91 (5): 1955–8. Bibcode:1994PNAS...91.1955G. doi:10.1073/pnas.91.5.1955. PMC 43283. PMID 8127914.
- ^ Peška V, Fajkus P, Fojtová M, Dvořáčková M, Hapala J, Dvořáček V, et al. (May 2015). "Characterisation of an unusual telomere motif (TTTTTTAGGG)n in the plant Cestrum elegans (Solanaceae), a species with a large genome". The Plant Journal. 82 (4): 644–54. doi:10.1111/tpj.12839. PMID 25828846.
- ^ Fajkus P, Peška V, Sitová Z, Fulnečková J, Dvořáčková M, Gogela R, et al. (February 2016). "Allium telomeres unmasked: the unusual telomeric sequence (CTCGGTTATGGG)n is synthesized by telomerase". The Plant Journal. 85 (3): 337–47. doi:10.1111/tpj.13115. PMID 26716914. S2CID 206331112.
- ^ Peska, Vratislav; Mátl, Martin; Mandákova, Terezie; Vitales, Daniel; Fajkus, Petr; Fajkus, Jiří; Garcia, Sònia (2020-03-12). "Human-like telomeres in Zostera marina reveal a mode of transition from the plant to the human telomeric sequences". doi:10.1101/2020.03.11.987156. S2CID 214725911.
{{cite journal}}: Cite journal requires|journal=(help) - ^ a b Price, Lawrence H.; Kao, Hung-Teh; Burgers, Darcy E.; Carpenter, Linda L.; Tyrka, Audrey R. (2013-01-01). "Telomeres and Early-Life Stress: An Overview". Biological Psychiatry. 73 (1): 15–23. doi:10.1016/j.biopsych.2012.06.025. ISSN 0006-3223. PMC 3495091. PMID 22831981.
- ^ Raynaud CM, Sabatier L, Philipot O, Olaussen KA, Soria JC (May 2008). "Telomere length, telomeric proteins and genomic instability during the multistep carcinogenic process". Critical Reviews in Oncology/Hematology. 66 (2): 99–117. doi:10.1016/j.critrevonc.2007.11.006. PMID 18243729.
- ^ Blasco MA, Lee HW, Hande MP, Samper E, Lansdorp PM, DePinho RA, Greider CW (October 1997). "Telomere shortening and tumor formation by mouse cells lacking telomerase RNA". Cell. 91 (1): 25–34. doi:10.1016/s0092-8674(01)80006-4. PMID 9335332.
- ^ Artandi SE, Chang S, Lee SL, Alson S, Gottlieb GJ, Chin L, DePinho RA (August 2000). "Telomere dysfunction promotes non-reciprocal translocations and epithelial cancers in mice". Nature. 406 (6796): 641–5. Bibcode:2000Natur.406..641A. doi:10.1038/35020592. PMID 10949306. S2CID 4420387.
- ^ a b Armanios M (March 2013). "Telomeres and age-related disease: how telomere biology informs clinical paradigms". The Journal of Clinical Investigation. 123 (3): 996–1002. doi:10.1172/JCI66370. PMC 3673231. PMID 23454763.
- ^ Willeit P, Willeit J, Mayr A, Weger S, Oberhollenzer F, Brandstätter A, et al. (July 2010). "Telomere length and risk of incident cancer and cancer mortality". JAMA. 304 (1): 69–75. doi:10.1001/jama.2010.897. PMID 20606151.
- ^ Ma H, Zhou Z, Wei S, Liu Z, Pooley KA, Dunning AM, et al. (2011). "Shortened telomere length is associated with increased risk of cancer: a meta-analysis". PLOS ONE. 6 (6): e20466. Bibcode:2011PLoSO...620466M. doi:10.1371/journal.pone.0020466. PMC 3112149. PMID 21695195.
{{cite journal}}: CS1 maint: unflagged free DOI (link) - ^ Wentzensen IM, Mirabello L, Pfeiffer RM, Savage SA (June 2011). "The association of telomere length and cancer: a meta-analysis". Cancer Epidemiology, Biomarkers & Prevention. 20 (6): 1238–50. doi:10.1158/1055-9965.epi-11-0005. PMC 3111877. PMID 21467229.
- ^ Allshire RC, Dempster M, Hastie ND (June 1989). "Human telomeres contain at least three types of G-rich repeat distributed non-randomly". Nucleic Acids Research. 17 (12): 4611–27. doi:10.1093/nar/17.12.4611. PMC 318019. PMID 2664709.
{{cite journal}}: Unknown parameter|displayauthors=ignored (|display-authors=suggested) (help) - ^ Rufer N, Dragowska W, Thornbury G, Roosnek E, Lansdorp PM (August 1998). "Telomere length dynamics in human lymphocyte subpopulations measured by flow cytometry". Nature Biotechnology. 16 (8): 743–7. doi:10.1038/nbt0898-743. PMID 9702772. S2CID 23833545.
{{cite journal}}: Unknown parameter|displayauthors=ignored (|display-authors=suggested) (help) - ^ Cawthon RM (May 2002). "Telomere measurement by quantitative PCR". Nucleic Acids Research. 30 (10): 47e–47. doi:10.1093/nar/30.10.e47. PMC 115301. PMID 12000852.
- ^ Ding Z (2014). "Estimating telomere length from whole genome sequence data". Nucleic Acids Research. 42 (9): e75. doi:10.1093/nar/gku181. PMC 4027178. PMID 24609383.
- ^ Farmery J (2018). "Telomerecat: A ploidy-agnostic method for estimating telomere length from whole genome sequencing data". Scientific Reports. 8 (1): 1300. Bibcode:2018NatSR...8.1300F. doi:10.1038/s41598-017-14403-y. PMC 5778012. PMID 29358629.
- ^ Feuerbach L (2019). "TelomereHunter–in silico estimation of telomere content and composition from cancer genomes". BMC Bioinformatics. 20 (1): 272. doi:10.1186/s12859-019-2851-0. PMC 6540518. PMID 31138115.
{{cite journal}}: CS1 maint: unflagged free DOI (link) - ^ Baerlocher, Gabriela M.; Vulto, Irma; de Jong, Gary; Lansdorp, Peter M. (December 2006). "Flow cytometry and FISH to measure the average length of telomeres (flow FISH)". Nature Protocols. 1 (5): 2365–2376. doi:10.1038/nprot.2006.263. ISSN 1750-2799.
- ^ Pollack, Andrew (May 18, 2011). "A Blood Test Offers Clues to Longevity". The New York Times.
- ^ von Zglinicki T (March 2012). "Will your telomeres tell your future?". BMJ. 344: e1727. doi:10.1136/bmj.e1727. PMID 22415954.
- ^ Marchant J (2011). "Spit test offers guide to health". Nature. doi:10.1038/news.2011.330.
- ^ Olsson M, Wapstra E, Friesen C (March 2018). "Ectothermic telomeres: it's time they came in from the cold". Philosophical Transactions of the Royal Society of London. Series B, Biological Sciences. 373 (1741): 20160449. doi:10.1098/rstb.2016.0449. PMC 5784069. PMID 29335373.
External links
- Telomeres and Telomerase: The Means to the End Nobel Lecture by Elizabeth Blackburn, which includes a reference to the impact of stress, and pessimism on telomere length
- Telomerase and the Consequences of Telomere Dysfunction Nobel Lecture by Carol Greider
- DNA Ends: Just the Beginning Nobel Lecture by Jack Szostak

