Photosystem I

| Photosystem I | |||||||||
|---|---|---|---|---|---|---|---|---|---|
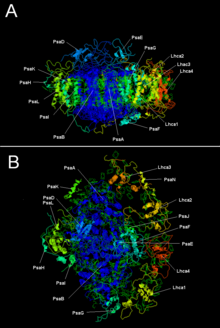 Plant photosystem I with LHC I | |||||||||
| Identifiers | |||||||||
| EC no. | 1.97.1.12 | ||||||||
| Databases | |||||||||
| IntEnz | IntEnz view | ||||||||
| BRENDA | BRENDA entry | ||||||||
| ExPASy | NiceZyme view | ||||||||
| KEGG | KEGG entry | ||||||||
| MetaCyc | metabolic pathway | ||||||||
| PRIAM | profile | ||||||||
| PDB structures | RCSB PDB PDBe PDBsum | ||||||||
| |||||||||

Photosystem I (PSI, or plastocyanin–ferredoxin oxidoreductase) is one of two photosystems in the photosynthetic light reactions of algae, plants, and cyanobacteria. Photosystem I [1] is an integral membrane protein complex that uses light energy to catalyze the transfer of electrons across the thylakoid membrane from plastocyanin to ferredoxin. Ultimately, the electrons that are transferred by Photosystem I are used to produce the moderate-energy hydrogen carrier NADPH.[2] The photon energy absorbed by Photosystem I also produces a proton-motive force that is used to generate ATP. PSI is composed of more than 110 cofactors, significantly more than Photosystem II.[3]
History
[edit]This photosystem is known as PSI because it was discovered before Photosystem II, although future experiments showed that Photosystem II is actually the first enzyme of the photosynthetic electron transport chain. Aspects of PSI were discovered in the 1950s, but the significance of these discoveries was not yet recognized at the time.[4] Louis Duysens first proposed the concepts of Photosystems I and II in 1960, and, in the same year, a proposal by Fay Bendall and Robert Hill assembled earlier discoveries into a coherent theory of serial photosynthetic reactions.[4] Hill and Bendall's hypothesis was later confirmed in experiments conducted in 1961 by the Duysens and Witt groups.[4]
Components and action
[edit]Two main subunits of PSI, PsaA and PsaB, are closely related proteins involved in the binding of the vital electron transfer cofactors P700, Acc, A0, A1, and Fx. PsaA and PsaB are both integral membrane proteins of 730 to 750 amino acids that contain 11 transmembrane segments. A [4Fe-4S] iron-sulfur cluster called Fx is coordinated by four cysteines; two cysteines are provided each by PsaA and PsaB. The two cysteines in each are proximal and located in a loop between the ninth and tenth transmembrane segments. A leucine zipper motif seems to be present [5] downstream of the cysteines and could contribute to dimerisation of PsaA/PsaB. The terminal electron acceptors FA and FB, also [4Fe-4S] iron-sulfur clusters, are located in a 9-kDa protein called PsaC that binds to the PsaA/PsaB core near FX.[6][7]
| Protein subunits | Description |
|---|---|
| PsaA | Related large transmembrane proteins involved in the binding of P700, A0, A1, and Fx. Part of the photosynthetic reaction centre protein family. |
| PsaB | |
| PsaC | Iron-sulfur center; apoprotein for Fa and Fb |
| PsaD | Required for assembly, helps bind ferredoxin. InterPro: IPR003685 |
| PsaE | InterPro: IPR003375 |
| PsaI | May stabilize PsaL. Stabilizes light-harvesting complex II binding.[9] InterPro: IPR001302 |
| PsaJ | InterPro: IPR002615 |
| PsaK | InterPro: IPR035982 |
| PsaL | InterPro: IPR036592 |
| PsaM | InterPro: IPR010010 |
| PsaX | InterPro: IPR012986 |
| cytochrome b6f complex | Soluble protein |
| Fa | From PsaC; In electron transport chain (ETC) |
| Fb | From PsaC; In ETC |
| Fx | From PsaAB; In ETC |
| Ferredoxin | Electron carrier in ETC |
| Plastocyanin | Soluble protein |
| Lipids | Description |
| MGDG II | Monogalactosyldiglyceride lipid |
| PG I | Phosphatidylglycerol phospholipid |
| PG III | Phosphatidylglycerol phospholipid |
| PG IV | Phosphatidylglycerol phospholipid |
| Pigments | Description |
| Chlorophyll a | 90 pigment molecules in antenna system |
| Chlorophyll a | 5 pigment molecules in ETC |
| Chlorophyll a0 | Early electron acceptor of modified chlorophyll in ETC |
| Chlorophyll a′ | 1 pigment molecule in ETC |
| β-Carotene | 22 carotenoid pigment molecules |
| Coenzymes and cofactors | Description |
| QK-A | Early electron acceptor vitamin K1 phylloquinone in ETC |
| QK-B | Early electron acceptor vitamin K1 phylloquinone in ETC |
| FNR | Ferredoxin-NADP+ oxidoreductase enzyme |
| Ca2+ |
Calcium ion |
| Mg2+ |
Magnesium ion |
Photon
[edit]Photoexcitation of the pigment molecules in the antenna complex induces electron and energy transfer.[10]
Antenna complex
[edit]The antenna complex is composed of molecules of chlorophyll and carotenoids mounted on two proteins.[11] These pigment molecules transmit the resonance energy from photons when they become photoexcited. Antenna molecules can absorb all wavelengths of light within the visible spectrum.[12] The number of these pigment molecules varies from organism to organism. For instance, the cyanobacterium Synechococcus elongatus (Thermosynechococcus elongatus) has about 100 chlorophylls and 20 carotenoids, whereas spinach chloroplasts have around 200 chlorophylls and 50 carotenoids.[12][3] Located within the antenna complex of PSI are molecules of chlorophyll called P700 reaction centers. The energy passed around by antenna molecules is directed to the reaction center. There may be as many as 120 or as few as 25 chlorophyll molecules per P700.[13]
P700 reaction center
[edit]The P700 reaction center is composed of modified chlorophyll a that best absorbs light at a wavelength of 700 nm.[14] P700 receives energy from antenna molecules and uses the energy from each photon to raise an electron to a higher energy level (P700*). These electrons are moved in pairs in an oxidation/reduction process from P700* to electron acceptors, leaving behind P700+. The pair of P700* - P700+ has an electric potential of about −1.2 volts. The reaction center is made of two chlorophyll molecules and is therefore referred to as a dimer.[11] The dimer is thought to be composed of one chlorophyll a molecule and one chlorophyll a′ molecule. However, if P700 forms a complex with other antenna molecules, it can no longer be a dimer.[13]
Modified chlorophyll A0 and A1
[edit]The two modified chlorophyll molecules are early electron acceptors in PSI. They are present one per PsaA/PsaB side, forming two branches electrons can take to reach Fx. A0 accepts electrons from P700*, passes it to A1 of the same side, which then passes the electron to the quinone on the same side. Different species seems to have different preferences for either A/B branch.[15]
Phylloquinone
[edit]A phylloquinone, sometimes called vitamin K1,[16] is the next early electron acceptor in PSI. It oxidizes A1 in order to receive the electron and in turn is re-oxidized by Fx, from which the electron is passed to Fb and Fa.[16][17] The reduction of Fx appears to be the rate-limiting step.[15]
Iron–sulfur complex
[edit]Three proteinaceous iron–sulfur reaction centers are found in PSI. Labeled Fx, Fa, and Fb, they serve as electron relays.[18] Fa and Fb are bound to protein subunits of the PSI complex and Fx is tied to the PSI complex.[18] Various experiments have shown some disparity between theories of iron–sulfur cofactor orientation and operation order.[18] In one model, Fx passes an electron to Fa, which passes it on to Fb to reach the ferredoxin.[15]
Ferredoxin
[edit]Ferredoxin (Fd) is a soluble protein that facilitates reduction of NADP+
to NADPH.[19] Fd moves to carry an electron either to a lone thylakoid or to an enzyme that reduces NADP+
.[19] Thylakoid membranes have one binding site for each function of Fd.[19] The main function of Fd is to carry an electron from the iron-sulfur complex to the enzyme ferredoxin–NADP+
reductase.[19]
Ferredoxin–NADP+
reductase (FNR)
[edit]This enzyme transfers the electron from reduced ferredoxin to NADP+
to complete the reduction to NADPH.[20] FNR may also accept an electron from NADPH by binding to it.[20]
Plastocyanin
[edit]Plastocyanin is an electron carrier that transfers the electron from cytochrome b6f to the P700 cofactor of PSI in its ionized state P700+.[10][21]
Ycf4 protein domain
[edit]The Ycf4 protein domain found on the thylakoid membrane is vital to photosystem I. This thylakoid transmembrane protein helps assemble the components of photosystem I. Without it, photosynthesis would be inefficient.[22]
Evolution
[edit]Molecular data show that PSI likely evolved from the photosystems of green sulfur bacteria. The photosystems of green sulfur bacteria and those of cyanobacteria, algae, and higher plants are not the same, but there are many analogous functions and similar structures. Three main features are similar between the different photosystems.[23] First, redox potential is negative enough to reduce ferredoxin.[23] Next, the electron-accepting reaction centers include iron–sulfur proteins.[23] Last, redox centres in complexes of both photosystems are constructed upon a protein subunit dimer.[23] The photosystem of green sulfur bacteria even contains all of the same cofactors of the electron transport chain in PSI.[23] The number and degree of similarities between the two photosystems strongly indicates that PSI and the analogous photosystem of green sulfur bacteria evolved from a common ancestral photosystem.
See also
[edit]References
[edit]- ^ Golbeck JH (1987). "Structure, function and organization of the Photosystem I reaction center complex". Biochimica et Biophysica Acta (BBA) - Reviews on Bioenergetics. 895 (3): 167–204. doi:10.1016/s0304-4173(87)80002-2. PMID 3333014.
- ^ Yamori W, Shikanai T (April 2016). "Physiological Functions of Cyclic Electron Transport Around Photosystem I in Sustaining Photosynthesis and Plant Growth". Annual Review of Plant Biology. 67: 81–106. doi:10.1146/annurev-arplant-043015-112002. PMID 26927905.
- ^ a b Nelson N, Yocum CF (2006). "Structure and function of photosystems I and II". Annual Review of Plant Biology. 57: 521–65. doi:10.1146/annurev.arplant.57.032905.105350. PMID 16669773.
- ^ a b c Fromme P, Mathis P (2004). "Unraveling the photosystem I reaction center: a history, or the sum of many efforts". Photosynthesis Research. 80 (1–3): 109–24. Bibcode:2004PhoRe..80..109F. doi:10.1023/B:PRES.0000030657.88242.e1. PMID 16328814. S2CID 13832448.
- ^ Webber AN, Malkin R (May 1990). "Photosystem I reaction-centre proteins contain leucine zipper motifs. A proposed role in dimer formation". FEBS Letters. 264 (1): 1–4. Bibcode:1990FEBSL.264....1W. doi:10.1016/0014-5793(90)80749-9. PMID 2186925. S2CID 42294700.
- ^ Jagannathan B, Golbeck JH (April 2009). "Breaking biological symmetry in membrane proteins: the asymmetrical orientation of PsaC on the pseudo-C2 symmetric Photosystem I core". Cellular and Molecular Life Sciences. 66 (7): 1257–70. doi:10.1007/s00018-009-8673-x. PMC 11131447. PMID 19132290. S2CID 32418758.
- ^ Jagannathan B, Golbeck JH (June 2009). "Understanding of the binding interface between PsaC and the PsaA/PsaB heterodimer in photosystem I". Biochemistry. 48 (23): 5405–16. doi:10.1021/bi900243f. PMID 19432395.
- ^ Saenger W, Jordan P, Krauss N (April 2002). "The assembly of protein subunits and cofactors in photosystem I". Current Opinion in Structural Biology. 12 (2): 244–54. doi:10.1016/S0959-440X(02)00317-2. PMID 11959504.
- ^ Plöchinger, Magdalena; Torabi, Salar; Rantala, Marjaana; Tikkanen, Mikko; Suorsa, Marjaana; Jensen, Poul-Erik; Aro, Eva Mari; Meurer, Jörg (September 2016). "The Low Molecular Weight Protein PsaI Stabilizes the Light-Harvesting Complex II Docking Site of Photosystem I". Plant Physiology. 172 (1): 450–463. doi:10.1104/pp.16.00647. PMC 5074619. PMID 27406169.
- ^ a b Raven PH, Evert RF, Eichhorn SE (2005). "Photosynthesis, Light, and Life". Biology of Plants (7th ed.). New York: W. H. Freeman. pp. 121–127. ISBN 978-0-7167-1007-3.
- ^ a b Zeiger E, Taiz L (2006). "Ch. 7: Topic 7.8: Photosystem I". Plant Physiology (4th ed.). Sunderland, MA: Sinauer Associates. ISBN 0-87893-856-7.[permanent dead link]
- ^ a b "The Photosynthetic Process". Archived from the original on 2009-02-19.
- ^ a b Shubin VV, Karapetyan NV, Krasnovsky AA (January 1986). "Molecular arrangement of pigment-protein complex of photosystem 1". Photosynthesis Research. 9 (1–2): 3–12. Bibcode:1986PhoRe...9....3S. doi:10.1007/BF00029726. PMID 24442279. S2CID 26158482.
- ^ Rutherford AW, Heathcote P (December 1985). "Primary photochemistry in photosystem-I". Photosynthesis Research. 6 (4): 295–316. Bibcode:1985PhoRe...6..295R. doi:10.1007/BF00054105. PMID 24442951. S2CID 21845584.
- ^ a b c Grotjohann, I; Fromme, P (2013). "Photosystem I". Encyclopedia of biological chemistry (Second ed.). London. pp. 503–507. doi:10.1016/B978-0-12-378630-2.00287-5. ISBN 978-0-12-378630-2.
{{cite book}}: CS1 maint: location missing publisher (link) - ^ a b Itoh S, Iwaki M (1989). "Vitamin K1 (Phylloquinone) Restores the Turnover of FeS centers of Ether-extracted Spinach PSI Particles". FEBS Letters. 243 (1): 47–52. doi:10.1016/0014-5793(89)81215-3. S2CID 84602152.
- ^ Palace GP, Franke JE, Warden JT (May 1987). "Is phylloquinone an obligate electron carrier in photosystem I?". FEBS Letters. 215 (1): 58–62. Bibcode:1987FEBSL.215...58P. doi:10.1016/0014-5793(87)80113-8. PMID 3552735. S2CID 42983611.
- ^ a b c Vassiliev IR, Antonkine ML, Golbeck JH (October 2001). "Iron-sulfur clusters in type I reaction centers". Biochimica et Biophysica Acta (BBA) - Bioenergetics. 1507 (1–3): 139–60. doi:10.1016/S0005-2728(01)00197-9. PMID 11687212.
- ^ a b c d Forti G, Maria P, Grubas G (1985). "Two Sites of Interaction of Ferredoxin with thylakoids". FEBS Letters. 186 (2): 149–152. Bibcode:1985FEBSL.186..149F. doi:10.1016/0014-5793(85)80698-0. S2CID 83495051.
- ^ a b Madoz J, Fernández Recio J, Gómez Moreno C, Fernández VM (November 1998). "Investigation of the Diaphorase Reaction of Ferredoxin–NADP+
Reductase by Electrochemical Methods" (PDF). Bioelectrochemistry and Bioenergetics. 47 (1): 179–183. doi:10.1016/S0302-4598(98)00175-5. - ^ Hope AB (January 2000). "Electron transfers amongst cytochrome f, plastocyanin and photosystem I: kinetics and mechanisms". Biochimica et Biophysica Acta (BBA) - Bioenergetics. 1456 (1): 5–26. doi:10.1016/S0005-2728(99)00101-2. PMID 10611452.
- ^ Boudreau E, Takahashi Y, Lemieux C, Turmel M, Rochaix JD (October 1997). "The chloroplast ycf3 and ycf4 open reading frames of Chlamydomonas reinhardtii are required for the accumulation of the photosystem I complex". The EMBO Journal. 16 (20): 6095–104. doi:10.1093/emboj/16.20.6095. PMC 1326293. PMID 9321389.
- ^ a b c d e Lockau W, Nitschke W (1993). "Photosystem I and its Bacterial Counterparts". Physiologia Plantarum. 88 (2): 372–381. doi:10.1111/j.1399-3054.1993.tb05512.x.
