CTP synthetase
| CTP synthase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
 | |||||||||
| Identifiers | |||||||||
| EC no. | 6.3.4.2 | ||||||||
| CAS no. | 9023-56-7 | ||||||||
| Databases | |||||||||
| IntEnz | IntEnz view | ||||||||
| BRENDA | BRENDA entry | ||||||||
| ExPASy | NiceZyme view | ||||||||
| KEGG | KEGG entry | ||||||||
| MetaCyc | metabolic pathway | ||||||||
| PRIAM | profile | ||||||||
| PDB structures | RCSB PDB PDBe PDBsum | ||||||||
| Gene Ontology | AmiGO / QuickGO | ||||||||
| |||||||||
CTP synthase is an enzyme involved in pyrimidine biosynthesis that interconverts UTP and CTP.[1][2]
Reaction mechanism
CTP (cytidine triphosphate) synthetase catalyzes the last committed step in pyrimidine nucleotide biosynthesis:[3]
ATP + UTP + glutamine → ADP + Pi + CTP + glutamate
It is the rate-limiting enzyme for the synthesis of cytosine nucleotides from both the de novo and uridine salvage pathways.[4]
The reaction proceeds by the ATP-dependent phosphorylation of UTP on the 4-oxygen atom, making the 4-carbon electrophilic and vulnerable to reaction with ammonia.[5] The source of the amino group in CTP is glutamine, which is hydrolysed in a glutamine amidotransferase domain to produce ammonia. This is then channeled through the interior of the enzyme to the synthetase domain.[6][7] Here, ammonia reacts with the intermediate 4-phosphoryl UTP.[8]

Isozymes
Two isozymes with CTP synthase activity exist in humans, encoded by the following genes:
Structure

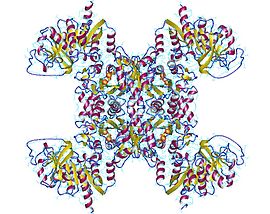
Active CTP synthase exists as a homotetrameric enzyme. At low enzyme concentrations and in the absence of ATP and UTP, CTP synthase exists as inactive monomer. As enzyme concentration increases, it polymerizes first to a dimer (such as the form shown to the left) and, in the presence of ATP and UTP, forms a tetramer.[5][9]
The enzyme contains two major domains, responsible for the aminotransferase and synthase activity, respectively. The amidotransferase domains are located away from the tetramer interfaces and are not affected by the oligomeric state. The ATP-binding site and CTP-binding site in the synthase domain are located at the tetramer interface. It is for this reason that ATP and UTP are required for tetramerization.[10]
Regulation
CTP synthase is precisely regulated by the intracellular concentrations of CTP and UTP, and both hCTPS1 and hCTPS2 have been seen to be maximally active at physiological concentrations of ATP, GTP, and glutamine.[11]
The activity of human CTPS1 isozyme has been demonstrated to be inhibited by phosphorylation.[12] One major example of this is phosphorylation of the Ser-571 residue by glycogen synthase kinase 3 (GSK3) in response to low serum conditions.[13] Additionally, Ser568 has been seen to be phosphorylated by casein kinase 1, inhibiting CTP synthase activity.[11]
CTP is also subject to various forms of allosteric regulation. GTP acts as an allosteric activator that strongly promotes the hydrolysis of glutamine, but is also inhibiting to glutamine-dependent CTP formation at high concentrations.[14] This acts to balance the relative amounts of purine and pyrimidine nucleotides. The reaction product CTP also serves as an allosteric inhibitor. The triphosphate binding site overlaps with that of UTP, but the nucleoside moiety of CTP binds in an alternative pocket opposite the binding site for UTP.[15]
CTP synthase levels have been shown to be dependent on levels of the transcription factor Myc. In turn, CTP synthase activity is required for Myc related phenotypes.[16]
The glutamine analog DON has also been seen to act as an irreversible inhibitor, and has been used as an anti-cancer agent.[17]
Filaments
CTP synthase has been reported to form filaments in several different organisms. These include bacteria (C. crescentus),[18] yeast (S. cerevisiae),[19] fruit flies (D. melanogaster)[20] and human cells.[21] These filamentous structures have been referred to as cytoplasmic rods and rings,[22] cytoophidia (from the Greek "cyto" meaning cell and "ophidium" meaning serpent, due to the structures morphology) or simply CTP synthase filaments. It has been shown that filamentation downregulates or upregulates CTP synthase activity depending on the species.[23][24][25][26][27] In Drosophila, only one of the CTP synthase isoform forms the filament.[28] Since the discovery of this novel mode of enzyme regulation in CTP synthase, multiple other enzymes have been shown to exhibit similar characteristics, suggesting that this is an important and well conserved strategy for enzymatic regulation.[29] CTP synthase remains a model enzyme for the study of filament formation.
Clinical significance
Upregulated CTP synthase activity has been widely seen in human and rodent tumors.[30]
Mutations in the CTP synthase have been seen to confer resistance to cytotoxic drugs such as cytosine arabinoside (ara-C) in a Chinese hamster ovary (CHO) cell model of leukemia though such mutations were not found in human patients with ara-C resistance.[31]
See also
References
- ^ Lieberman I (October 1956). "Enzymatic amination of uridine triphosphate to cytidine triphosphate". The Journal of Biological Chemistry. 222 (2): 765–75. PMID 13367044.
- ^ Long CW, Levitzki A, Koshland DE (January 1970). "The subunit structure and subunit interactions of cytidine triphosphate synthetase". The Journal of Biological Chemistry. 245 (1): 80–7. PMID 5411547.
- ^ Koshland DE, Levitzki A (1974). "CTP Synthetase and Related Enzymes". In Boyer PD (ed.). The Enzymes (3rd ed.). New York: Academic Press. pp. 539–59. ISBN 978-0-12-122710-4.
- ^ van Kuilenburg AB, Meinsma R, Vreken P, Waterham HR, van Gennip AH (2000). "Isoforms of human CTP synthetase". Advances in Experimental Medicine and Biology. 486: 257–61. doi:10.1007/0-306-46843-3_50. ISBN 978-0-306-46515-4. PMID 11783495.
- ^ a b von der Saal W, Anderson PM, Villafranca JJ (December 1985). "Mechanistic investigations of Escherichia coli cytidine-5'-triphosphate synthetase. Detection of an intermediate by positional isotope exchange experiments". The Journal of Biological Chemistry. 260 (28): 14993–7. PMID 2933396.
- ^ Levitzki A, Koshland DE (August 1971). "Cytidine triphosphate synthetase. Covalent intermediates and mechanisms of action". Biochemistry. 10 (18): 3365–71. doi:10.1021/bi00794a008. PMID 4940761.
- ^ Endrizzi JA, Kim H, Anderson PM, Baldwin EP (June 2004). "Crystal structure of Escherichia coli cytidine triphosphate synthetase, a nucleotide-regulated glutamine amidotransferase/ATP-dependent amidoligase fusion protein and homologue of anticancer and antiparasitic drug targets". Biochemistry. 43 (21): 6447–63. doi:10.1021/bi0496945. PMC 2891762. PMID 15157079.
- ^ Lewis DA, Villafranca JJ (October 1989). "Investigation of the mechanism of CTP synthetase using rapid quench and isotope partitioning methods". Biochemistry. 28 (21): 8454–9. doi:10.1021/bi00447a027. PMID 2532543.
- ^ Anderson PM (June 1983). "CTP synthetase from Escherichia coli: an improved purification procedure and characterization of hysteretic and enzyme concentration effects on kinetic properties". Biochemistry. 22 (13): 3285–92. doi:10.1021/bi00282a038. PMID 6349684.
- ^ Lauritsen I, Willemoës M, Jensen KF, Johansson E, Harris P (February 2011). "Structure of the dimeric form of CTP synthase from Sulfolobus solfataricus". Acta Crystallographica Section F. 67 (Pt 2): 201–8. doi:10.1107/S1744309110052334. PMC 3034608. PMID 21301086.
- ^ a b Kassel KM, Au DR, Higgins MJ, Hines M, Graves LM (October 2010). "Regulation of human cytidine triphosphate synthetase 2 by phosphorylation". The Journal of Biological Chemistry. 285 (44): 33727–36. doi:10.1074/jbc.M110.178566. PMC 2962471. PMID 20739275.
{{cite journal}}: CS1 maint: unflagged free DOI (link) - ^ Carman GM, Kersting MC (February 2004). "Phospholipid synthesis in yeast: regulation by phosphorylation". Biochemistry and Cell Biology. 82 (1): 62–70. doi:10.1139/o03-064. PMID 15052328.
- ^ Higgins MJ, Graves PR, Graves LM (October 2007). "Regulation of human cytidine triphosphate synthetase 1 by glycogen synthase kinase 3". The Journal of Biological Chemistry. 282 (40): 29493–503. doi:10.1074/jbc.M703948200. PMID 17681942.
- ^ Lunn FA, MacDonnell JE, Bearne SL (January 2008). "Structural requirements for the activation of Escherichia coli CTP synthase by the allosteric effector GTP are stringent, but requirements for inhibition are lax". The Journal of Biological Chemistry. 283 (4): 2010–20. doi:10.1074/jbc.M707803200. PMID 18003612.
- ^ Endrizzi JA, Kim H, Anderson PM, Baldwin EP (October 2005). "Mechanisms of product feedback regulation and drug resistance in cytidine triphosphate synthetases from the structure of a CTP-inhibited complex". Biochemistry. 44 (41): 13491–9. doi:10.1021/bi051282o. PMC 2891682. PMID 16216072.
- ^ Aughey GN, Grice SJ, Liu JL (February 2016). "The Interplay between Myc and CTP Synthase in Drosophila". PLOS Genetics. 12 (2): e1005867. doi:10.1371/journal.pgen.1005867. PMC 4759343. PMID 26889675.
{{cite journal}}: CS1 maint: unflagged free DOI (link) - ^ Ahluwalia GS, Grem JL, Hao Z, Cooney DA (1990). "Metabolism and action of amino acid analog anti-cancer agents". Pharmacology & Therapeutics. 46 (2): 243–71. doi:10.1016/0163-7258(90)90094-I. PMID 2108451.
- ^ Ingerson-Mahar M, Briegel A, Werner JN, Jensen GJ, Gitai Z (August 2010). "The metabolic enzyme CTP synthase forms cytoskeletal filaments". Nature Cell Biology. 12 (8): 739–46. doi:10.1038/ncb2087. PMC 3210567. PMID 20639870.
- ^ Noree C, Sato BK, Broyer RM, Wilhelm JE (August 2010). "Identification of novel filament-forming proteins in Saccharomyces cerevisiae and Drosophila melanogaster". The Journal of Cell Biology. 190 (4): 541–51. doi:10.1083/jcb.201003001. PMC 2928026. PMID 20713603.
- ^ Liu JL (May 2010). "Intracellular compartmentation of CTP synthase in Drosophila". Journal of Genetics and Genomics = Yi Chuan Xue Bao. 37 (5): 281–96. doi:10.1016/S1673-8527(09)60046-1. PMID 20513629.
- ^ Chen K, Zhang J, Tastan ÖY, Deussen ZA, Siswick MY, Liu JL (September 2011). "Glutamine analogs promote cytoophidium assembly in human and Drosophila cells". Journal of Genetics and Genomics = Yi Chuan Xue Bao. 38 (9): 391–402. doi:10.1016/j.jgg.2011.08.004. PMID 21930098.
- ^ Carcamo WC, Satoh M, Kasahara H, Terada N, Hamazaki T, Chan JY, Yao B, Tamayo S, Covini G, von Mühlen CA, Chan EK (2011). "Induction of cytoplasmic rods and rings structures by inhibition of the CTP and GTP synthetic pathway in mammalian cells". PLOS ONE. 6 (12): e29690. doi:10.1371/journal.pone.0029690. PMC 3248424. PMID 22220215.
{{cite journal}}: CS1 maint: unflagged free DOI (link) - ^ Lynch EM, Hicks DR, Shepherd M, Endrizzi JA, Maker A, Hansen JM, Barry RM, Gitai Z, Baldwin EP, Kollman JM (June 2017). "Human CTP synthase filament structure reveals the active enzyme conformation". Nature Structural & Molecular Biology. 24 (6): 507–514. doi:10.1038/nsmb.3407. PMC 5472220. PMID 28459447.
- ^ Barry RM, Bitbol AF, Lorestani A, Charles EJ, Habrian CH, Hansen JM, Li HJ, Baldwin EP, Wingreen NS, Kollman JM, Gitai Z (July 2014). "Large-scale filament formation inhibits the activity of CTP synthetase". eLife. 3: e03638. doi:10.7554/eLife.03638. PMC 4126345. PMID 25030911.
{{cite journal}}: CS1 maint: unflagged free DOI (link) - ^ Aughey GN, Grice SJ, Shen QJ, Xu Y, Chang CC, Azzam G, Wang PY, Freeman-Mills L, Pai LM, Sung LY, Yan J, Liu JL (October 2014). "Nucleotide synthesis is regulated by cytoophidium formation during neurodevelopment and adaptive metabolism". Biology Open. 3 (11): 1045–56. doi:10.1242/bio.201410165. PMC 4232762. PMID 25326513.
- ^ Noree C, Monfort E, Shiau AK, Wilhelm JE (August 2014). "Common regulatory control of CTP synthase enzyme activity and filament formation". Molecular Biology of the Cell. 25 (15): 2282–90. doi:10.1091/mbc.E14-04-0912. PMC 4116302. PMID 24920825.
- ^ Barry RM, Gitai Z (December 2011). "Self-assembling enzymes and the origins of the cytoskeleton". Current Opinion in Microbiology. 14 (6): 704–11. doi:10.1016/j.mib.2011.09.015. PMC 3234109. PMID 22014508.
- ^ Azzam G, Liu JL (February 2013). "Only one isoform of Drosophila melanogaster CTP synthase forms the cytoophidium". PLOS Genetics. 9 (2): e1003256. doi:10.1371/journal.pgen.1003256. PMC 3573105. PMID 23459760.
{{cite journal}}: CS1 maint: unflagged free DOI (link) - ^ Aughey GN, Liu JL (2015). "Metabolic regulation via enzyme filamentation". Critical Reviews in Biochemistry and Molecular Biology. 51 (4): 282–93. doi:10.3109/10409238.2016.1172555. PMC 4915340. PMID 27098510.
- ^ Kizaki H, Williams JC, Morris HP, Weber G (November 1980). "Increased cytidine 5'-triphosphate synthetase activity in rat and human tumors". Cancer Research. 40 (11): 3921–7. PMID 7471043.
- ^ Whelan J, Smith T, Phear G, Rohatiner A, Lister A, Meuth M (February 1994). "Resistance to cytosine arabinoside in acute leukemia: the significance of mutations in CTP synthetase". Leukemia. 8 (2): 264–5. PMID 8309250.
Further reading
External links
- CTP+synthetase at the U.S. National Library of Medicine Medical Subject Headings (MeSH)
