Haplogroup R1a
| Haplogroup R1a | |
|---|---|
| File:Distribution Haplogroup R1a Y-DNA.svg | |
| Possible time of origin | Less than 18,500 YBP (Sharma 2009) |
| Possible place of origin | Eurasia (see text). |
| Ancestor | Haplogroup R1 |
| Descendants | Haplogroup R1a1 |
| Defining mutations | L62, L63, L120, M420, M449, M511, M513 |
| Highest frequencies | See List of R1a frequency by population |
Haplogroup R1a, or haplogroup R-M420, is a common Y DNA haplogroup in many parts of Eurasia. One sub-clade (branch) of R1a, haplogroup R1a1, is much more common than the others in all major geographical regions. R1a1, defined by the single-nucleotide polymorphism (SNP) mutation M17, (and sometimes alternatively defined as R-M198), is particularly common in a large region extending from South Asia and southern Siberia to Central Europe and Scandinavia (Underhill 2009).
The R1a family is defined most broadly by the SNP mutation M420, which was discovered after M17. The discovery of M420 resulted in a reorganization of the lineage in particular establishing a new paragroup (designated R-M420*) for the relatively rare lineages which are not in the R-SRY10831.2 (R1a1) branch leading to R-M17.
A large, 2014 study by Underhill et al., using 16,244 individuals from over 126 populations from across Eurasia, concluded there was compelling evidence, that R1a-M420 originated in Iran.[1] Another large, 2012 study on ethnic groups of Afghanistan, using 204 Afghan samples along with at least 8,500 samples from surrounding populations, have found the Mid-Holocene R1a1a7 M-458 sublineage to be absent there which does not support the view of R1a1a's relation with the intrusional Indo-European languages to Central Asia and South Asia, as previously thought.[2]
The data on DNA-archeology
Haplogroup R1a was found in the remains of the Corded Ware culture[3][4] and Urnfield culture;[5] as well as the burial of the remains of the Andronovo culture,[6] the Pazyryk culture,[7] Tagar culture[8] and Tashtyk culture,[8] the inhabitants of ancient Tanais,[9] in the Tarim mummies,[10] the aristocracy Xiongnu.[11]
Phylogeny
The R1a family tree now has three major levels of branching, with the largest number of defined subclades within the dominant and best known branch, R1a1a (which will be found with various names; in particular, as "R1a1" in relatively recent but not the latest literature.)
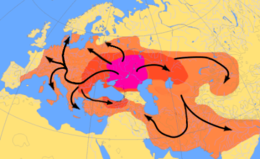
Roots of R1a
|
R1a, distinguished by several unique markers including the M420 mutation, is a subclade of Haplogroup R-M173 (previously called R1), which is defined by SNP mutation M173. Besides R-M420, R-M173 also has the subclades Haplogroup R1b, defined by the M343 mutation, and the paragroup R-M173*.
R-M420 (R1a)
R-M420, defined by the mutation M420, has two branches: R-SRY1532.2, defined by the mutation SRY1532.2, which makes up the vast majority; and R-M420*, the paragroup, defined as M420 positive but SRY1532.2 negative. (In the 2002 scheme, this SRY1532.2 negative minority was one part of the relatively rare group classified as the paragroup R1*.) Mutations understood to be equivalent to M420 include M449, M511, M513, L62, and L63.[12][13]
Only isolated samples of the new paragroup R-M420* were found by Underhill 2009, mostly in the Middle East and Caucasus: 1/121 Omanis, 2/150 Iranians, 1/164 in the United Arab Emirates, and 3/612 in Turkey. Testing of 7224 more males in 73 other Eurasian populations showed no sign of this category.[14]
R-SRY1532.2 (R1a1)
R-SRY1532.2 is defined by SRY1532.2, also referred to as SRY10831.2. SNP mutations understood to be always occurring with SRY1532.2 include SRY10831.2, M448, L122, M459, and M516.[12][15] This family of lineages is dominated by the R-M17 branch, which is positive for M17 and M198. The paragroup R-SRY1532.2* is positive for the SRY1532.2 marker, but lacks either the M17 or M198 markers.
The R-SRY1532.2* paragroup is apparently less rare than R1*, but still relatively unusual, though it has been tested in more than one survey. Underhill[14] reported 1/51 in Norway, 3/305 in Sweden, 1/57 Greek Macedonians, 1/150 Iranians, 2/734 ethnic Armenians, and 1/141 Kabardians. While Sahoo[16] reported R-SRY1532.2* for 1/15 Himachal Pradesh Rajput samples.
R-M17/M198 (R1a1a)
R-M17 makes up the vast majority of all R-M420 over its entire geographic range. It is defined by SNP mutations M17 or M198, which have always appeared together in the same men so far. SNP mutations understood to be always occurring with M17 and M198 include M417, M512, M514, M515.[17] R-M17 has many subclades of its own defined by mutations. Two important subclades appear to broadly divide the European and Asian parts of this large clade:
R-Z283 (R1a1a1b1)
This large subclade appears to encompass most of the R1a1a found in Europe.[18]
R-Z93 (R1a1a1b2)
This large subclade appears to encompass most of the R1a1a found in Asia.[18]
Popular science
Bryan Sykes in his book Blood of the Isles gives imaginative names to the founders or "clan patriarchs" of major British Y haplogroups, much as he did for mitochondrial haplogroups in his work The Seven Daughters of Eve. He named R1a1a in Europe the "clan" of a "patriarch" Sigurd, reflecting the theory that R1a1a in the British Isles has Norse origins.
Historic meanings of "R1a"
The historic naming system commonly used for R1a was inconsistent in different published sources, because it changed often, this requires some explanation.
In 2002, the Y chromosome consortium (YCC) proposed a new naming system for haplogroups, which has now become standard.(YCC 2002) In this system, names with the format "R1" and "R1a" are "phylogenetic" names, aimed at marking positions in a family tree. Names of SNP mutations can also be used to name clades or haplogroups. For example, as M173 is currently the defining mutation of R1, R1 is also R-M173, a "mutational" clade name. When a new branching in a tree is discovered, some phylogenetic names will change, but by definition all mutational names will remain the same.
The widely occurring haplogroup defined by mutation M17 was known by various names, such as "Eu19", as used in (Semino 2000) in the older naming systems. The 2002 YCC proposal assigned the name R1a to the haplogroup defined by mutation SRY1532.2. This included Eu19 (i.e. R-M17) as a subclade, so Eu19 was named R1a1. Note, SRY1532.2 is also known as SRY10831.2[citation needed] The discovery of M420 in 2009 has caused a reassignment of these phylogenetic names.(Underhill 2009 and ISOGG 2012) R1a is now defined by the M420 mutation: in this updated tree, the subclade defined by SRY1532.2 has moved from R1a to R1a1, and Eu19 (R-M17) from R1a1 to R1a1a.
More recent updates recorded at the ISOGG reference webpage involve branches of R-M17, including one major branch, R-M417.
| 2002 Scheme proposed in (YCC 2002) | 2009 Scheme as per (2009) | Latest ISOGG tree as per January 2011 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
|
|
|
See also
Y-DNA R-M207 subclades
Y-DNA backbone tree
In art
Artem Lukichev created an animation based on the Bashkir epic about the Ural, which outlined the history of the clusters of haplogroup R1: R1a and R1b.[19]
References
- "Y-DNA Haplogroup R and its Subclades". International Society of Genetic Genealogy (ISOGG). Retrieved 8 January 2011.
- Krahn, Thomas; FTDNA; Genetic Genealogy Community. "Family Tree DNA Draft Y-Chromosome Tree".
- Pamjav, Horolma; Fehér, Tibor; Németh, Endre; Pádár, Zsolt (2012). "Brief communication: new Y-chromosome binary markers improve phylogenetic resolution within haplogroup R1a1". American Journal of Physical Anthropology. 149 (4): 611–615. doi:10.1002/ajpa.22167. PMID 23115110.
- Sahoo, S; Singh, A; Himabindu, G; Banerjee, J; Sitalaximi, T; Gaikwad, S; Trivedi, R; Endicott, P; et al. (2006). "A prehistory of Indian Y chromosomes: Evaluating demic diffusion scenarios". Proceedings of the National Academy of Sciences. 103 (4): 843–848. Bibcode:2006PNAS..103..843S. doi:10.1073/pnas.0507714103. PMC 1347984. PMID 16415161.
- Sharma, S; Rai, E; Sharma, P; Jena, M; Singh, S; Darvishi, K; Bhat, AK; Bhanwer, AJ; et al. (2009). "The Indian origin of paternal haplogroup R1a1(*)substantiates the autochthonous origin of Brahmins and the caste system". Journal of Human Genetics. 54 (1): 47–55. doi:10.1038/jhg.2008.2. PMID 19158816.
- Semino, O.; Passarino, G; Oefner, PJ; Lin, AA; Arbuzova, S; Beckman, LE; De Benedictis, G; Francalacci, P; et al. (2000). "The Genetic Legacy of Paleolithic Homo sapiens sapiens in Extant Europeans: A Y Chromosome Perspective" (PDF). Science. 290 (5494): 1155–59. Bibcode:2000Sci...290.1155S. doi:10.1126/science.290.5494.1155. PMID 11073453.. Copy can be found at http://www.historyofmacedonia.org/ConciseMacedonia/Y_Hromosomes.pdf.
- Underhill, Peter A; Myres, Natalie M; Rootsi, Siiri; Metspalu, Mait; Zhivotovsky, Lev A; King, Roy J; Lin, Alice A; Chow, Cheryl-Emiliane T; et al. (2009). "Separating the post-Glacial coancestry of European and Asian Y chromosomes within haplogroup R1a". European Journal of Human Genetics. 18 (4): 479–84. doi:10.1038/ejhg.2009.194. PMC 2987245. PMID 19888303.
- Y Chromosome Consortium "YCC" (2002). "A Nomenclature System for the Tree of Human Y-Chromosomal Binary Haplogroups". Genome Research. 12 (2): 339–348. doi:10.1101/gr.217602. PMC 155271. PMID 11827954.
- ^ http://www.nature.com/ejhg/journal/v23/n1/pdf/ejhg201450a.pdf
- ^ http://www.plosone.org/article/info%3Adoi%2F10.1371%2Fjournal.pone.0034288
- ^ Haak W. Ancient DNA, Strontium isotopes, and osteological analyses shed light on social and kinship organization of the Later Stone Age//Stanford University, Stanford, CA, and approved October 3, 2008 (received for review August 5, 2008)
- ^ Brandit, G (2013). "Ancient DNA Reveals Key Stages in the Formation of Central European Mitochondrial Genetic Diversity". Science. 342 (6155): 257–261. doi:10.1126/science.1241844.
- ^ Schweitzer D. Lichtenstein Cave Data Analysis, 2008.
- ^ [Keyser C., etc. Ancient DNA provides new insights into the history of south Siberian Kurgan people//Hum Genet (2009) 126:395-410DOI 10.1007/s00439-009-0683-0]
- ^ Ricaut, F.; et al. (2004). "Genetic Analysis of a Scytho-Siberian Skeleton and Its Implications for Ancient Central Asian Migrations". Human Biology. 76: 1.
{{cite journal}}: Explicit use of et al. in:|last2=(help) - ^ a b Keyser C. etc. Ancient DNA provides new insights into the history of south Siberian Kurgan people//Hum Genet (2009) 126:395-410DOI 10.1007/s00439-009-0683-0
- ^ Корниенко И. В., Водолажский Д. И. Использование нерекомбинантных маркеров Y-хромосомы в исследованиях древних популяций (на примере поселения Танаис)//Материалы Донских антропологических чтений. Ростов-на-Дону, Ростовский научно-исследовательский онкологический институт, Ростов-на-Дону, 2013.
- ^ Chunxiang Li, etc. Evidence that a West-East admixed population lived in the Tarim Basin as early as the early Bronze Age
- ^ Kim, K.; et al. (Jul 2010). "A western Eurasian male is found in 2000-year-old elite Xiongnu cemetery in Northeast Mongolia". Am J Phys Anthropol. 142 (3): 429–40. doi:10.1002/ajpa.21242. PMID 20091844.
{{cite journal}}: Explicit use of et al. in:|last2=(help) - ^ a b Underhill 2009
- ^ ISOGG 2012)
- ^ a b (Underhill 2009)
- ^ Krahn 2012)
- ^ (Sahoo 2006)
- ^ (Underhill 2009).
- ^ a b (Pamjav 2012).
- ^ About R1a and R1b from Ural epic story. Artem Lukichev (c)
Further reading
- Adams, Susan M.; Bosch, E; Balaresque, PL; Ballereau, SJ; Lee, AC; Arroyo, E; López-Parra, AM; Aler, M; et al. (2008). "The Genetic Legacy of Religious Diversity and Intolerance: Paternal Lineages of Christians, Jews, and Muslims in the Iberian Peninsula". American Journal of Human Genetics. 83 (6): 725–36. doi:10.1016/j.ajhg.2008.11.007. PMC 2668061. PMID 19061982.
- Al Zahery, N.; Semino, O.; Benuzzi, G.; Magri, C.; Passarino, G.; Torroni, A.; Santachiara-Benerecetti, A.S. (2003). "Y-chromosome and mtDNA polymorphisms in Iraq, a crossroad of the early human dispersal and of post-Neolithic migrations" (PDF). Molecular Phylogenetics and Evolution. 28 (3): 458–72. doi:10.1016/S1055-7903(03)00039-3. PMID 12927131.
- Balanovsky, O; Rootsi, S; Pshenichnov, A; Kivisild, T; Churnosov, M; Evseeva, I; Pocheshkhova, E; Boldyreva, M; et al. (2008). "Two Sources of the Russian Patrilineal Heritage in Their Eurasian Context". AJHG. 82 (1): 236–250. doi:10.1016/j.ajhg.2007.09.019. PMC 2253976. PMID 18179905.
{{cite journal}}: Invalid|ref=harv(help) - Bamshad, M.; Kivisild, T; Watkins, WS; Dixon, ME; Ricker, CE; Rao, BB; Naidu, JM; Prasad, BV; et al. (2001). "Genetic evidence on the origins of Indian caste populations". Genome Research. 11 (6): 994–1004. doi:10.1101/gr.GR-1733RR. PMC 311057. PMID 11381027.
{{cite journal}}: Invalid|ref=harv(help). - Barać, Lovorka; Pericić, Marijana; Klarić, Irena Martinović; Rootsi, Siiri; Janićijević, Branka; Kivisild, Toomas; Parik, Jüri; Rudan, Igor; et al. (July 2003). "Y chromosomal heritage of Croatian population and its island isolates" (PDF). European Journal of Human Genetics. 11 (7): 535–42. doi:10.1038/sj.ejhg.5200992. PMID 12825075.
{{cite journal}}: Invalid|ref=harv(help)CS1 maint: postscript (link) - Battaglia, Vincenza; Fornarino, S; Al-Zahery, N; Olivieri, A; Pala, M; Myres, NM; King, RJ; Rootsi, S; et al. (2008). "Y-chromosomal evidence of the cultural diffusion of agriculture in southeast Europe". European Journal of Human Genetics. 17 (6): 820–30. doi:10.1038/ejhg.2008.249. PMC 2947100. PMID 19107149.
- Behar, D; Thomas, MG; Skorecki, K; Hammer, MF; Bulygina, E; Rosengarten, D; Jones, AL; Held, K; et al. (2003). "Multiple Origins of Ashkenazi Levites: Y Chromosome Evidence for Both Near Eastern and European Ancestries" (– Scholar search). American Journal of Human Genetics. 73 (4): 768–779. doi:10.1086/378506. PMC 1180600. PMID 13680527.
{{cite journal}}: External link in|format=|ref=harv(help) [dead link]. Also at http://www.ucl.ac.uk/tcga/tcgapdf/Behar-AJHG-03.pdf and http://www.familytreedna.com/pdf/400971.pdf - Bouakaze, C.; Keyser, C; Amory, S; Crubézy, E; Ludes, B (2007). "First successful assay of Y-SNP typing by SNaPshot minisequencing on ancient DNA". International Journal of Legal Medicine. 121 (6): 493–9. doi:10.1007/s00414-007-0177-3. PMID 17534642.
- Bowden, G. R.; Balaresque, P; King, TE; Hansen, Z; Lee, AC; Pergl-Wilson, G; Hurley, E; Roberts, SJ; et al. (2008). "Excavating Past Population Structures by Surname-Based Sampling: The Genetic Legacy of the Vikings in Northwest England". Molecular Biology and Evolution. 25 (2): 301–309. doi:10.1093/molbev/msm255. PMC 2628767. PMID 18032405.
- Braya, Steven; Mullea, Jennifer; Dodda, Anne; Pulver, Ann; Wooding, Stephen; Warren, Stephen (2010). "Signatures of founder effects, admixture, and selection in the Ashkenazi Jewish population". PNAS. 107 (37): 16222–16227. Bibcode:2010PNAS..10716222B. doi:10.1073/pnas.1004381107. PMC 2941333. PMID 20798349.
- Capelli, C; Redhead, N; Abernethy, JK; Gratrix, F; Wilson, JF; Moen, T; Hervig, T; Richards, M; et al. (2003). "A Y Chromosome Census of the British Isles". Current Biology. 13 (11): 979–84. doi:10.1016/S0960-9822(03)00373-7. PMID 12781138.
{{cite journal}}: Invalid|ref=harv(help) also at "University College London" (PDF). - Cinnioğlu, C; King, R; Kivisild, T; Kalfoğlu, E; Atasoy, S; Cavalleri, GL; Lillie, AS; Roseman, CC; et al. (2004). "Excavating Y-chromosome haplotype strata in Anatolia" (PDF). Hum Genet. 114 (2): 127–48. doi:10.1007/s00439-003-1031-4. PMID 14586639.
{{cite journal}}: Invalid|ref=harv(help) - Cordaux, Richard; Aunger, R; Bentley, G; Nasidze, I; Sirajuddin, SM; Stoneking, M (2004). "Independent Origins of Indian Caste and Tribal Paternal Lineages". Current Biology. 14 (3): 231–235. doi:10.1016/j.cub.2004.01.024. PMID 14761656.
- Dupuy, Berit Myhre; Stenersen, M; Lu, TT; Olaisen, B (2005). "Geographical heterogeneity of Y-chromosomal lineages in Norway" (PDF). Forensic Science International. 164 (1): 10–19. doi:10.1016/j.forsciint.2005.11.009. PMID 16337760.
- Firasat, Sadaf; Khaliq, S; Mohyuddin, A; Papaioannou, M; Tyler-Smith, C; Underhill, PA; Ayub, Q (2006). "Y-chromosomal evidence for a limited Greek contribution to the Pathan population of Pakistan". European Journal of Human Genetics. 15 (1): 121–126. doi:10.1038/sj.ejhg.5201726. PMC 2588664. PMID 17047675.
{{cite journal}}: Invalid|ref=harv(help) - Flores, Carlos; Maca-Meyer, N; Larruga, JM; Cabrera, VM; Karadsheh, N; Gonzalez, AM (2005). "Isolates in a corridor of migrations: a high-resolution analysis of Y-chromosome variation in Jordan". Journal of Human Genetics. 50 (9): 435–441. doi:10.1007/s10038-005-0274-4. PMID 16142507.
{{cite journal}}: Invalid|ref=harv(help) - Fornarino, Simona; Pala, Maria; Battaglia, Vincenza; Maranta, Ramona; Achilli, Alessandro; Modiano, Guido; Torroni, Antonio; Semino, Ornella; Santachiara-Benerecetti, Silvana A (2009). "Mitochondrial and Y-chromosome diversity of the Tharus (Nepal): a reservoir of genetic variation". BMC Evolutionary Biology. 9: 154. doi:10.1186/1471-2148-9-154. PMC 2720951. PMID 19573232.
{{cite journal}}: CS1 maint: unflagged free DOI (link) - Gimbutas (1970). Indo-European and Indo-Europeans. Univ. of Pennsylvania Press, Philadelphia, PA. pp. 155–195.
- Gwozdz (2009). "Y-STR Mountains in Haplospace, Part II: Application to Common Polish Clades" (PDF). Journal of Genetic Genealogy. 5 (2).
- Haak, W.; Brandt, G.; Jong, H. N. d.; Meyer, C.; Ganslmeier, R.; Heyd, V.; Hawkesworth, C.; Pike, A. W. G.; et al. (2008). "Ancient DNA, Strontium isotopes, and osteological analyses shed light on social and kinship organization of the Later Stone Age". Proceedings of the National Academy of Sciences. 105 (47): 18226–18231. Bibcode:2008PNAS..10518226H. doi:10.1073/pnas.0807592105. PMC 2587582. PMID 19015520.
- Hammer, Michael F.; Behar, Doron M.; Karafet, Tatiana M.; Mendez, Fernando L.; Hallmark, Brian; Erez, Tamar; Zhivotovsky, Lev A.; Rosset, Saharon; Skorecki, Karl (2009). "Response" (PDF). Human Genetics. 126 (5): 725–726. doi:10.1007/s00439-009-0747-1.
- Helgason, A; Sigureardottir, S; Nicholson, J; Sykes, B; Hill, E; Bradley, D; Bosnes, V; Gulcher, J; et al. (2000). "Estimating Scandinavian and Gaelic Ancestry in the Male Settlers of Iceland". American Journal of Human Genetics. 67 (3): 697–717. doi:10.1086/303046. PMC 1287529. PMID 10931763.
- Karafet, TM; Mendez, FL; Meilerman, MB; Underhill, PA; Zegura, SL; Hammer, MF (May 2008). Abstract "New Binary Polymorphisms Reshape and Increase Resolution of the Human Y-Chromosomal Haplogroup Tree". Genome Research. 18 (5): 830–8. doi:10.1101/gr.7172008. PMC 2336805. PMID 18385274.
{{cite journal}}: Check|url=value (help); Invalid|ref=harv(help). Published online April 2, 2008. See also Supplementary Material. - Kasperaviciūte, D.; Kucinskas, V.; Stoneking, M. (2005). "Y Chromosome and Mitochondrial DNA Variation in Lithuanians". Annals of Human Genetics. 68 (5): 438–452. doi:10.1046/j.1529-8817.2003.00119.x. PMID 15469421.
- Kayser, M; Lao, O; Anslinger, K; Augustin, C; Bargel, G; Edelmann, J; Elias, S; Heinrich, M; et al. (2005). "Significant genetic differentiation between Poland and Germany follows present-day political borders, as revealed by Y-chromosome analysis". Human Genetics. 117 (5): 428–443. doi:10.1007/s00439-005-1333-9. PMID 15959808.
{{cite journal}}: Invalid|ref=harv(help) A copy can be found here [1]. - Keyser; Bouakaze, Caroline; Crubézy, Eric; Nikolaev, Valery G.; Montagnon, Daniel; Reis, Tatiana; Ludes, Bertrand (2009). "Ancient DNA provides new insights into the history of south Siberian Kurgan people". Human Genetics. 126 (3): 395–410. doi:10.1007/s00439-009-0683-0. PMID 19449030.
{{cite journal}}: Unknown parameter|displayauthors=ignored (|display-authors=suggested) (help) - Kharkov, V. N.; Stepanov, V. A.; Borinskaya, S. A.; Kozhekbaeva, Zh. M.; Gusar, V. A.; Grechanina, E. Ya.; Puzyrev, V. P.; Khusnutdinova, E. K.; Yankovsky, N. K. (2004). "Gene Pool Structure of Eastern Ukrainians as Inferred from the Y-Chromosome Haplogroups". Russian Journal of Genetics. 40 (3): 326–331. doi:10.1023/B:RUGE.0000021635.80528.2f.
{{cite journal}}: Invalid|ref=harv(help) A copy can be found here [2]. - Kharkov, V. N.; Stepanov, V. A.; Feshchenko, S. P.; Borinskaya, S. A.; Yankovsky, N. K.; Puzyrev, V. P. (2005). "Frequencies of Y Chromosome Binary Haplogroups in Belarusians". Russian Journal of Genetics. 41 (8): 928–931. doi:10.1007/s11177-005-0182-x.
{{cite journal}}: Invalid|ref=harv(help) A copy can be found here [3]. - Kharkov, V. N.; Stepanov, V. A.; Medvedeva, O. F.; Spiridonova, M. G.; Voevoda, M. I.; Tadinova, V. N.; Puzyrev, V. P. (2007). "Gene Pool Differences between Northern and Southern Altaians Inferred from the Data on Y-Chromosomal Haplogroups" (PDF). Russian Journal of Genetics. 43 (5): 551–562. doi:10.1134/S1022795407050110.
- King, RJ; Ozcan, SS; Carter, T; Kalfoğlu, E; Atasoy, S; Triantaphyllidis, C; Kouvatsi, A; Lin, AA; et al. (2008). "Differential Y-chromosome Anatolian Influences on the Greek and Cretan Neolithic" (PDF). Annals of Human Genetics. 72 (Pt 2): 205–214. doi:10.1111/j.1469-1809.2007.00414.x. PMID 18269686.
{{cite journal}}: Invalid|ref=harv(help) - Kivisild, T; Rootsi, S; Metspalu, M; Mastana, S; Kaldma, K; Parik, J; Metspalu, E; Adojaan, M; et al. (2003). "The Genetic Heritage of the Earliest Settlers Persists Both in Indian Tribal and Caste Populations". AJHG. 72 (2): 313–32. doi:10.1086/346068. PMC 379225. PMID 12536373.
{{cite journal}}: Invalid|ref=harv(help). - Lalueza-Fox, C.; Robello, M; Mao, C; Mainardi, P; Besio, G; Pettener, D.; Bertranpetit, J. (2004). "Unravelling migrations in the steppe: mitochondrial DNA sequences from ancient central Asians". Proc Biol Sci. 271 (1542): 941–947. doi:10.1098/rspb.2004.2698. PMC 1691686. PMID 15255049.
- Lell, JT; Sukernik, RI; Starikovskaya, YB; Su, B; Jin, L; Schurr, TG; Underhill, PA; Wallace, DC (2002). "The Dual Origin and Siberian Affinities of Native American Y Chromosomes" (PDF). American Journal of Human Genetics. 70 (1): 192–206. doi:10.1086/338457. PMC 384887. PMID 11731934.
{{cite journal}}: Invalid|ref=harv(help) - Luca, F; Di Giacomo, F; Benincasa, T; Popa, LO; Banyko, J; Kracmarova, A; Malaspina, P; Novelletto, A; Brdicka, R (2006). "Y-Chromosomal Variation in the Czech Republic". American Journal of Physical Anthropology. 132 (1): 132–9. doi:10.1002/ajpa.20500. PMID 17078035.
{{cite journal}}: Invalid|ref=harv(help) - Malaspina (2003). "Analysis of Y-chromosome variation in modern populations at the European-Asian border" (PDF): 309–313.
{{cite journal}}: Cite journal requires|journal=(help) in K. Boyle, C. Renfrew, and M. Levine, eds. Ancient interactions: east and west in Eurasia. McDonald Institute for Archaeological Research Monograph Series, Cambridge University Press, Cambridge - Marjanovic, D; Fornarino, S; Montagna, S; Primorac, D.; Hadziselimovic, R.; Vidovic, S.; Pojskic, N.; Battaglia, V.; et al. (November 2005). "The peopling of modern Bosnia-Herzegovina: Y-chromosome haplogroups in the three main ethnic groups". Annals of Human Genetics. 69 (Pt 6): 757–63. doi:10.1111/j.1529-8817.2005.00190.x. PMID 16266413.
{{cite journal}}: Invalid|ref=harv(help)CS1 maint: postscript (link) - Mirabal, Sheyla; Regueiro, M; Cadenas, AM; Cavalli-Sforza, LL; Underhill, PA; Verbenko, DA; Limborska, SA; Herrera, RJ (2009). "Y-Chromosome distribution within the geo-linguistic landscape of northwestern Russia". European Journal of Human Genetics. 17 (10): 1260–1273. doi:10.1038/ejhg.2009.6. PMC 2986641. PMID 19259129.
- Mukherjee, Namita; Nebel, Almut; Oppenheim, Ariella; Majumder, Partha P. (2001). "High-resolution analysis of Y-chromosomal polymorphisms reveals signatures of population movements from central Asia and West Asia into India". Journal of Genetics. 80 (3) (published December 2001): 125–135. doi:10.1007/BF02717908. PMID 11988631.
{{cite journal}}: Invalid|ref=harv(help). - Nasidze, I; Ling, EY; Quinque, D; Dupanloup, I; Cordaux, R; Rychkov, S; Naumova, O; Zhukova, O; et al. (2004). "Mitochondrial DNA and Y-Chromosome Variation in the Caucasus" (PDF). Annals of Human Genetics. 68 (Pt 3): 205–221. doi:10.1046/j.1529-8817.2004.00092.x. PMID 15180701.
{{cite journal}}: Invalid|ref=harv(help) - Nasidze, Ivan; Quinque, D; Ozturk, M; Bendukidze, N; Stoneking, M (2005). "MtDNA and Y-chromosome Variation in Kurdish Groups" (PDF). Annals of Human Genetics. 69 (Pt 4): 401–412. doi:10.1046/j.1529-8817.2005.00174.x. PMID 15996169.
- Nebel, Almut; Filon, Dvora; Brinkmann, Bernd; Majumder, Partha; Faerman, Marina; Oppenheim, Ariella (2001). "The Y Chromosome Pool of Jews as Part of the Genetic Landscape of the Middle East". American Journal of Human Genetics. 69 (5): 1095–112. doi:10.1086/324070. PMC 1274378. PMID 11573163.
- Passarino, G; Semino, Ornella; Magria, Chiara; Al-Zahery, Nadia; Benuzzi, Giorgia; Quintana-Murci, Lluis; Andellnovic, Slmun; Bullc-Jakus, Floriana; et al. (2001). "The 49a,f haplotype 11 is a new marker of the EU19 lineage that traces migrations from northern regions of the black sea". Hum. Immunol. 62 (9): 922–932. doi:10.1016/S0198-8859(01)00291-9. PMID 11543894.
{{cite journal}}: Invalid|ref=harv(help) - Passarino, Giuseppe; Cavalleri, GL; Lin, AA; Cavalli-Sforza, LL; Børresen-Dale, AL; Underhill, PA (2002). "Different genetic components in the Norwegian population revealed by the analysis of mtDNA and Y chromosome polymorphisms". European Journal of Human Genetics. 10 (9): 521–9. doi:10.1038/sj.ejhg.5200834. PMID 12173029.
{{cite journal}}: Invalid|ref=harv(help). - Pawlowski, R; Dettlaff-Kakol, A; MacIejewska, A; Paszkowska, R; Reichert, M; Jezierski, G (2002). "Population genetics of 9 Y-chromosome STR loci w Northern Poland". Arch. Med. Sadowej Kryminol. 52 (4): 261–277. PMID 14669672.
- Pericić, M.; Lauc, LB; Klarić, IM; Rootsi, S; Janićijević, B; Rudan, I; Terzić, R; Colak, I; et al. (2005). "High-resolution phylogenetic analysis of southeastern Europe traces major episodes of paternal gene flow among Slavic populations". Mol. Biol. Evol. 22 (10): 1964–75. doi:10.1093/molbev/msi185. PMID 15944443.
{{cite journal}}: Invalid|ref=harv(help). - Qamar, R; Ayub, Q; Mohyuddin, A; Helgason, A; Mazhar, K; Mansoor, A; Zerjal, T; Tylersmith, C; Mehdi, S (2002). "Y-Chromosomal DNA Variation in Pakistan". American Journal of Human Genetics. 70 (5): 1107–24. doi:10.1086/339929. PMC 447589. PMID 11898125.
- Quintana-Murci, L; Krausz, C; Zerjal, T; Sayar, SH; Hammer, MF; Mehdi, SQ; Ayub, Q; Qamar, R; et al. (2001). "Y-chromosome lineages trace diffusion of people and languages in southwestern Asia". American Journal of Human Genetics. 68 (2): 537–542. doi:10.1086/318200. PMC 1235289. PMID 11133362.
- Rebala, Krzysztof; Mikulich, AI; Tsybovsky, IS; Siváková, D; Džupinková, Z; Szczerkowska-Dobosz, A; Szczerkowska, Z (2007). "Y-STR variation among Slavs: evidence for the Slavic homeland in the middle Dnieper basin". Journal of Human Genetics. 52 (5): 406–414. doi:10.1007/s10038-007-0125-6. PMID 17364156.
- Regueiro, M; Cadenas, AM; Gayden, T; Underhill, PA; Herrera, RJ (2006). "Iran: Tricontinental Nexus for Y-Chromosome Driven Migration" (PDF). Hum Hered. 61 (3): 132–143. doi:10.1159/000093774. PMID 16770078.
{{cite journal}}: Invalid|ref=harv(help) - Rosser, ZH; Zerjal, T; Hurles, ME; Adojaan, M; Alavantic, D; Amorim, A; Amos, W; Armenteros, M; et al. (2000). "Y-Chromosomal Diversity in Europe Is Clinal and Influenced Primarily by Geography, Rather than by Language". American Journal of Human Genetics. 67 (6): 1526–1543. doi:10.1086/316890. PMC 1287948. PMID 11078479.
{{cite journal}}: Invalid|ref=harv(help) - Saha, Anjana; Sharma, S; Bhat, A; Pandit, A; Bamezai, R (2005). "Genetic affinity among five different population groups in India reflecting a Y-chromosome gene flow". Journal of Human Genetics. 50 (1): 49–51. doi:10.1007/s10038-004-0219-3. PMID 15611834.
{{cite journal}}: Invalid|ref=harv(help). - Sanchez, J; Børsting, C; Hallenberg, C; Buchard, A; Hernandez, A; Morling, N (2003). "Multiplex PCR and minisequencing of SNPs—a model with 35 Y chromosome SNPs". Forensic Sci Int. 137 (1): 74–84. doi:10.1016/S0379-0738(03)00299-8. PMID 14550618.
- Scozzari, R; Cruciani, F; Pangrazio, A; Santolamazza, P; Vona, G; Moral, P; Latini, V; Varesi, L; et al. (2001). "Human Y-Chromosome Variation in the Western Mediterranean Area: Implications for the Peopling of the Region" (PDF). Human Immunology. 62 (9): 871–84. doi:10.1016/S0198-8859(01)00286-5. PMID 11543889.
- Sengupta, S; Zhivotovsky, LA; King, R; Mehdi, SQ; Edmonds, CA; Chow, CE; Lin, AA; Mitra, M; et al. (2005). "Polarity and Temporality of High-Resolution Y-Chromosome Distributions in India Identify Both Indigenous and Exogenous Expansions and Reveal Minor Genetic Influence of Central Asian Pastoralists". American Journal of Human Genetics. 78 (2): 202–21. doi:10.1086/499411. PMC 1380230. PMID 16400607.
{{cite journal}}: Invalid|ref=harv(help). - Sharma (2007). "The Autochthonous Origin and a Tribal Link of Indian Brahmins: Evaluation Through Molecular Genetic Markers" (PDF). THE AMERICAN SOCIETY OF HUMAN GENETICS 57th Annual Meeting.
{{cite journal}}: Invalid|ref=harv(help) - Shilz (2006). "Molekulargenetische Verwandtschaftsanalysen am prähistorischen Skelettkollektiv der Lichtensteinhöhle, Dissertation, Göttingen" (PDF).
{{cite journal}}: Cite journal requires|journal=(help) - Soares, Pedro; Achilli, Alessandro; Semino, Ornella; Davies, William; MacAulay, Vincent; Bandelt, Hans-JüRgen; Torroni, Antonio; Richards, Martin B. (2010). "The Archaeogenetics of Europe" (PDF). Current Biology. 20 (4): R174–83. doi:10.1016/j.cub.2009.11.054. PMID 20178764.
- Tambets, K; Rootsi, S; Kivisild, T; Help, H; Serk, P; Loogväli, EL; Tolk, HV; Reidla, M; et al. (2004). "The Western and Eastern Roots of the Saami—the Story of Genetic 'Outliers' Told by Mitochondrial DNA and Y Chromosomes". American Journal of Human Genetics. 74 (4): 661–682. doi:10.1086/383203. PMC 1181943. PMID 15024688.
{{cite journal}}: Invalid|ref=harv(help) - Thangaraj, Kumarasamy; Naidu, B. Prathap; Crivellaro, Federica; Tamang, Rakesh; Upadhyay, Shashank; Sharma, Varun Kumar; Reddy, Alla G.; Walimbe, S. R.; et al. (2010). Cordaux, Richard (ed.). "The Influence of Natural Barriers in Shaping the Genetic Structure of Maharashtra Populations". PLoS ONE. 5 (12): e15283. Bibcode:2010PLoSO...515283T. doi:10.1371/journal.pone.0015283. PMC 3004917. PMID 21187967.
{{cite journal}}: CS1 maint: unflagged free DOI (link) - Thanseem, Ismail; Thangaraj, Kumarasamy; Chaubey, Gyaneshwer; Singh, Vijay Kumar; Bhaskar, Lakkakula VKS; Reddy, B Mohan; Reddy, Alla G; Singh, Lalji (2006). "Genetic affinities among the lower castes and tribal groups of India: inference from Y chromosome and mitochondrial DNA". BMC Genetics. 7: 42. doi:10.1186/1471-2156-7-42. PMC 1569435. PMID 16893451.
{{cite journal}}: CS1 maint: unflagged free DOI (link) - Varzari, Alexander (2006). "Population History of the Dniester-Carpathians: Evidence from Alu Insertion and Y-Chromosome Polymorphisms" (PDF). Dissertation der Fakultät für Biologie der Ludwig-Maximilians-Universität München.
{{cite journal}}: Invalid|ref=harv(help) - Völgyi, Antónia; Zalán, Andrea; Szvetnik, Enikő; Pamjav, Horolma (2008). "Hungarian population data for 11 Y-STR and 49 Y-SNP markers". Forensic Science International: Genetics. 3 (2): e27–8. doi:10.1016/j.fsigen.2008.04.006. PMID 19215861.
- Wang, Wei; Wise, Cheryl; Baric, Tom; Black, Michael L.; Bittles, Alan H. (2003). "The origins and genetic structure of three co-resident Chinese Muslim populations: The Salar, Bo'an and Dongxiang". Human Genetics. 113 (3): 244–52. doi:10.1007/s00439-003-0948-y. PMID 12759817.
{{cite journal}}: Invalid|ref=harv(help) - Weale, Michael; Yepiskoposyan, L; Jager, RF; Hovhannisyan, N; Khudoyan, A; Burbage-Hall, O; Bradman, N; Thomas, MG (2001). "Armenian Y chromosome haplotypes reveal strong regional structure within a single ethno-national group" (PDF). Hum Genet. 109 (6): 659–674. doi:10.1007/s00439-001-0627-9. PMID 11810279.
- Weale, S; Zhivotovsky, LA; King, R; Mehdi, SQ; Edmonds, CA; Chow, CE; Lin, AA; Mitra, M; Sil, SK (2002). "Y Chromosome Evidence for Anglo-Saxon Mass Migration" (PDF). Mol. Biol. Evol. 19 (7): 1008–1021. doi:10.1093/oxfordjournals.molbev.a004160. PMID 12082121.
{{cite journal}}: Invalid|ref=harv(help). - Wells, R. S.; Yuldasheva, N; Ruzibakiev, R; Underhill, PA; Evseeva, I; Blue-Smith, J; Jin, L; Su, B; et al. (2001). "The Eurasian Heartland: A continental perspective on Y-chromosome diversity". Proc. Natl. Acad. Sci. U.S.A. 98 (18): 10244–9. Bibcode:2001PNAS...9810244W. doi:10.1073/pnas.171305098. PMC 56946. PMID 11526236.
{{cite journal}}: Invalid|ref=harv(help). Also at http://www.pnas.org/cgi/reprint/98/18/10244.pdf - Wells, Spencer (2002). The Journey of Man: A Genetic Odyssey. Princeton University Press. ISBN 0-691-11532-X.
{{cite book}}: Invalid|ref=harv(help). - Wilson, J. F.; Weiss, DA; Richards, M; Thomas, MG; Bradman, N; Goldstein, DB (2001). "Genetic evidence for different male and female roles during cultural transitions in the British Isles". Proc. Natl. Acad. Sci. U.S.A. 98 (9): 5078–5083. Bibcode:2001PNAS...98.5078W. doi:10.1073/pnas.071036898. PMC 33166. PMID 11287634.
- Zerjal, T; Beckman, L; Beckman, G; Mikelsaar, AV; Krumina, A; Kucinskas, V; Hurles, ME; Tyler-Smith, C (2001). "Geographical, linguistic, and cultural influences on genetic diversity: Y-chromosomal distribution in Northern European populations". Mol Biol Evol. 18 (6): 1077–1087. doi:10.1093/oxfordjournals.molbev.a003879. PMID 11371596.
- Zerjal, T; Wells, RS; Yuldasheva, N; Ruzibakiev, R; Tyler-Smith, C (2002). "A Genetic Landscape Reshaped by Recent Events: Y-Chromosomal Insights into Central Asia". American Journal of Human Genetics. 71 (3): 466–482. doi:10.1086/342096. PMC 419996. PMID 12145751.
{{cite journal}}: Invalid|ref=harv(help) - Zhou, Ruixia; An, Lizhe; Wang, Xunling; Shao, Wei; Lin, Gonghua; Yu, Weiping; Yi, Lin; Xu, Shijian; et al. (2007). "Testing the hypothesis of an ancient Roman soldier origin of the Liqian people in northwest China: a Y-chromosome perspective". Journal of Human Genetics. 52 (7): 584–91. doi:10.1007/s10038-007-0155-0. PMID 17579807.
{{cite journal}}: Invalid|ref=harv(help) - Zhao, Zhongming; Khan, Faisal; Borkar, Minal; Herrera, Rene; Agrawal, Suraksha (2009). "Presence of three different paternal lineages among North Indians: A study of 560 Y chromosomes". Annals of Human Biology. 36 (1): 1–14. doi:10.1080/03014460802558522. PMC 2755252. PMID 19058044.
- Zhivotovsky, L; Underhill, PA; Cinnioğlu, C; Kayser, M; Morar, B; Kivisild, T; Scozzari, R; Cruciani, F; et al. (2004). "The effective mutation rate at Y chromosome short tandem repeats, with application to human population-divergence time". American Journal of Human Genetics. 74 (1): 50–61. doi:10.1086/380911. PMC 1181912. PMID 14691732.
