Haplogroup A (Y-DNA)
| Haplogroup A | |
|---|---|
| Possible time of origin | 237,000YBP - 581,000 YBP [1] |
| Possible place of origin | Central-Northwest Africa[2] |
| Ancestor | Human Y-MRCA |
| Descendants | A0, A1 |
| Highest frequencies | Namibia (Tsumkwe San, Nama) 60-70% Southern Sudan (Dinka, Shilluk, Nuer) 33%-61.5% Ethiopia (Beta Israel ) 41%-46% |
In human genetics, Haplogroup A is the lineage of all human males. Because of a mistake in phylogeny, it used to refer to a group of y-chromosome lineages that were later found to be among the first to branch off from the root of the human y-chromosome phylogeny, and are now referred to as "haplogroup A proper" or "haplogroup A(xBT)". No mutations define Haplogroup A, but since this nomenclature only deals with Homo sapiens sapiens, the "Y-Chromosomal Adam" can be considered its founder.
Origin

Many proposals for haplogroup A's origin suggest it was associated with the ancestral population of Southern Africa's hunter-gatherers. This is because Haplogroup A lineages are frequent among the San people. In addition, the most basal mitochondrial DNA lineages are also largely restricted to the San.
However the A lineages of Southern Africa are sub-clades of A lineages found in other parts of Africa. This suggests that A lineages arrived in Southern Africa from elsewhere.[3] The two most basal lineages of Haplogroup A, A0 and A1, have been detected in West Africa, Northwest Africa and Central Africa. Cruciani et al. suggest that these lineages may have emerged somewhere in between Central and Northwest Africa, though such an interpretation is still preliminary due to the incomplete geographic coverage of African y-chromosomes.[2]
Initial studies reported that Haplogroup A lineages emerged around 60,000 years ago which was significantly more recent than TMRCA for mitochondrial DNA lineages which coalesce to between 150-200kya. But Cruciani et al. 2011 pushed back the root of the Y-chromosome tree to 142,000 years ago.[2]
In November 2012, a new study by Scozzari et al. reinforced "the hypothesis of an origin in the north-western quadrant of the African continent for the A1b haplogroup, and, together with recent findings of ancient Y-lineages in central-western Africa, provide new evidence regarding the geographical origin of human MSY diversity".[4]
Distribution
Haplogroup A(xBT) is largely restricted to parts of Africa, though a handful of cases have been reported in Europe and Western Asia. The clade achieves its highest modern frequencies in the Bushmen hunter-gatherer populations of Southern Africa, followed closely by many Nilotic groups in Eastern Africa. However, haplogroup A's oldest sub-clades are exclusively found in Central-Northwest Africa, where it, and consequently Y-chromosomal Adam, is believed to have originated about 140,000 years ago.[2] The clade has also been observed at notable frequencies in certain populations in Ethiopia, as well as some Pygmy groups in Central Africa.
Haplogroup A is less common among Niger–Congo speakers, who largely belong to the E1b1a clade. Haplogroup E in general is believed to have originated in Northeast Africa,[5] and was later introduced to West Africa from where it spread around 5,000 years ago to Central, Southern and Southeastern Africa with the Bantu expansion.[6][7] According to Wood et al. (2005) and Rosa et al. (2007), such relatively recent population movements from West Africa changed the pre-existing population Y chromosomal diversity in Central, Southern and Southeastern Africa, replacing the previous haplogroups in these areas with the now dominant E1b1a lineages. Traces of ancestral inhabitants, however, can be observed today in these regions via the presence of the Y DNA haplogroups A-M91 and B-M60 that are common in certain relict populations, such as the Mbuti Pygmies and the Khoisan.[8][9][10]
| Africa | ||
| Study population | Freq. (in %) | |
| [9] | Tsumkwe San (Namibia) | 66% |
| [9] | Nama (Namibia) | 64 |
| [11] | Dinka (Sudan) | 62 |
| [11] | Shilluk (Sudan) | 53 |
| [11] | Nuba (Sudan) | 46 |
| [12] | Khoisan | 44 |
| [13][14] | Ethiopian Jews | 41 |
| [9][13] | !Kung/Sekele | ~40 |
| [11] | Borgu (Sudan) | 35 |
| [11] | Nuer (Sudan) | 33 |
| [11] | Fur (Sudan) | 31 |
| [9] | Maasai (Kenya) | 27 |
| [15] | Nara (Eritrea) | 20 |
| [11] | Masalit (Sudan) | 19 |
| [9][16] | Amhara (Ethiopia) | ~16 |
| [12] | Ethiopians | 14 |
| [17] | Bantu (Kenya) | 14 |
| [9] | Mandara (Cameroon) | 14 |
| [11] | Hausa (Sudan) | 13 |
| [13] | Khwe (South Africa) | 12 |
| [13] | Fulbe (Cameroon) | 12 |
| [9] | Dama (Namibia) | 11 |
| [16] | Oromo (Ethiopia) | 10 |
| [15] | Kunama (Eritrea) | 10 |
| [9] | South Semitic (Ethiopia) | 10 |
| [17] | Arabs (Egypt) | 3 |
In a composite sample of 3551 African men, Haplogroup A had a frequency of 5.4%.[18] The highest frequencies of haplogroup A have been reported among the Khoisan of Southern Africa, Beta Israel, and Nilo-Saharans from Sudan.
Africa —Central
Haplogroup A3b2-M13 has been observed in populations of northern Cameroon (2/9 = 22% Tupuri,[9] 4/28 = 14% Mandara,[9] 2/17 = 12% Fulbe[13]) and eastern DRC (2/9 = 22% Alur,[9] 1/18 = 6% Hema,[9] 1/47 = 2% Mbuti[9]).
Haplogroup A-M91(xA1a-M31, A2-M6/M14/P3/P4, A3-M32) has been observed in the Bakola people of southern Cameroon (3/33 = 9%).[9]
Without testing for any subclade, haplogroup A Y-DNA has been observed in samples of several populations of Gabon, including 9% (3/33) of a sample of Baka, 3% (1/36) of a sample of Ndumu, 2% (1/46) of a sample of Duma, 2% (1/57) of a sample of Nzebi, and 2% (1/60) of a sample of Tsogo.[7]
Africa —Eastern
Haplogroup A3b2-M13 is common among the Southern Sudanese (53%),[11] especially the Dinka Sudanese (61.5%).[19] Haplogroup A3b2-M13 also has been observed in another sample of a South Sudanese population at a frequency of 45% (18/40), including 1/40 A3b2a-M171.[12] Haplogroup A also has been reported in 14.6% (7/48) of an Amhara sample,[16] 10.3% (8/78) of an Oromo sample,[16] 13.6% (12/88) of another sample from Ethiopia,[12] and 41% of a sample of the Beta Israel (Cruciani et al. 2002), and important percentages are also shared by Bantus in Kenya (14%, Luis et al. 2004) and Iraqw in Tanzania (3/43 = 7.0% (Luis et al. 2004) to 1/6 = 17% (Knight et al. 2003)).
Africa —Northern
The subclade A1 has been observed in Moroccan Berbers, while the subclade A3b2 has been observed in approximately 3% of Egyptian males.
Africa —Southern
One study has found haplogroup A in samples of various Khoisan-speaking tribes with frequency ranging from 10% to 70%.[9] Surprisingly, this particular haplogroup was not found in a sample of the Hadzabe from Tanzania, a population traditionally considered an ancient remnant of Khoisans due to the presence of click consonants in their language.
Eurasia
Haplogroup A has been observed as A1 in European men in England. As A3b2, it has been observed with low frequency in Asia Minor, the Middle East, and some Mediterranean islands, among Aegean Turks, Sardinians, Palestinians, Jordanians, Yemenites, and Omanis. Without testing for any subclade, haplogroup A has been observed in a sample of Greeks from Mitilini on the Aegean island of Lesvos[20] and in samples of Portuguese from southern Portugal, central Portugal, and Madeira.[21] The authors of one study have reported finding what appears to be haplogroup A in 3.1% (2/65) of a sample of Cypriots,[22] though they have not definitively excluded the possibility that either of these individuals may belong to haplogroup B or haplogroup C.
Subclades

A0-P305
A0 or A1b-P305 is found only in Bakola Pygmies (South Cameroon) at 8.3% and Berbers from Algeria at 1.5%.[2] Also found in Ghana.[4]
A1a-M31
The subclade A1a-M31 has been found in approximately 2.8% (8/282) of a pool of seven samples of various ethnic groups in Guinea-Bissau, especially among the Papel-Manjaco-Mancanha (5/64 = 7.8%).[8] In an earlier study published in 2003, Gonçalves et al. have reported finding A1a-M31 in 5.1% (14/276) of a sample from Guinea-Bissau and in 0.5% (1/201) of a pair of samples from Cabo Verde.[23] The authors of another study have reported finding haplogroup A1a-M31 in 5% (2/39) of a sample of Mandinka from Senegambia and 2% (1/55) of a sample of Dogon from Mali.[9] Haplogroup A1a-M31 also has been found in 3% (2/64) of a sample of Berbers from Morocco[13] and 2.3% (1/44) of a sample of unspecified ethnic affiliation from Mali.[12]
In 2007, seven men from Yorkshire, England sharing a distinctive surname were identified as being from the A1a-M31 subgroup of haplogroup A. It was discovered that these men had a common male-line ancestor from the 18th century, but no previous information about African ancestry was known. [18]
A-M6
The subclade A1b1a1a-M6 (formerly A2) is typically found among Khoisan peoples. The authors of one study have reported finding haplogroup A-M6(xA-P28) in 28% (8/29) of a sample of Tsumkwe San and 16% (5/32) of a sample of !Kung/Sekele, and haplogroup A2b-P28 in 17% (5/29) of a sample of Tsumkwe San, 9% (3/32) of a sample of !Kung/Sekele, 9% (1/11) of a sample of Nama, and 6% (1/18) of a sample of Dama.[9] The authors of another study have reported finding haplogroup A2 in 15.4% (6/39) of a sample of Khoisan males, including 5/39 A2-M6/M14/M23/M29/M49/M71/M135/M141(xA2a-M114) and 1/39 A2a-M114.[12]
A-M32
The clade A1b1b-M32 (formerly A3) contains the most populous branches of haplogroup A and is mainly found in Eastern Africa and Southern Africa.
A-M28
The subclade A1b1b1-M28 (formerly A3a) has only been rarely observed in the Horn of Africa. In 5% (1/20) of a mixed sample of speakers of South Semitic languages from Ethiopia,[9] 1.1% (1/88) of a sample of Ethiopians,[12] and 0.5% (1/201) in Somalis.[24]
A-M51
The subclade A1b1b2a-M51 (formerly A3b1) occurs most frequently among Khoisan peoples (6/11 = 55% Nama,[9] 11/39 = 28% Khoisan,[12] 7/32 = 22% !Kung/Sekele,[9] 6/29 = 21% Tsumkwe San,[9] 1/18 = 6% Dama[9]). However, it also has been found with lower frequency among Bantu peoples of Southern Africa, including 2/28 = 7% Sotho–Tswana,[9] 3/53 = 6% non-Khoisan Southern Africans,[12] 4/80 = 5% Xhosa,[9] and 1/29 = 3% Zulu.[9]
A-M13
The subclade A1b1b2b-M13 (formerly A3b2) that is commonly found in East Africa and northern Cameroon is different from those found in the Khoisan samples and only remotely related to them (it is actually only one of many subclades within haplogroup A). This finding suggests an ancient divergence.
In Sudan, haplogroup A-M13 has been found in 28/53 = 52.8% of Southern Sudanese, 13/28 = 46.4% of the Nuba of central Sudan, 25/90 = 27.8% of Western Sudanese, 4/32 = 12.5% of local Hausa people, and 5/216 = 2.3% of Northern Sudanese.[25]
In Ethiopia, one study has reported finding haplogroup A-M13 in 14.6% (7/48) of a sample of Amhara and 10.3% (8/78) of a sample of Oromo.[16] Another study has reported finding haplogroup A3b2b-M118 in 6.8% (6/88) and haplogroup A3b2*-M13(xA3b2a-M171, A3b2b-M118) in 5.7% (5/88) of a mixed sample of Ethiopians, amounting to a total of 12.5% (11/88) A3b2-M13.[12]
Haplogroup A-M13 also has been observed occasionally outside of Central and Eastern Africa, as in the Aegean Region of Turkey (2/30 = 6.7%[26]), Yemenite Jews (1/20 = 5%[14]), Egypt (4/147 = 2.7%,[17] 3/92 = 3.3%[9]), Palestinian Arabs (2/143 = 1.4%[27]), Sardinia (1/77 = 1.3%,[28] 1/22 = 4.5%[12]), the capital of Jordan, Amman (1/101=1%[29]), and Oman (1/121 = 0.8%[17]).
Phylogenetics
Phylogenetic History
Prior to 2002, there were in academic literature at least seven naming systems for the Y-Chromosome Phylogenetic tree. This lead to considerable confusion. In 2002, the major research groups came together and formed the Y-Chromosome Consortium (YCC). They published a joint paper that created a single new tree that all agreed to use. Latter, a group of citizen scientists with an interest in population genetics and genetic genealogy formed a working group to create an amateur tree aiming at being above all timely. The table below brings together all of these works at the point of the landmark 2002 YCC Tree. This allows a researcher reviewing older published literature to quickly move between nomenclatures.
| YCC 2002/2008 (Shorthand) | (α) | (β) | (γ) | (δ) | (ε) | (ζ) | (η) | YCC 2002 (Longhand) | YCC 2005 (Longhand) | YCC 2008 (Longhand) | YCC 2010r (Longhand) | ISOGG 2006 | ISOGG 2007 | ISOGG 2008 | ISOGG 2009 | ISOGG 2010 | ISOGG 2011 | ISOGG 2012 |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| A-M31 | 7 | I | 1A | 1 | – | H1 | A | A1 | A1 | A1 | A1a | A1 | A1 | A1a | A1a | A1a | A1a | A1a |
| A-M6 | 27 | I | 2 | 3 | – | H1 | A | A2* | A2 | A2 | A2 | A2 | A2 | A2 | A2 | A2 | A2 | A1b1a1a |
| A-M114 | 27 | I | 2 | 3 | – | H1 | A | A2a | A2a | A2a | A2a | A2a | A2a | A2a | A2a | A2a | A2a | A1b1a1a1a |
| A-P28 | 27 | I | 2 | 4 | – | H1 | A | A2b | A2b | A2b | A2b | A2b | A2b | A2b | A2b | A2b | A2b | A1b1a1a1b |
| A-M32 | * | * | * | * | * | * | * | * | A3 | A3 | A3 | A3 | A3 | A3 | A3 | A3 | A3 | A1b1b |
| A-M28 | 7 | I | 1A | 1 | – | H1 | A | A3a | A3a | A3a | A3a | A3a | A3a | A3a | A3a | A3a | A3a | A1b1b1 |
| A-M51 | 7 | I | 1A | 1 | – | H1 | A | A3b1 | A3b1 | A3b1 | A3b1 | A3b1 | A3b1 | A3b1 | A3b1 | A3b1 | A3b1 | A1b1b2a |
| A-M13 | 7 | I | 1A | 2 | Eu1 | H1 | A | A3b2* | A3b2 | A3b2 | A3b2 | A3b2 | A3b2 | A3b2 | A3b2 | A3b2 | A3b2 | A1b1b2b |
| A-M171 | 7 | I | 1A | 2 | Eu1 | H1 | A | A3b2a | A3b2a | A3b2a | A3b2a | A3b2a | A3b2a | A3b2a | A3b2a | A3b2a | A3b2a | removed |
| A-M118 | 7 | I | 1A | 2 | Eu1 | H1 | A | A3b2b | A3b2b | A3b2b | A3b2b | A3b2b | A3b2b | A3b2b | A3b2b | A3b2b | A3b2b | A1b1b2b1 |
Original Research Publications
The following research teams per their publications were represented in the creation of the YCC Tree.
Cruciani 2011
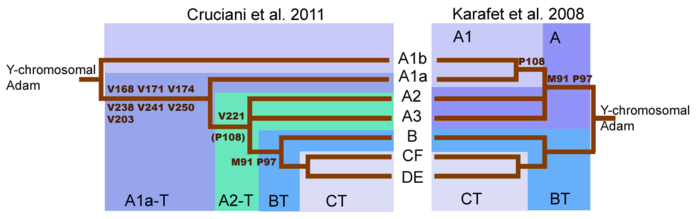
A major shift in understanding of the Haplogroup A tree came with the publication of (Cruciani 2011). Initial sequencing of the human y-chromosome suggested that first split in the Y-Chromosome family tree occurred with the M91 mutation that separated Haplogroup A from Haplogroup BT.[30] However, it is now known that deepest split in the Y-chromosome tree is found between two previously reported subclades of Haplogroup A, rather than between Haplogroup A and Haplogroup BT. Subclades A1b and A1a-T now descend directly from the root of the tree. The rearrangement of the Y-chromosome family tree implies that lineages classified as Haplogroup A do not necessarily form a monophyletic clade.[2] Haplogroup A therefore refers to a collection of lineages that do not possess the markers that define Haplogroup BT, though many lineages within haplogroup A are only very distantly related.
The M91 and P97 mutations distinguish Haplogroup A from Haplogroup BT . Within Haplogroup A chromosomes, the M91 marker consists of a stretch of 8 T nucleobase units. In Haplogroup BT and chimpanzee chromosomes, this marker consists of 9 T nucleobase units. This pattern suggested that the 9T stretch of Haplogroup BT was the ancestral version and that Haplogroup A was formed by the deletion of one nucleobase.[30][2]
But according to Cruciani et al. 2011, the region surrounding the M91 marker is a mutational hotspot prone to recurrent mutations. It is therefore possible that the 8T stretch of Haplogroup A may be the ancestral state of M91 and the 9T of Haplogroup BT may be the derived state that arose by an insertion of 1T. This would explain why subclades A1b and A1a-T, the deepest branches of Haplogroup A, both possess the 8T stretch. Furthermore Cruciani et al. 2011 determined that the P97 marker, which is also used to identify haplogroup A, possessed the ancestral state in haplogroup A but the derived state in Haplogroup BT.[2]
Phylogenetic Trees
This phylogenetic tree of haplogroup subclades is based on the Y-Chromosome Consortium (YCC) Tree,[31] the ISOGG Y-DNA Haplogroup Tree,[6] and subsequent published research.
Y-chromosomal Adam
- A0 (formerly A1b) (P305, V148, V149, V154, V164, V166, V172, V173, V177, V190, V196, V223, V225, V229, V233, V239)
- A1 (A1a-T according Cruciani 2011) (L985, L989, L990, L1002, L1003, L1004, L1009, L1013, L1053, V161, V168, V171, V174, V203, V238, V241, V250, V238, V241, V250)
- A1a (M31, P82, V4, V14, V15, V25, V26, V28, V30, V40, V48, V53, V57, V58, V63, V76, V191, V201, V204, V214, V215, V236)
- A1b (A2-T according Cruciani 2011) (P108, V221)
- A1b1 (L419)
- A1b1a (V50, V82, V198, V224)
- A1b1a1 formerly A2 (M14, M23, L968/M29/P3/PN3, M71, M135, M141, M206, M276/P247, M277/P248, MEH1, P4, P5, P36.1, Page71, Page87, Page95)
- A1b1a1a (M6, M196)
- A1b1a1a1 (M212)
- A1b1a1a1a formerly A2a (M114)
- A1b1a1a1b formerly A2b (P28)
- A1b1a1a1c formerly A2c (P262)
- A1b1a1a1 (M212)
- A1b1a1a (M6, M196)
- A1b1a1 formerly A2 (M14, M23, L968/M29/P3/PN3, M71, M135, M141, M206, M276/P247, M277/P248, MEH1, P4, P5, P36.1, Page71, Page87, Page95)
- A1b1b formerly A3 (M32)
- A1b1b1 formerly A3a (M28, M59)
- A1b1b2 formerly A3b (M144, M190, M220, P289)
- A1b1b2a formerly A3b1 (M51, P100, P291)
- A1b1b2a1 formerly A3b1a (P71, P102)
- A1b1b2b formerly A3b2 (M13, M127, M202, M219, M305):
- A1b1b2b1 (M118)
- A1b1b2a formerly A3b1 (M51, P100, P291)
- A1b1a (V50, V82, V198, V224)
- BT (M42, M94, M139, M299, M60, M181/Page32, P85, P90, P97, Page65.1/SRY1532.1/SRY10831.1, V21, V29, V31, V59, V64, V102, V187, V202, V216, V235)
- A1b1 (L419)
See also
Genetics
Y-DNA A Subclades
Y-DNA Backbone Tree
References
- ^ http://www.sciencedirect.com/science/article/pii/S0002929713000736
- ^ a b c d e f g h Cruciani, Fulvio (19 May 2011). "A Revised Root for the Human Y Chromosomal Phylogenetic Tree: The Origin of Patrilineal Diversity in Africa". 88 (6). doi:10.1016/j.ajhg.2011.05.002. PMC 3113241. PMID 21601174. Retrieved 11 June 2011.
{{cite journal}}: Cite journal requires|journal=(help); Unknown parameter|coauthors=ignored (|author=suggested) (help) - ^ Batini; et al. (2011). "Signatures of the pre-agricultural peopling processes in sub-Saharan Africa as revealed by the phylogeography of early Y chromosome lineages" (PDF). doi:10.1093/molbev/msr089.
{{cite journal}}: Cite journal requires|journal=(help); Explicit use of et al. in:|last=(help) - ^ a b Scozzari R, Massaia A, D’Atanasio E, Myres NM, Perego UA, et al. (2012) Molecular Dissection of the Basal Clades in the Human Y Chromosome Phylogenetic Tree. PLoS ONE 7(11): e49170. doi:10.1371/journal.pone.0049170
- ^ K. Abu-Amero, Khaled (2009). "Saudi Arabian Y-Chromosome diversity and its relationship with nearby regions". BMC Genetics. 10 (59). doi:10.1186/1471-2156-10-59. PMC 2759955. PMID 19772609.
{{cite journal}}: Unknown parameter|coauthors=ignored (|author=suggested) (help)CS1 maint: unflagged free DOI (link) - ^ a b International Society of Genetic Genealogy. "Y-DNA Haplogroup Tree". Retrieved 2012.
{{cite web}}: Check date values in:|accessdate=(help) - ^ a b Berniell-Lee, Gemma (2010-07-26). "Genetic and Demographic Implications of the Bantu Expansion: Insights from Human Paternal Lineages" (Epub). Molecular Biology and Evolution. 26 (7): 1581–1589. doi:10.1093/molbev/msp069. PMID 19369595. Retrieved 2010-11-30.
{{cite journal}}: Unknown parameter|coauthors=ignored (|author=suggested) (help) - ^ a b Rosa Alexandra, Ornelas Carolina, Jobling Mark A; et al. (2007). "Y-chromosomal diversity in the population of Guinea-Bissau: a multiethnic perspective". BMC Evolutionary Biology. 7: 124. doi:10.1186/1471-2148-7-124. PMC 1976131. PMID 17662131.
{{cite journal}}: Explicit use of et al. in:|author=(help)CS1 maint: multiple names: authors list (link) CS1 maint: unflagged free DOI (link) - ^ a b c d e f g h i j k l m n o p q r s t u v w x y z aa Elizabeth T Wood, Daryn A Stover, Christopher Ehret et al., "Contrasting patterns of Y chromosome and mtDNA variation in Africa: evidence for sex-biased demographic processes," European Journal of Human Genetics (2005) 13, 867–876. (cf. Appendix A: Y Chromosome Haplotype Frequencies)
- ^ Underhill, P.A (January 2001). "The phylogeography of Y chromosome binary haplotypes and the origins of modern human populations". Annals of Human Genetics. 65 (1): 43–62. doi:10.1046/j.1469-1809.2001.6510043.x. PMID 11415522.
{{cite journal}}: Unknown parameter|coauthors=ignored (|author=suggested) (help) - ^ a b c d e f g h i 28/53 (Dinka, Nuer, and Shilluk), Hassan HY, Underhill PA, Cavalli-Sforza LL, Ibrahim ME (2008). "Y-chromosome variation among Sudanese: restricted gene flow, concordance with language, geography, and history" (PDF). Am. J. Phys. Anthropol. 137 (3): 316–23. doi:10.1002/ajpa.20876. PMID 18618658.
{{cite journal}}: Unknown parameter|month=ignored (help)CS1 maint: multiple names: authors list (link) - ^ a b c d e f g h i j k Underhill PA, Shen P, Lin AA; et al. (2000). "Y chromosome sequence variation and the history of human populations". Nat. Genet. 26 (3): 358–61. doi:10.1038/81685. PMID 11062480.
{{cite journal}}: Explicit use of et al. in:|author=(help); Unknown parameter|month=ignored (help)CS1 maint: multiple names: authors list (link) - ^ a b c d e f Cruciani Fulvio, Santolamazza Piero, Shen Peidong; et al. "A Back Migration from Asia to Sub-Saharan Africa Is Supported by High-Resolution Analysis of Human Y-Chromosome Haplotypes". American Journal of Human Genetics. 70 (1197–1214): 2002.
{{cite journal}}: Explicit use of et al. in:|author=(help)CS1 maint: multiple names: authors list (link) - ^ a b Peidong Shen, Tal Lavi, Toomas Kivisild et al., "Reconstruction of Patrilineages and Matrilineages of Samaritans and Other Israeli Populations From Y-Chromosome and Mitochondrial DNA Sequence Variation," Human Mutation 24:248-260 (2004).
- ^ a b Fulvio Cruciani, Beniamino Trombetta, Daniele Sellitto et al., "Human Y chromosome haplogroup R-V88: a paternal genetic record of early mid Holocene trans-Saharan connections and the spread of Chadic languages," European Journal of Human Genetics (2010), 1–8
- ^ a b c d e Semino O, Santachiara-Benerecetti AS, Falaschi F, Cavalli-Sforza LL, Underhill PA (2002). "Ethiopians and Khoisan share the deepest clades of the human Y-chromosome phylogeny". Am. J. Hum. Genet. 70 (1): 265–8. doi:10.1086/338306. PMC 384897. PMID 11719903.
{{cite journal}}: Unknown parameter|month=ignored (help)CS1 maint: multiple names: authors list (link) - ^ a b c d Luis J. R., Rowold D. J., Regueiro M.; et al. "The Levant versus the Horn of Africa: Evidence for Bidirectional Corridors of Human Migrations". American Journal of Human Genetics. 74 (532–544): 2004.
{{cite journal}}: Explicit use of et al. in:|author=(help)CS1 maint: multiple names: authors list (link) - ^ a b King TE, Parkin EJ, Swinfield G; et al. (2007). "Africans in Yorkshire? The deepest-rooting clade of the Y phylogeny within an English genealogy". Eur. J. Hum. Genet. 15 (3): 288–93. doi:10.1038/sj.ejhg.5201771. PMC 2590664. PMID 17245408.
{{cite journal}}: Explicit use of et al. in:|author=(help); Unknown parameter|month=ignored (help)CS1 maint: multiple names: authors list (link)
News article: "Yorkshire clan linked to Africa". BBC News. 2007-01-24. Retrieved 2007-01-27. - ^ 16/26, Hassan et al. 2008
- ^ Giacomo F. Di, Luca F., Anagnou N.; et al. (2003). "Clinal patterns of human Y chromosomal diversity in continental Italy and Greece are dominated by drift and founder effects". Molecular Phylogenetics and Evolution. 28 (3): 387–395. doi:10.1016/S1055-7903(03)00016-2. PMID 12927125.
{{cite journal}}: Explicit use of et al. in:|author=(help)CS1 maint: multiple names: authors list (link) - ^ Gonçalves Rita, Freitas Ana, Branco Marta; et al. (2005). "Y-chromosome Lineages from Portugal, Madeira and Açores Record Elements of Sephardim and Berber Ancestry". Annals of Human Genetics. 69 (Pt 4): 443–454. doi:10.1111/j.1529-8817.2005.00161.x. PMID 15996172.
{{cite journal}}: Explicit use of et al. in:|author=(help)CS1 maint: multiple names: authors list (link) - ^ C. Capelli, N. Redhead, V. Romano et al., "Population Structure in the Mediterranean Basin: A Y Chromosome Perspective," Annals of Human Genetics (2005)
- ^ Rita Gonçalves, Alexandra Rosa, Ana Freitas et al., "Y-chromosome lineages in Cabo Verde Islands witness the diverse geographic origin of its first male settlers," Human Genetics (2003) 113 : 467–472
- ^ Abu-Amero, Khaled (22 September 2009). "Saudi Arabian Y-Chromosome diversity and its relationship with nearby regions". BMC Genetics. 10: 59. doi:10.1186/1471-2156-10-59. PMC 2759955. PMID 19772609.
{{cite journal}}: Unknown parameter|coauthors=ignored (|author=suggested) (help)CS1 maint: unflagged free DOI (link) - ^ Hisham Y. Hassan et al. (2008). "Southern Sudanese" includes 26 Dinka, 15 Shilluk, and 12 Nuer. "Western Sudanese" includes 26 Borgu, 32 Masalit, and 32 Fur. "Northern Sudanese" includes 39 Nubians, 42 Beja, 33 Copts, 50 Gaalien, 28 Meseria, and 24 Arakien.
- ^ Cengiz Cinnioglu, Roy King, Toomas Kivisild et al., "Excavating Y-chromosome haplotype strata in Anatolia," Human Genetics (2004) 114 : 127–148. DOI 10.1007/s00439-003-1031-4
- ^ Nebel Almut, Filon Dvora, Brinkmann Bernd; et al. "The Y Chromosome Pool of Jews as Part of the Genetic Landscape of the Middle East". American Journal of Human Genetics. 69 (1095–1112): 2001.
{{cite journal}}: Explicit use of et al. in:|author=(help)CS1 maint: multiple names: authors list (link) - ^ Ornella Semino, Giuseppe Passarino, Peter J. Oefner et al., "The Genetic Legacy of Paleolithic Homo sapiens sapiens in Extant Europeans: A Y Chromosome Perspective," Science Vol 290 (10 November 2000).
- ^ Flores, C; Maca-Meyer, N; Larruga, JM; Cabrera, VM; Karadsheh, N; Gonzalez, AM; et al. (2005). "Isolates in a corridor of migrations: a high-resolution analysis of Y-chromosome variation in Jordan". J Hum Genet. 50 (9): 435–441. doi:10.1007/s10038-005-0274-4. PMID 16142507.
{{cite journal}}: Explicit use of et al. in:|last=(help) - ^ a b Karafet TM, Mendez FL, Meilerman MB, Underhill PA, Zegura SL, Hammer MF (2008). "New binary polymorphisms reshape and increase resolution of the human Y chromosomal haplogroup tree". Genome Research. 18 (5): 830–8. doi:10.1101/gr.7172008. PMC 2336805. PMID 18385274.
{{cite journal}}: CS1 maint: multiple names: authors list (link) - ^ Krahn, Thomas. "YCC Tree". Houston, Texas: FTDNA. Retrieved 16 May 2011.
