DNA-binding protein: Difference between revisions
Citation bot (talk | contribs) m Alter: template type. Add: year, pages, issue, volume, journal, title, pmc, pmid, doi, isbn, series, citeseerx, author pars. 1-8. Converted bare reference to cite template. Removed parameters. Formatted dashes. | You can use this bot yourself. Report bugs here. | User-activated. |
|||
| Line 10: | Line 10: | ||
==Non-specific DNA-protein interactions== |
==Non-specific DNA-protein interactions== |
||
Structural proteins that bind DNA are well-understood examples of non-specific DNA-protein interactions. Within chromosomes, DNA is held in complexes with structural proteins. These proteins organize the DNA into a compact structure called [[chromatin]]. In [[eukaryote]]s, this structure involves DNA binding to a complex of small basic proteins called [[histone]]s. In [[prokaryote]]s, multiple types of proteins are involved.<ref>{{cite journal |vauthors=Sandman K, Pereira S, Reeve J |title=Diversity of prokaryotic chromosomal proteins and the origin of the nucleosome |journal=Cell Mol Life Sci |volume=54 |issue=12 |pages=1350–64 |year=1998 |pmid=9893710 |doi=10.1007/s000180050259}}</ref><ref>{{cite journal |author=Dame RT |title=The role of nucleoid-associated proteins in the organization and compaction of bacterial chromatin |journal=Mol. Microbiol. |volume=56 |issue=4 |pages=858–70 |year=2005 |pmid=15853876 |doi=10.1111/j.1365-2958.2005.04598.x}}</ref> The histones form a disk-shaped complex called a [[nucleosome]], which contains two complete turns of double-stranded DNA wrapped around its surface. These non-specific interactions are formed through basic residues in the histones making [[ionic bond]]s to the acidic sugar-phosphate backbone of the DNA, and are therefore largely independent of the base sequence.<ref>{{cite journal |vauthors=Luger K, Mäder A, Richmond R, Sargent D, Richmond T |title=Crystal structure of the nucleosome core particle at 2.8 A resolution |journal=Nature |volume=389 |issue=6648 |pages=251–60 |year=1997 |pmid=9305837 |doi= 10.1038/38444|bibcode=1997Natur.389..251L }}</ref> [[Chemical]] modifications of these basic [[amino acid]] residues include [[methylation]], [[phosphorylation]] and [[acetylation]].<ref>{{cite journal |vauthors=Jenuwein T, Allis C |title=Translating the histone code |journal=Science |volume=293 |issue=5532 |pages=1074–80 |year=2001 |pmid=11498575 |doi=10.1126/science.1063127}}</ref> These chemical changes alter the strength of the interaction between the DNA and the histones, making the DNA more or less accessible to [[transcription factor]]s and changing the rate of transcription.<ref>{{cite |
Structural proteins that bind DNA are well-understood examples of non-specific DNA-protein interactions. Within chromosomes, DNA is held in complexes with structural proteins. These proteins organize the DNA into a compact structure called [[chromatin]]. In [[eukaryote]]s, this structure involves DNA binding to a complex of small basic proteins called [[histone]]s. In [[prokaryote]]s, multiple types of proteins are involved.<ref>{{cite journal |vauthors=Sandman K, Pereira S, Reeve J |title=Diversity of prokaryotic chromosomal proteins and the origin of the nucleosome |journal=Cell Mol Life Sci |volume=54 |issue=12 |pages=1350–64 |year=1998 |pmid=9893710 |doi=10.1007/s000180050259}}</ref><ref>{{cite journal |author=Dame RT |title=The role of nucleoid-associated proteins in the organization and compaction of bacterial chromatin |journal=Mol. Microbiol. |volume=56 |issue=4 |pages=858–70 |year=2005 |pmid=15853876 |doi=10.1111/j.1365-2958.2005.04598.x}}</ref> The histones form a disk-shaped complex called a [[nucleosome]], which contains two complete turns of double-stranded DNA wrapped around its surface. These non-specific interactions are formed through basic residues in the histones making [[ionic bond]]s to the acidic sugar-phosphate backbone of the DNA, and are therefore largely independent of the base sequence.<ref>{{cite journal |vauthors=Luger K, Mäder A, Richmond R, Sargent D, Richmond T |title=Crystal structure of the nucleosome core particle at 2.8 A resolution |journal=Nature |volume=389 |issue=6648 |pages=251–60 |year=1997 |pmid=9305837 |doi= 10.1038/38444|bibcode=1997Natur.389..251L }}</ref> [[Chemical]] modifications of these basic [[amino acid]] residues include [[methylation]], [[phosphorylation]] and [[acetylation]].<ref>{{cite journal |vauthors=Jenuwein T, Allis C |title=Translating the histone code |journal=Science |volume=293 |issue=5532 |pages=1074–80 |year=2001 |pmid=11498575 |doi=10.1126/science.1063127|citeseerx=10.1.1.453.900 }}</ref> These chemical changes alter the strength of the interaction between the DNA and the histones, making the DNA more or less accessible to [[transcription factor]]s and changing the rate of transcription.<ref>{{cite book |author=Ito T |title=Nucleosome assembly and remodelling |journal=Curr Top Microbiol Immunol |volume=274 |pages=1–22 |pmid=12596902 |year=2003 |doi=10.1007/978-3-642-55747-7_1|series=Current Topics in Microbiology and Immunology |isbn=978-3-642-62909-9 }}</ref> Other non-specific DNA-binding proteins in chromatin include the high-mobility group (HMG) proteins, which bind to bent or distorted DNA.<ref>{{cite journal |author=Thomas J |title=HMG1 and 2: architectural DNA-binding proteins |journal=Biochem Soc Trans |volume=29 |issue=Pt 4 |pages=395–401 |year=2001 |pmid=11497996 |doi=10.1042/BST0290395}}</ref> Biophysical studies show that these architectural HMG proteins bind, bend and loop DNA to perform its biological functions.<ref>{{Cite journal |doi = 10.1093/nar/gku635|pmid = 25063301|pmc = 4132745|title = DNA bridging and looping by HMO1 provides a mechanism for stabilizing nucleosome-free chromatin|journal = Nucleic Acids Research|volume = 42|issue = 14|pages = 8996–9004|year = 2014|last1 = Murugesapillai|first1 = Divakaran|last2 = McCauley|first2 = Micah J.|last3 = Huo|first3 = Ran|last4 = Nelson Holte|first4 = Molly H.|last5 = Stepanyants|first5 = Armen|last6 = Maher|first6 = L. James|last7 = Israeloff|first7 = Nathan E.|last8 = Williams|first8 = Mark C.}}</ref><ref>{{Cite journal | doi=10.1007/s12551-016-0236-4| pmid=28303166|title = Single-molecule studies of high-mobility group B architectural DNA bending proteins| journal=Biophysical Reviews| volume=9| pages=17–40|year = 2017|last1 = Murugesapillai|first1 = Divakaran| last2=McCauley| first2=Micah J.| last3=Maher| first3=L. James| last4=Williams| first4=Mark C.}}</ref> These proteins are important in bending arrays of nucleosomes and arranging them into the larger structures that form chromosomes.<ref>{{cite journal |vauthors=Grosschedl R, Giese K, Pagel J |title=HMG domain proteins: architectural elements in the assembly of nucleoprotein structures |journal=Trends Genet |volume=10 |issue=3 |pages=94–100 |year=1994 |pmid=8178371 |doi=10.1016/0168-9525(94)90232-1}}</ref> |
||
==Proteins that specifically bind single-stranded DNA== |
==Proteins that specifically bind single-stranded DNA== |
||
| Line 24: | Line 24: | ||
Protein–DNA interactions occur when a [[protein]] binds a molecule of [[DNA]], often to regulate the [[Function (biology)|biological function]] of DNA, usually the [[Gene expression|expression]] of a [[gene]]. Among the proteins that bind to DNA are [[transcription factors]] that activate or repress gene expression by binding to DNA motifs and [[histones]] that form part of the structure of DNA and bind to it less specifically. Also proteins that [[DNA repair|repair DNA]] such as [[uracil-DNA glycosylase]] interact closely with it. |
Protein–DNA interactions occur when a [[protein]] binds a molecule of [[DNA]], often to regulate the [[Function (biology)|biological function]] of DNA, usually the [[Gene expression|expression]] of a [[gene]]. Among the proteins that bind to DNA are [[transcription factors]] that activate or repress gene expression by binding to DNA motifs and [[histones]] that form part of the structure of DNA and bind to it less specifically. Also proteins that [[DNA repair|repair DNA]] such as [[uracil-DNA glycosylase]] interact closely with it. |
||
In general, proteins bind to DNA in the [[major groove]]; however, there are exceptions.<ref name="pmid9646864">{{cite journal|vauthors=Bewley CA, Gronenborn AM, Clore GM|year=1998|title=Minor groove-binding architectural proteins: structure, function, and DNA recognition|url=|journal=Annu Rev Biophys Biomol Struct|volume=27|issue=|pages=105–31|doi=10.1146/annurev.biophys.27.1.105|pmid=9646864|pmc=4781445}}</ref> Protein–DNA interaction are of mainly two types, either specific interaction, or non-specific interaction. Recent single-molecule experiments showed that DNA binding proteins undergo of rapid rebinding in order to bind in correct orientation for recognizing the target site.<ref>{{Cite journal|last=Ganji|first=Mahipal|last2=Docter|first2=Margreet|last3=Le Grice|first3=Stuart F. J.|last4=Abbondanzieri|first4=Elio A.|date=2016-09-30|title=DNA binding proteins explore multiple local configurations during docking via rapid rebinding |
In general, proteins bind to DNA in the [[major groove]]; however, there are exceptions.<ref name="pmid9646864">{{cite journal|vauthors=Bewley CA, Gronenborn AM, Clore GM|year=1998|title=Minor groove-binding architectural proteins: structure, function, and DNA recognition|url=|journal=Annu Rev Biophys Biomol Struct|volume=27|issue=|pages=105–31|doi=10.1146/annurev.biophys.27.1.105|pmid=9646864|pmc=4781445}}</ref> Protein–DNA interaction are of mainly two types, either specific interaction, or non-specific interaction. Recent single-molecule experiments showed that DNA binding proteins undergo of rapid rebinding in order to bind in correct orientation for recognizing the target site.<ref>{{Cite journal|last=Ganji|first=Mahipal|last2=Docter|first2=Margreet|last3=Le Grice|first3=Stuart F. J.|last4=Abbondanzieri|first4=Elio A.|date=2016-09-30|title=DNA binding proteins explore multiple local configurations during docking via rapid rebinding|journal=Nucleic Acids Research|volume=44|issue=17|pages=8376–8384|doi=10.1093/nar/gkw666|issn=0305-1048|pmc=5041478|pmid=27471033}}</ref> |
||
=== Design === |
=== Design === |
||
| Line 40: | Line 40: | ||
=== Manipulating the interactions === |
=== Manipulating the interactions === |
||
The protein–DNA interactions can be modulated using stimuli like ionic strength of the buffer, macromolecular crowding,<ref>{{Cite journal|last=Ganji|first=Mahipal|last2=Docter|first2=Margreet|last3=Grice|first3=Stuart F.J. Le|last4=Abbondanzieri|first4=Elio A.|date=2016-09-30|title=DNA binding proteins explore multiple local configurations during docking via rapid rebinding |
The protein–DNA interactions can be modulated using stimuli like ionic strength of the buffer, macromolecular crowding,<ref>{{Cite journal|last=Ganji|first=Mahipal|last2=Docter|first2=Margreet|last3=Grice|first3=Stuart F.J. Le|last4=Abbondanzieri|first4=Elio A.|date=2016-09-30|title=DNA binding proteins explore multiple local configurations during docking via rapid rebinding|journal=Nucleic Acids Research|language=en|volume=44|issue=17|pages=8376–8384|doi=10.1093/nar/gkw666|issn=0305-1048|pmc=5041478|pmid=27471033}}</ref> temperature, pH and electric field. This can lead to reversible dissociation/association of the protein–DNA complex.<ref>Hianik, T. and Wang, J. (2009), Electrochemical Aptasensors – Recent Achievements and Perspectives. Electroanalysis, 21: 1223–1235. {{doi|10.1002/elan.200904566}}</ref><ref>Gosai, A. et al. (2016) Electrical Stimulus Controlled Binding/Unbinding of Human Thrombin-Aptamer Complex. Sci. Rep. 6, 37449; {{doi|10.1038/srep37449}} {{PMC|5118750}}</ref> |
||
== See also == |
== See also == |
||
* [[bZIP domain]] |
* [[bZIP domain]] |
||
Revision as of 11:13, 15 February 2019

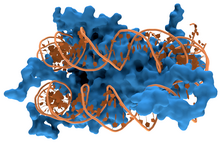


DNA-binding proteins are proteins that have DNA-binding domains and thus have a specific or general affinity for single- or double-stranded DNA.[3][4][5] Sequence-specific DNA-binding proteins generally interact with the major groove of B-DNA, because it exposes more functional groups that identify a base pair. However, there are some known minor groove DNA-binding ligands such as netropsin,[6] distamycin, Hoechst 33258, pentamidine, DAPI and others.[7]
Examples
DNA-binding proteins include transcription factors which modulate the process of transcription, various polymerases, nucleases which cleave DNA molecules, and histones which are involved in chromosome packaging and transcription in the cell nucleus. DNA-binding proteins can incorporate such domains as the zinc finger, the helix-turn-helix, and the leucine zipper (among many others) that facilitate binding to nucleic acid. There are also more unusual examples such as transcription activator like effectors.
Non-specific DNA-protein interactions
Structural proteins that bind DNA are well-understood examples of non-specific DNA-protein interactions. Within chromosomes, DNA is held in complexes with structural proteins. These proteins organize the DNA into a compact structure called chromatin. In eukaryotes, this structure involves DNA binding to a complex of small basic proteins called histones. In prokaryotes, multiple types of proteins are involved.[8][9] The histones form a disk-shaped complex called a nucleosome, which contains two complete turns of double-stranded DNA wrapped around its surface. These non-specific interactions are formed through basic residues in the histones making ionic bonds to the acidic sugar-phosphate backbone of the DNA, and are therefore largely independent of the base sequence.[10] Chemical modifications of these basic amino acid residues include methylation, phosphorylation and acetylation.[11] These chemical changes alter the strength of the interaction between the DNA and the histones, making the DNA more or less accessible to transcription factors and changing the rate of transcription.[12] Other non-specific DNA-binding proteins in chromatin include the high-mobility group (HMG) proteins, which bind to bent or distorted DNA.[13] Biophysical studies show that these architectural HMG proteins bind, bend and loop DNA to perform its biological functions.[14][15] These proteins are important in bending arrays of nucleosomes and arranging them into the larger structures that form chromosomes.[16]
Proteins that specifically bind single-stranded DNA
A distinct group of DNA-binding proteins are the DNA-binding proteins that specifically bind single-stranded DNA. In humans, replication protein A is the best-understood member of this family and is used in processes where the double helix is separated, including DNA replication, recombination and DNA repair.[17] These binding proteins seem to stabilize single-stranded DNA and protect it from forming stem-loops or being degraded by nucleases.
Binding to specific DNA sequences
In contrast, other proteins have evolved to bind to specific DNA sequences. The most intensively studied of these are the various transcription factors, which are proteins that regulate transcription. Each transcription factor binds to one specific set of DNA sequences and activates or inhibits the transcription of genes that have these sequences near their promoters. The transcription factors do this in two ways. Firstly, they can bind the RNA polymerase responsible for transcription, either directly or through other mediator proteins; this locates the polymerase at the promoter and allows it to begin transcription.[18] Alternatively, transcription factors can bind enzymes that modify the histones at the promoter. This alters the accessibility of the DNA template to the polymerase.[19]
These DNA targets can occur throughout an organism's genome. Thus, changes in the activity of one type of transcription factor can affect thousands of genes.[20] Thus, these proteins are often the targets of the signal transduction processes that control responses to environmental changes or cellular differentiation and development. The specificity of these transcription factors' interactions with DNA come from the proteins making multiple contacts to the edges of the DNA bases, allowing them to read the DNA sequence. Most of these base-interactions are made in the major groove, where the bases are most accessible.[21] Mathematical descriptions of protein-DNA binding taking into account sequence-specificity, and competitive and cooperative binding of proteins of different types are usually performed with the help of the lattice models.[22] Computational methods to identify the DNA binding sequence specificity have been proposed to make a good use of the abundant sequence data in the post-genomic era.[23]
Protein–DNA interactions
Protein–DNA interactions occur when a protein binds a molecule of DNA, often to regulate the biological function of DNA, usually the expression of a gene. Among the proteins that bind to DNA are transcription factors that activate or repress gene expression by binding to DNA motifs and histones that form part of the structure of DNA and bind to it less specifically. Also proteins that repair DNA such as uracil-DNA glycosylase interact closely with it.
In general, proteins bind to DNA in the major groove; however, there are exceptions.[24] Protein–DNA interaction are of mainly two types, either specific interaction, or non-specific interaction. Recent single-molecule experiments showed that DNA binding proteins undergo of rapid rebinding in order to bind in correct orientation for recognizing the target site.[25]
Design
Designing DNA-binding proteins that have a specified DNA-binding site has been an important goal for biotechnology. Zinc finger proteins have been designed to bind to specific DNA sequences and this is the basis of zinc finger nucleases. Recently transcription activator-like effector nucleases (TALENs) have been created which are based on natural proteins secreted by Xanthomonas bacteria via their type III secretion system when they infect various plant species.[26]
Detection methods
There are many in vitro and in vivo techniques which are useful in detecting DNA-Protein Interactions. The following lists some methods currently in use:[27]
- Electrophoretic mobility shift assay is a widespread technique to identify protein–DNA interactions.
- DNase footprinting assay can be used to identify the specific site of binding of a protein to DNA.
- Chromatin immunoprecipitation is used to identify the sequence of the DNA fragments which bind to a known transcription factor. This technique when combined with high throughput sequencing is known as ChIP-Seq and when combined with microarrays it is known as ChIP-chip.
- Yeast One-hybrid System (Y1H) is used to identify which protein binds to a particular DNA fragment.
- Bacterial one-hybrid system (B1H) is used to identify which protein binds to a particular DNA fragment.
- Structure determination using X-ray crystallography has been used to give a highly detailed atomic view of protein–DNA interactions.
Manipulating the interactions
The protein–DNA interactions can be modulated using stimuli like ionic strength of the buffer, macromolecular crowding,[28] temperature, pH and electric field. This can lead to reversible dissociation/association of the protein–DNA complex.[29][30]
See also
- bZIP domain
- ChIP-exo
- Comparison of nucleic acid simulation software
- DNA-binding domain
- Helix-loop-helix
- Helix-turn-helix
- HMG-box
- Leucine zipper
- Lexitropsin (a semi-synthetic DNA-binding ligand)
- Deoxyribonucleoprotein
- Protein–DNA interaction site prediction software
- RNA-binding protein
- Single-strand binding protein
- Zinc finger
References
- ^ Created from PDB 1LMB
- ^ Created from PDB 1RVA
- ^ Travers, A. A. (1993). DNA-protein interactions. London: Springer. ISBN 978-0-412-25990-6.
- ^ Pabo CO, Sauer RT (1984). "Protein-DNA recognition". Annu. Rev. Biochem. 53 (1): 293–321. doi:10.1146/annurev.bi.53.070184.001453. PMID 6236744.
- ^ Dickerson R.E. (1983). "The DNA helix and how it is read". Sci Am. 249 (6): 94–111. Bibcode:1983SciAm.249f..94D. doi:10.1038/scientificamerican1283-94.
- ^ Zimmer C, Wähnert U (1986). "Nonintercalating DNA-binding ligands: specificity of the interaction and their use as tools in biophysical, biochemical and biological investigations of the genetic material". Prog. Biophys. Mol. Biol. 47 (1): 31–112. doi:10.1016/0079-6107(86)90005-2. PMID 2422697.
- ^ Dervan PB (April 1986). "Design of sequence-specific DNA-binding molecules". Science. 232 (4749): 464–71. Bibcode:1986Sci...232..464D. doi:10.1126/science.2421408. PMID 2421408.
- ^ Sandman K, Pereira S, Reeve J (1998). "Diversity of prokaryotic chromosomal proteins and the origin of the nucleosome". Cell Mol Life Sci. 54 (12): 1350–64. doi:10.1007/s000180050259. PMID 9893710.
- ^ Dame RT (2005). "The role of nucleoid-associated proteins in the organization and compaction of bacterial chromatin". Mol. Microbiol. 56 (4): 858–70. doi:10.1111/j.1365-2958.2005.04598.x. PMID 15853876.
- ^ Luger K, Mäder A, Richmond R, Sargent D, Richmond T (1997). "Crystal structure of the nucleosome core particle at 2.8 A resolution". Nature. 389 (6648): 251–60. Bibcode:1997Natur.389..251L. doi:10.1038/38444. PMID 9305837.
- ^ Jenuwein T, Allis C (2001). "Translating the histone code". Science. 293 (5532): 1074–80. CiteSeerX 10.1.1.453.900. doi:10.1126/science.1063127. PMID 11498575.
- ^ Ito T (2003). Nucleosome assembly and remodelling. Current Topics in Microbiology and Immunology. Vol. 274. pp. 1–22. doi:10.1007/978-3-642-55747-7_1. ISBN 978-3-642-62909-9. PMID 12596902.
{{cite book}}:|journal=ignored (help) - ^ Thomas J (2001). "HMG1 and 2: architectural DNA-binding proteins". Biochem Soc Trans. 29 (Pt 4): 395–401. doi:10.1042/BST0290395. PMID 11497996.
- ^ Murugesapillai, Divakaran; McCauley, Micah J.; Huo, Ran; Nelson Holte, Molly H.; Stepanyants, Armen; Maher, L. James; Israeloff, Nathan E.; Williams, Mark C. (2014). "DNA bridging and looping by HMO1 provides a mechanism for stabilizing nucleosome-free chromatin". Nucleic Acids Research. 42 (14): 8996–9004. doi:10.1093/nar/gku635. PMC 4132745. PMID 25063301.
- ^ Murugesapillai, Divakaran; McCauley, Micah J.; Maher, L. James; Williams, Mark C. (2017). "Single-molecule studies of high-mobility group B architectural DNA bending proteins". Biophysical Reviews. 9: 17–40. doi:10.1007/s12551-016-0236-4. PMID 28303166.
- ^ Grosschedl R, Giese K, Pagel J (1994). "HMG domain proteins: architectural elements in the assembly of nucleoprotein structures". Trends Genet. 10 (3): 94–100. doi:10.1016/0168-9525(94)90232-1. PMID 8178371.
- ^ Iftode C, Daniely Y, Borowiec J (1999). "Replication protein A (RPA): the eukaryotic SSB". Crit Rev Biochem Mol Biol. 34 (3): 141–80. doi:10.1080/10409239991209255. PMID 10473346.
- ^ Myers L, Kornberg R (2000). "Mediator of transcriptional regulation". Annu Rev Biochem. 69 (1): 729–49. doi:10.1146/annurev.biochem.69.1.729. PMID 10966474.
- ^ Spiegelman B, Heinrich R (2004). "Biological control throughs regulated transcriptional coactivators". Cell. 119 (2): 157–67. doi:10.1016/j.cell.2004.09.037. PMID 15479634.
- ^ Li Z, Van Calcar S, Qu C, Cavenee W, Zhang M, Ren B (2003). "A global transcriptional regulatory role for c-Myc in Burkitt's lymphoma cells". Proc Natl Acad Sci USA. 100 (14): 8164–9. Bibcode:2003PNAS..100.8164L. doi:10.1073/pnas.1332764100. PMC 166200. PMID 12808131.
- ^ Pabo C, Sauer R (1984). "Protein-DNA recognition". Annu Rev Biochem. 53 (1): 293–321. doi:10.1146/annurev.bi.53.070184.001453. PMID 6236744.
- ^ Teif V.B.; Rippe K. (2010). "Statistical-mechanical lattice models for protein-DNA binding in chromatin". Journal of Physics: Condensed Matter. 22 (41): 414105. arXiv:1004.5514. Bibcode:2010JPCM...22O4105T. doi:10.1088/0953-8984/22/41/414105. PMID 21386588.
- ^ Wong KC, Chan TM, Peng C., Li Y., and Zhang Z. "DNA Motif Elucidation using belief propagation" Nucleic Acids Research Advanced Online June 2013; doi:10.1093/nar/gkt574 PMID 23814189
- ^ Bewley CA, Gronenborn AM, Clore GM (1998). "Minor groove-binding architectural proteins: structure, function, and DNA recognition". Annu Rev Biophys Biomol Struct. 27: 105–31. doi:10.1146/annurev.biophys.27.1.105. PMC 4781445. PMID 9646864.
- ^ Ganji, Mahipal; Docter, Margreet; Le Grice, Stuart F. J.; Abbondanzieri, Elio A. (2016-09-30). "DNA binding proteins explore multiple local configurations during docking via rapid rebinding". Nucleic Acids Research. 44 (17): 8376–8384. doi:10.1093/nar/gkw666. ISSN 0305-1048. PMC 5041478. PMID 27471033.
- ^ Clark KJ, Voytas DF, Ekker SC (September 2011). "A TALE of two nucleases: gene targeting for the masses?". Zebrafish. 8 (3): 147–9. doi:10.1089/zeb.2011.9993. PMC 3174730. PMID 21929364.
- ^ Cai YH, Huang H (July 2012). "Advances in the study of protein–DNA interaction". Amino Acids. 43 (3): 1141–6. doi:10.1007/s00726-012-1377-9. PMID 22842750.
- ^ Ganji, Mahipal; Docter, Margreet; Grice, Stuart F.J. Le; Abbondanzieri, Elio A. (2016-09-30). "DNA binding proteins explore multiple local configurations during docking via rapid rebinding". Nucleic Acids Research. 44 (17): 8376–8384. doi:10.1093/nar/gkw666. ISSN 0305-1048. PMC 5041478. PMID 27471033.
- ^ Hianik, T. and Wang, J. (2009), Electrochemical Aptasensors – Recent Achievements and Perspectives. Electroanalysis, 21: 1223–1235. doi:10.1002/elan.200904566
- ^ Gosai, A. et al. (2016) Electrical Stimulus Controlled Binding/Unbinding of Human Thrombin-Aptamer Complex. Sci. Rep. 6, 37449; doi:10.1038/srep37449 PMC 5118750
External links
- Protein-DNA binding: data, tools & models (annotated list, constantly updated)
- Abalone tool for modeling DNA-ligand interactions.
- DBD database of predicted transcription factors Uses a curated set of DNA-binding domains to predict transcription factors in all completely sequenced genomes
- DNA-Binding+Proteins at the U.S. National Library of Medicine Medical Subject Headings (MeSH)
