Breast cancer classification
Breast cancer classification divides breast cancer into categories according to different schemes criteria and serving a different purpose. The major categories are the histopathological type, the grade of the tumor, the stage of the tumor, and the expression of proteins and genes. As knowledge of cancer cell biology develops these classifications are updated.
The purpose of classification is to select the best treatment. The effectiveness of a specific treatment is demonstrated for a specific breast cancer (usually by randomized, controlled trials). That treatment may not be effective in a different breast cancer. Some breast cancers are aggressive and life-threatening, and must be treated with aggressive treatments that have major adverse effects. Other breast cancers are less aggressive and can be treated with less aggressive treatments, such as lumpectomy.
Treatment algorithms rely on breast cancer classification to define specific subgroups that are each treated according to the best evidence available. Classification aspects must be carefully tested and validated, such that confounding effects are minimized, making them either true prognostic factors, which estimate disease outcomes such as disease-free or overall survival in the absence of therapy, or true predictive factors, which estimate the likelihood of response or lack of response to a specific treatment.[1]
Classification of breast cancer is usually, but not always, primarily based on the histological appearance of tissue in the tumor. A variant from this approach, defined on the basis of physical exam findings, is that inflammatory breast cancer (IBC), a form of ductal carcinoma or malignant cancer in the ducts, is distinguished from other carcinomas by the inflamed appearance of the affected breast, which correlates with increased cancer aggressivity.[2]
Schemes or aspects[edit]
Overview[edit]
Breast cancers can be classified by different schemata. Each of these aspects influences treatment response and prognosis. Description of a breast cancer would optimally include all of these classification aspects, as well as other findings, such as signs found on physical exam. A full classification includes histopathological type, grade, stage (TNM), receptor status, and the presence or absence of genes as determined by DNA testing:
- Histopathology. Although breast cancer has many different histologies, the considerable majority of breast cancers are derived from the epithelium lining the ducts or lobules, and are classified as mammary ductal carcinoma. Carcinoma in situ is proliferation of cancer cells within the epithelial tissue without invasion of the surrounding tissue. In contrast, invasive carcinoma invades the surrounding tissue.[3] Perineural and/or lymphovascular space invasion is usually considered as part of the histological description of a breast cancer, and when present may be associated with more aggressive disease.
- Grade. Grading focuses on the appearance of the breast cancer cells compared to the appearance of normal breast tissue. Normal cells in an organ like the breast become differentiated, meaning that they take on specific shapes and forms that reflect their function as part of that organ. Cancerous cells lose that differentiation. In cancer, the cells that would normally line up in an orderly way to make up the milk ducts become disorganized. Cell division becomes uncontrolled. Cell nuclei become less uniform. Pathologists describe cells as well differentiated (low-grade), moderately differentiated (intermediate-grade), and poorly differentiated (high-grade) as the cells progressively lose the features seen in normal breast cells. Poorly differentiated cancers have a worse prognosis.
- Stage. The TNM classification for staging breast cancer is based on the size of the cancer where it originally started in the body and the locations to which it has travelled. These cancer characteristics are described as the size of the tumor (T), whether or not the tumor has spread to the lymph nodes (N) in the armpits, neck, and inside the chest, and whether the tumor has metastasized (M) (i.e. spread to a more distant part of the body). Larger size, nodal spread, and metastasis have a larger stage number and a worse prognosis. The main stages are:
- Stage 0 which is in situ disease or Paget's disease of the nipple. Stage 0 is a pre-cancerous or marker condition, either ductal carcinoma in situ (DCIS) or lobular carcinoma in situ (LCIS).
- Stages 1–3 are within the breast or regional lymph nodes.
- Stage 4 is a metastatic cancer. Metastatic breast cancer has a less favorable prognosis.
- Receptor status. Cells have receptors on their surface and in their cytoplasm and nucleus. Chemical messengers such as hormones bind to receptors, and this causes changes in the cell. Breast cancer cells may or may not have many different types of receptors, the three most important in the present classification being: estrogen receptor (ER), progesterone receptor (PR), and HER2/neu. Cells with or without these receptors are called ER positive (ER+), ER negative (ER-), PR positive (PR+), PR negative (PR-), HER2 positive (HER2+), and HER2 negative (HER2-). Cells with none of these receptors are called basal-like or triple negative. HER2-low has some HER2 proteins on the cell surface, but not enough to be classified as HER2-positive.[4] Trastuzumab deruxtecan is the first approved therapy by the US Food and Drug Administration (FDA) targeted to people with the HER2-low breast cancer subtype.[4]
- DNA-based classification. Understanding the specific details of a particular breast cancer may include looking at the cancer cell DNA or RNA by several different laboratory approaches. When specific DNA mutations or gene expression profiles are identified in the cancer cells this may guide the selection of treatments, either by targeting these changes, or by predicting from these alterations which non-targeted therapies are most effective.
- Other classification approaches.
- Computer models such as Adjuvant can combine the various classification aspects according to validated algorithms and present visually appealing graphics that assist in treatment decisions.
- The USC/Van Nuys prognostic index (VNPI) classifies ductal carcinoma in situ (DCIS) into dissimilar risk categories that may be treated accordingly.
- The choice of which treatment to receive can be substantially influenced by comorbidity assessments.
- Familial breast cancers may potentially undergo dissimilar treatment (such as mastectomy).
Histopathology[edit]


Histopathologic classification is based upon characteristics seen upon light microscopy of biopsy specimens. They can broadly be classified into:
- Carcinoma in situ . This group constitutes about 15-30% of breast biopsies, more so in countries with high coverage of breast screening programs.[6] These have favorable prognosis, with 5-year survival rates of 97-99%.[7]
- Invasive carcinoma. This group constitutes the other 70-85%.[6] The most common type in this group is invasive ductal carcinoma, representing about 80% of invasive carcinomas.[6] In the US, 55% of breast cancers are invasive ductal carcinoma.[8] Invasive lobular carcinoma represent about 10% of invasive carcinomas,[6] and 5% of all breast cancers in the US.[8] The overall 5-year survival rate for both invasive ductal carcinoma and invasive lobular carcinoma was approximately 85% in 2003.[9] Ductal carcinoma in situ, on the other hand, is in itself harmless, although if untreated approximately 60% of these low-grade DCIS lesions will become invasive over the course of 40 years in follow-up.[10]
WHO classification[edit]
The 2012 World Health Organization (WHO) classification of tumors of the breast[11] which includes benign (generally harmless) tumors and malignant (cancerous) tumors, recommends the following pathological types:
|
Invasive breast carcinomas
Classic Solid Mixed Alveolar Tubulolobular Pleomorphic
(Other well-accepted subtypes of metaplastic mammary carcinoma thought to have clinical significance but not included in the decade old WHO classification:
Mesenchymal tumors (including sarcoma)
|
Tumors of the male breast
Malignant lymphoma Metastatic tumors to the breast from other places in the body Precursor lesions
Benign epithelial lesions
Myoepithelial lesions
Fibroepithelial tumours
Benign tumors of the nipple
Malignant tumors of the nipple
|
Grade[edit]
The grading of a cancer in the breast depends on the microscopic similarity of breast cancer cells to normal breast tissue, and classifies the cancer as well differentiated (low-grade), moderately differentiated (intermediate-grade), and poorly differentiated (high-grade), reflecting progressively less normal appearing cells that have a worsening prognosis. Although grading is fundamentally based on how biopsied, cultured cells behave, in practice the grading of a given cancer is derived by assessing the cellular appearance of the tumor. The closer the appearance of the cancer cells to normal cells, the slower their growth and the better the prognosis. If cells are not well differentiated, they will appear immature, will divide more rapidly, and will tend to spread. Well differentiated is given a grade of 1, moderate is grade 2, while poor or undifferentiated is given a higher grade of 3 or 4 (depending upon the scale used).
The Nottingham system[12] is recommended for breast cancer grading.[13] The Nottingham system is also called the Bloom–Richardson–Elston system (BRE),[14] or the Elston-Ellis modification[15] of the Scarff-Bloom-Richardson grading system.[16][17] It grades breast carcinomas by adding up scores for tubule formation, nuclear pleomorphism, and mitotic count, each of which is given 1 to 3 points. The scores for each of these three criteria are then added together to give an overall final score and corresponding grade. It is not applicable to medullary carcinomas which are histologically high-grade by definition, while being clinically low-grade if lymph nodes are negative.[18] It is also not applicable to metaplastic carcinomas.[19]
The grading criteria are as follows:
Tubule formation[edit]

This parameter assesses what percent of the tumor forms normal duct structures. In cancer, there is a breakdown of the mechanisms that cells use to attach to each other and communicate with each other, to form tissues such as ducts, so the tissue structures become less orderly.
Note: The overall appearance of the tumor has to be considered.[20]
- 1 point: tubular formation in more than 75% of the tumor (it may in addition be termed "majority of tumor")
- 2 points: tubular formation in 10 to 75% of the tumor ("moderate")
- 3 points: tubular formation in less than 10% of the tumor ("little or none")
Nuclear pleomorphism[edit]
This parameter assesses whether the cell nuclei are uniform like those in normal breast duct epithelial cells, or whether they are larger, darker, or irregular (pleomorphic). In cancer, the mechanisms that control genes and chromosomes in the nucleus break down, and irregular nuclei and pleomorphic changes are signs of abnormal cell reproduction.
Note: The cancer areas having cells with the greatest cellular abnormalities should be evaluated.
- 1 point: nuclei with minimal or mild variation in size and shape
- 2 points: nuclei with moderate variation in size and shape
- 3 points: nuclei with marked variation in size and shape
-
Ductal carcinoma with mild nuclear pleomorphism.
-
Invasive ductal carcinoma with moderate nuclear pleomorphism.
-
Invasive lobular carcinoma with moderate nuclear pleomorphism.
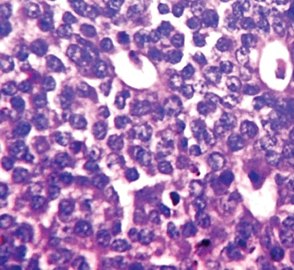
-
Invasive ductal carcinoma with marked nuclear pleomorphism.
Mitotic count[edit]

This parameter assesses how many mitotic figures (dividing cells) the pathologist sees in 10x high power microscope field. One of the hallmarks of cancer is that cells divide uncontrollably. The more cells that are dividing, the worse the cancer.
Note: Mitotic figures are counted only at the periphery of the tumor, and counting should begin in the most mitotically active areas.
| Area per HPF | Score | ||||
|---|---|---|---|---|---|
| 0.096 mm2[notes 1] | 0.12 mm2[notes 1] | 0.16 mm2[notes 1] | 0.27 mm2[notes 1] | 0.31 mm2[notes 1] | |
| 0-3 | 0-4 | 0-5 | 0-9 | 0-11 | 1 |
| 4-7 | 5-8 | 6-10 | 10-19 | 12-22 | 2 |
| >7 | >8 | >10 | >19 | >22 | 3 |
Overall grade[edit]
The scores for each of these three criteria are added together to give a final overall score and a corresponding grade as follows:
- 3-5 Grade 1 tumor (well-differentiated). Best prognosis.
- 6-7 Grade 2 tumor (moderately differentiated). Medium prognosis.
- 8-9 Grade 3 tumor (poorly differentiated). Worst prognosis.
Lower-grade tumors, with a more favorable prognosis, can be treated less aggressively, and have a better survival rate. Higher-grade tumors are treated more aggressively, and their intrinsically worse survival rate may warrant the adverse effects of more aggressive medications.
Stage[edit]
Staging[22] is the process of determining how much cancer there is in the body and where it is located. The underlying purpose of staging is to describe the extent or severity of an individual's cancer, and to bring together cancers that have similar prognosis and treatment.[22] Staging of breast cancer is one aspect of breast cancer classification that assists in making appropriate treatment choices, when considered along with other classification aspects such as estrogen receptor and progesterone receptor levels in the cancer tissue, the human epidermal growth factor receptor 2 (HER2/neu) status, menopausal status, and the person's general health.[23]
Staging information that is obtained prior to surgery, for example by mammography, x-rays and CT scans, is called clinical staging and staging by surgery is known as pathological staging.
Pathologic staging is more accurate than clinical staging, but clinical staging is the first and sometimes the only staging type. For example, if clinical staging reveals stage IV disease, extensive surgery may not be helpful, and (appropriately) incomplete pathological staging information will be obtained.
The American Joint Committee on Cancer (AJCC) and the International Union Against Cancer (UICC) recommend TNM staging, which is a two step procedure. Their TNM system, which they now develop jointly, first classifies cancer by several factors, T for tumor, N for nodes, M for metastasis, and then groups these TNM factors into overall stages.
Primary Tumor (T)[edit]
Tumor – The tumor values (TX, T0, Tis, T1, T2, T3 or T4) depend on the cancer at the primary site of origin in the breast, as follows:[24]
- TX: inability to assess that site
- Tis: ductal carcinoma in situ (DCIS), lobular carcinoma in situ (LCIS) or Paget's disease
- T1: Less than 2 cm
- T1a: 0.1 to 0.5 cm
- T1b: 0.5 to 1.0 cm
- T1c: 1.0 to 2.0 cm
- T2: 2 to 5 cm
- T3: Larger than 5 cm
- T4
- T4a: Chest wall involvement
- T4b: Skin involvement
- T4c: Both 4a and 4b
- T4d: Inflammatory breast cancer, a clinical circumstance where typical skin changes involve at least a third of the breast.
Regional Lymph Nodes (N)[edit]
Lymph Node – The lymph node values (NX, N0, N1, N2 or N3) depend on the number, size and location of breast cancer cell deposits in various regional lymph nodes, such as the armpit (axillary lymph nodes), the collar area (supraclavicular lymph nodes), and inside the chest (internal mammary lymph nodes.)[25][26] The armpit is designated as having three levels: level I is the low axilla, and is below or outside the lower edge of the pectoralis minor muscle; level II is the mid-axilla which is defined by the borders of the pectoralis minor muscle; and level III, or high (apical) axilla which is above the pectoralis minor muscle. Each stage is as follows:[24]
- N0: There is some nuance to the official definitions for N0 disease, which includes:
- N0(i+) : Isolated Tumor Cell clusters (ITC), which are small clusters of cells not greater than 0.2 mm, or single tumor cells, or a cluster of fewer than 200 cells in a single histologic cross-section, whether detected by routine histology or immunohistochemistry.[27]
- N0(mol-): regional lymph nodes have no metastases histologically, but have positive molecular findings (RT-PCR).[27]
- N1: Metastases in 1-3 axillary lymph nodes and/or in internal mammary nodes; and/or in clinically negative internal mammary nodes with micrometastasis, or macrometastasis on sentinel lymph node biopsy.[27]
- N1mi: Micrometastasis, that is, lymph node clusters at least 2 mm or 200 cells, but less than 2.0 mm.[27] At least one carcinoma focus over 2.0 mm is called "Lymph node metastasis". If one node qualifies as metastasis, all other nodes even with smaller foci are counted as metastases as well.
- N2: Fixed/matted ipsilateral axillary nodes.
- N3
- N3a – Ipsilateral infraclavicular nodes
- N3b – Ipsilateral internal mammary nodes
- N3c – Ipsilateral supraclavicular nodes
Distant Metastases (M)[edit]
- M0: No clinical or radiographic evidence of distant metastases
- M0(i+): Molecularly or microscopically detected tumor cells in circulating blood, bone marrow or non-regional nodal tissue, no larger than 0.2 mm, and without clinical or radiographic evidence or symptoms or signs of metastases, and which, perhaps counter-intuitively, does not change the stage grouping, as staging for in M0(i+) is done according to the T and N values
- M1: Distant detectable metastases as determined by classic clinical and radiographic means, and/or metastasis that are histologically larger than 0.2 mm.
Overall stage[edit]
A combination of T, N and M, as follows:[24]
- Stage 0: Tis
- Stage I: T1N0
- Stage II: T2N0, T3N0 T0N1, T1N1, or T2N1
- Stage III: Invasion into skin and/or ribs, matted lymph nodes, T3N1, T0N2, T1N2, T2N2, T3N2, AnyT N3, T4 any N, locally advanced breast cancer
- Stage IV: M1, advanced breast cancer
| Stage | 5-year survival |
|---|---|
| Stage 0 | 100% |
| Stage I | 100% |
| Stage II | 90% |
| Stage III | 70% |
| Stage IV | 30% |
Staging and prognosis[edit]
The impact of different stages on outcome can be appreciated in the following table, taken from patient data in the 2013-2015 period, and using the AJCC 8th edition for staging.[28] It does not show the influence of important additional factors such as estrogen receptor (ER) or HER2/neu receptor status, and does not reflect the impact of newer treatments.
Previous editions[edit]
Although TNM classification is an internationally agreed system, it has gradually evolved through its different editions; the dates of publication and of adoption for use of AJCC editions is summarized in the table in this article; past editions are available from AJCC for web download.[29]
Several factors are important when reviewing reports for individual breast cancers or when reading the medical literature, and applying staging data.
| AJCC edition | published[29] | went into effect[29] | Breast cancer link(s) and page numbers in the original |
|---|---|---|---|
| 7 | 2009 | 2010 | AJCC[30] or NCI[23] |
| 6 | 2002 | 2003 | AJCC;[31] original pages 223-240 |
| 5 | 1997 | 1998 | AJCC;[32] original pages 171-180 |
| 4 | 1992 | 1993 | AJCC;[33] original pages 149-154 |
| 3 | 1988 | 1989 | AJCC;[34] original pages 145-150 |
| 2 | 1983 | 1984 | AJCC;[35] original pages 127-134 |
| 1 | 1977 | 1978 | AJCC;[36] original pages 101-108 |
It is crucial to be aware that the TNM system criteria have varied over time, sometimes fairly substantially, according to the different editions that AJCC and UICC have released.[29] Readers are assisted by the provision in the table of direct links to the breast cancer chapters of these various editions.
As a result, a given stage may have quite a different prognosis depending on which staging edition is used, independent of any changes in diagnostic methods or treatments, an effect that can contribute to "stage migration".[37] For example, differences in the 1998 and 2003 categories resulted in many cancers being assigned differently, with apparent improvement in survival rates.[38]
As a practical matter, reports often use the staging edition that was in place when the study began, rather than the date of acceptance or publication. However, it is worth checking whether the author updated the staging system during the study, or modified the usual classification rules for specific use in the investigation.
A different effect on staging arises from evolving technologies that are used to assign patients to particular categories, such that increasingly sensitive methods tend to cause individual cancers to be reassigned to higher stages, making it improper to compare that cancer's prognosis to the historical expectations for that stage.
Finally, of course, a further important consideration is the effect of improving treatments over time as well.
Previous editions featured three metastatic values (MX, M0 and M1) which referred respectively to absence of adequate information, the confirmed absence, or the presence of breast cancer cells in locations other than the breast and regional lymph nodes, such as to bone, brain, lung.
AJCC has provided web accessible poster versions of the current versions of these copyrighted TNM descriptors and groups,[30] and readers should refer to that up to date, accurate information[30] or to the National Cancer Institute (NCI)[23] or National Comprehensive Cancer Network[39] sites which reprints these with AJCC permission.
For accurate, complete, current details refer to the accessible copyrighted documentation from AJCC,[30] or to the authorized documentation from NCI[23] or NCCN;[39] for past editions refer to AJCC.[29]
Receptor status[edit]
The receptor status of breast cancers has traditionally been identified by immunohistochemistry (IHC), which stains the cells based on the presence of estrogen receptors (ER), progesterone receptors (PR) and HER2. This remains the most common method of testing for receptor status, but DNA multi-gene expression profiles can categorize breast cancers into molecular subtypes that generally correspond to IHC receptor status; one commercial source is the BluePrint test, as discussed in the following section.
Receptor status is a critical assessment for all breast cancers as it determines the suitability of using targeted treatments such as tamoxifen and or trastuzumab. These treatments are now some of the most effective adjuvant treatments of breast cancer. Estrogen receptor positive (ER+) cancer cells depend on estrogen for their growth, so they can be treated with drugs to reduce either the effect of estrogen (e.g. tamoxifen) or the actual level of estrogen (e.g. aromatase inhibitors), and generally have a better prognosis. Generally, prior to modern treatments, HER+ had a worse prognosis,[40] however HER2+ cancer cells respond to drugs such as the monoclonal antibody, trastuzumab, (in combination with conventional chemotherapy) and this has improved the prognosis significantly.[41] Conversely, triple negative cancer (i.e. no positive receptors), lacking targeted treatments, now has a comparatively poor prognosis.[42][43]
Androgen receptor is expressed in 80-90% of ER+ breast cancers and 40% of "triple negative" breast cancers. Activation of androgen receptors appears to suppress breast cancer growth in ER+ cancer while in ER- breast it appears to act as growth promoter. Efforts are underway to utilize this as prognostic marker and treatment.[44][45]
Molecular subtype[edit]
Receptor status was traditionally considered by reviewing each individual receptor (ER, PR, her2) in turn, but newer approaches look at these together, along with the tumor grade, to categorize breast cancer into several conceptual molecular classes[46] that have different prognoses[39] and may have different responses to specific therapies.[47] DNA microarrays have assisted this approach, as discussed in the following section. Proposed molecular subtypes include:
- Basal-like: ER-, PR- and HER2-; also called triple negative breast cancer (TNBC).[48] Most BRCA1 breast cancers are basal-like TNBC.
- Luminal A: ER+ and low grade
- Luminal B: ER+ but often high grade
- Luminal ER-/AR+: (overlapping with apocrine and so called molecular apocrine) - recently identified androgen responsive subtype which may respond to antihormonal treatment with bicalutamide[44]
- ERBB2/HER2-amplified: has overexpressed HER2/neu[40]
- Normal breast-like[39][46][49]
- Claudin-low: a more recently described class; often triple-negative, but distinct in that there is low expression of cell-cell junction proteins[48] including E-cadherin[48] and frequently there is infiltration with lymphocytes.[46][50][51]
| Luminal A | Luminal B | ERBB2/HER2-amplified | Basal-like | |
|---|---|---|---|---|
| Overall gene expression | High expression of:
|
Compared to Luminal A, higher expression of:
Lower expression of:
|
|
High expression of:
Low expression of:
|
| Light microscopy types |
|
|
|
|
| Immunohistochemistry |
|
|
|
|
| Gene mutations |
|
Similar to luminal A but:
|
|
|
DNA classification[edit]
Traditional DNA classification[edit]
Traditional DNA classification was based on the general observation that cells that are dividing more quickly have a worse prognosis, and relied on either the presence of protein Ki67 or the percentage of cancer cell DNA in S phase. These methods, and scoring systems that used DNA ploidy, are used much less often now, as their predictive and prognostic power was less substantial than other classification schemes such as the TNM stage. In contrast, modern DNA analyses are increasingly relevant in defining underlying cancer biology and in helping choose treatments.[53][54][55][56]
HER2/neu[edit]
HER2/neu status can be analyzed by fluorescent in-situ hybridization (FISH) assays.[39] Some commentators prefer this approach, claiming a higher correlation than receptor immunohistochemistry with response to trastuzumab, a targeted therapy, but guidelines permit either testing method.[39]
DNA microarrays[edit]
Background[edit]

DNA microarrays have compared normal cells to breast cancer cells and found differences in the expression of hundreds of genes. Although the significance of many of those genetic differences is unknown, independent analyses by different research groups has found that certain groups of genes have a tendency to co-express. These co-expressing clusters have included hormone receptor-related genes, HER2-related genes, a group of basal-like genes, and proliferation genes. As might therefore be anticipated, there is considerable similarity between the receptor and microarray classifications, but assignment of individual tumors is by no means identical. By way of illustration, some analyses have suggested that approximately 75% of receptor classified triple-negative breast cancers (TNBC) basal-like tumors have the expected DNA expression profile, and a similar 75% of tumors with a typical basal-like DNA expression profile are receptor TNBC as well. To say this in a different way to emphasize things, this means that 25% of triple-negative breast cancer (TNBC) basal-like tumors as defined by one or other classification are excluded from the alternative classification's results. Which classification scheme (receptor IHC or DNA expression profile) more reliably assorts particular cancers to effective therapies is under investigation.
Several commercially marketed DNA microarray tests analyze clusters of genes and may help decide which possible treatment is most effective for a particular cancer.[57] The use of these assays in breast cancers is supported by Level II evidence or Level III evidence. No tests have been verified by Level I evidence, which is rigorously defined as being derived from a prospective, randomized controlled trial where patients who used the test had a better outcome than those who did not. Acquiring extensive Level I evidence would be clinically and ethically challenging. However, several validation approaches[58][59] are being actively pursued.
Numerous genetic profiles have been developed.[60][61] The most heavily marketed are:
- Oncotype DX is supported by Level II evidence, and was originally designed for use in estrogen receptor (ER) positive tumors,[62] and has been endorsed by the American Society of Clinical Oncology (ASCO)[60][63] and the NCCN.[39]
- MammaPrint is supported only by Level III evidence, can be performed on estrogen receptor (ER) positive and negative tumors, and has FDA approval.
- Two other tests also only have Level III evidence: Theros and MapQuant Dx.[64][65][66]
These multigene assays, some partially and some completely commercialized, have been scientifically reviewed to compare them with other standard breast cancer classification methods such as grade and receptor status.[49][61] Although these gene-expression profiles look at different individual genes, they seem to classify a given tumor into similar risk groups and thus provide concordant predictions of outcome.[39][62]
Although there is considerable evidence that these tests can refine the treatment decisions in a meaningful proportion of breast cancers[60][61] they are fairly expensive; proposed selection criteria for which particular tumors may benefit by being interrogated by these assays[39] remain controversial, particularly with lymph node positive cancers.[39] One review characterized these genetic tests collectively as adding "modest prognostic information for patients with HER2-positive and triple-negative tumors, but when measures of clinical risk are equivocal (e.g., intermediate expression of ER and intermediate histologic grade), these assays could guide clinical decisions".[40]
Oncotype DX[edit]
Oncotype DX assesses 16 cancer-related genes and 5 normal comparator reference genes, and is therefore sometimes known as the 21-gene assay. It was designed for use in estrogen receptor (ER) positive tumors. The test is run on formalin fixed, paraffin-embedded tissue. Oncotype results are reported as a Recurrence Score (RS), where a higher RS is associated with a worse prognosis, referring to the likelihood of recurrence without treatment. In addition to that prognostic role, a higher RS is also associated with a higher probability of response to chemotherapy, which is termed a positive predictive factor.
These results suggest that not only does Oncotype stratify estrogen-receptor positive breast cancer into different prognostic groups, but also suggest that cancers that have a particularly favorable Oncotype DX microarray result tend to derive minimal benefit from adjuvant chemotherapy and so it may be appropriate to choose to avoid side effects from that additional treatment. As an additional example, a neoadjuvant clinical treatment program that included initial chemotherapy followed by surgery and subsequent additional chemotherapy, radiotherapy, and hormonal therapy found a strong correlation of the Oncotype classification with the likelihood of a complete response (CR) to the presurgical chemotherapy.[67]
Since high risk features may already be evident in many high risk cancers, for example hormone-receptor negativity or HER-2 positive disease, the Oncotype test may especially improve the risk assessment that is derived from routine clinical variables in intermediate risk disease.[68] Results from both the US[69] and internationally[70] suggest that Oncotype may assist in treatment decisions.[71]
Oncotype DX has been endorsed by the American Society of Clinical Oncology (ASCO)[60][63] and the National Comprehensive Cancer Network (NCCN).[39] The NCCN Panel considers the 21-gene assay as an option when evaluating certain tumors[39] to assist in estimating likelihood of recurrence and benefit from chemotherapy, emphasizing that the recurrence score should be used along with other breast cancer classification elements when stratifying risk.[39] Oncotype fulfilled all California Technology Assessment Forum (CTAF) criteria in October 2006.[72] The U.S. Food and Drug Administration (FDA) does not mandate approval of this class of tests if they are performed at a single, company-operated laboratory[73] Genomic Health, which developed Oncotype DX, offers the test under these so-called home brew rules and, accordingly, to that extent the Oncotype DX assay is not specifically FDA approved.[73]
MammaPrint and BluePrint[edit]
The MammaPrint gene pattern is a commercial-stage 70-gene panel marketed by Agendia,[74] that was developed in patients under age 55 years who had lymph node negative breast cancers (N0).[72] The commercial test is marketed for use in breast cancer irrespective of estrogen receptor (ER) status.[72] The test is run on formalin fixed, paraffin-embedded tissue. MammaPrint traditionally used rapidly frozen tissue[39] but a room temperature, molecular fixative is available for use within 60 minutes of obtaining fresh tissue samples.[75]
A summary of clinical trials using MammaPrint is included in the MammaPrint main article. The available evidence for Mammaprint was reviewed by California Technology Assessment Forum (CTAF) in June 2010; the written report indicated that MammaPrint had not yet fulfilled all CTAF criteria.[72] MammaPrint has 5 FDA clearances and is the only FDA cleared microarray assay available. To be eligible for the MammaPrint gene expression profile, a breast cancer should have the following characteristics: stage 1 or 2, tumor size less than 5.0 cm, estrogen receptor positive (ER+) or estrogen receptor negative (ER-). In the US, the tumor should also be lymph node negative (N0), but internationally the test may be performed if the lymph node status is negative or positive with up to 3 nodes.[76]
One method of assessing the molecular subtype of a breast cancer is by BluePrint,[77] a commercial-stage 80-gene panel marketed by Agendia, either as a standalone test, or combined with the MammaPrint gene profile.
Other DNA assays and choice of treatment[edit]
The choice of established chemotherapy medications, if chemotherapy is needed, may also be affected by DNA assays that predict relative resistance or sensitivity. Topoisomerase II (TOP2A) expression predicts whether doxorubicin is relatively useful.[78][79] Expression of genes that regulate tubulin may help predict the activity of taxanes.
Various molecular pathway targets and DNA results are being incorporated in the design of clinical trials of new medicines.[80] Specific genes such as p53, NME1, BRCA and PIK3CA/Akt may be associated with responsiveness of the cancer cells to innovative research pharmaceuticals. BRCA1 and BRCA2 polymorphic variants can increase the risk of breast cancer, and these cancers tend to express a pr ofile of genes, such as p53, in a pattern that has been called "BRCA-ness." Cancers arising from BRCA1 and BRCA2 mutations, as well as other cancers that share a similar "BRCA-ness" profile, including some basal-like receptor triple negative breast cancers, may respond to treatment with PARP inhibitors[81] such as olaparib. Combining these newer medicines with older agents such as 6-Thioguanine (6TG) may overcome the resistance that can arise in BRCA cancers to PARP inhibitors or platinum-based chemotherapy.[82] mTOR inhibitors such as everolimus may show more effect in PIK3CA/Akt e9 mutants than in e20 mutants or wild types.[83]
DNA methylation patterns can epigenetically affect gene expression in breast cancer and may contribute to some of the observed differences between genetic subtypes.[84]
Tumors overexpressing the Wnt signaling pathway co-receptor low-density lipoprotein receptor-related protein 6 (LRP6) may represent a distinct subtype of breast cancer and a potential treatment target.[85]
Numerous clinical investigations looked at whether testing for variant genotype polymorphic alleles of several genes could predict whether or not to prescribe tamoxifen; this was based on possible differences in the rate of conversion of tamoxifen to the active metabolite, endoxifen. Although some studies had suggested a potential advantage from CYP2D6 testing, data from two large clinical trials found no benefit.[86][87] Testing for the CYP2C19*2 polymorphism gave counterintuitive results.[88] The medical utility of potential biomarkers of tamoxifen responsiveness such as HOXB13,[89] PAX2,[90] and estrogen receptor (ER) alpha and beta isoforms interaction with SRC3[91][92] have all yet[when?] to be fully defined.
Other classification approaches[edit]
Computer models[edit]
Computer models consider several traditional factors concurrently to derive individual survival predictions and calculations of potential treatment benefits. The validated algorithms can present visually appealing graphics that assist in treatment decisions. In addition, other classifications of breast cancers do exist and no uniform system has been consistently adopted worldwide.
Adjuvant! is based on US cohorts[93] and presents colored bar charts that display information that may assist in decisions regarding systemic adjuvant therapies. Successful validation was seen with Canadian[94] and Dutch[95] cohorts. Adjuvant! seemed less applicable to a British cohort[96] and accordingly PREDICT is being developed in the United Kingdom.[97]
Other immunohistochemical tests[edit]
Among the immunohistochemical tests that may further stratify prognosis, BCL2 has shown promise in preliminary studies.[98]
Van Nuys prognostic index[edit]
The USC/Van Nuys prognostic index (VNPI) classifies ductal carcinoma in situ (DCIS) into dissimilar risk categories that may be treated accordingly.[99]
Notes[edit]
- ^ a b c d e Area per high-power field for some microscope types:
- Olympus BX50, BX40 or BH2 or AO or Nikon with 15x eyepiece: 0.096 mm2
- AO with 10x eyepiece: 0.12 mm2
- Nikon or Olympus with 10x eyepiece: 0.16 mm2
- Leitz Ortholux: 0.27 mm2
- Leitz Diaplan: 0.31 mm2
- "Infiltrating Ductal Carcinoma of the Breast (Carcinoma of No Special Type)". Stanford University School of Medicine. Archived from the original on 11 September 2019. Retrieved 2 October 2019.
References[edit]
- ^ Gonzalez-Angulo AM, Morales-Vasquez F, Hortobagyi GN (2007). "Overview of Resistance to Systemic Therapy in Patients with Breast Cancer". In Yu D, Hung MC (eds.). Breast Cancer Chemosensitivity. Advances in Experimental Medicine and Biology. Vol. 608. pp. 1–22. doi:10.1007/978-0-387-74039-3_1. ISBN 978-0-387-74037-9. PMID 17993229.
- ^ Giordano SH, Hortobagyi GN (2003). "Inflammatory breast cancer: Clinical progress and the main problems that must be addressed". Breast Cancer Res. 5 (6): 284–8. doi:10.1186/bcr608. PMC 314400. PMID 14580242.
- ^ Merck Manual, Professional Edition Archived 1 July 2010 at the Wayback Machine, Ch. 253, Breast Cancer.
- ^ a b "FDA Approves First Targeted Therapy for HER2-Low Breast Cancer". U.S. Food and Drug Administration (FDA) (Press release). 5 August 2022. Archived from the original on 6 August 2022. Retrieved 5 August 2022.
 This article incorporates text from this source, which is in the public domain.
This article incorporates text from this source, which is in the public domain.
- ^ References for pie chart is located at Image description page in commons
- ^ a b c d Page 1084 Archived 6 August 2022 at the Wayback Machine in: Robbins, Stanley (2010). Robbins and Cotran pathologic basis of disease. Philadelphia, PA: Saunders/Elsevier. ISBN 978-1-4377-2182-9. OCLC 489074868.
- ^ Ductal carcinoma in situ: 99% - Kerlikowske, K (2010). "Epidemiology of ductal carcinoma in situ". Journal of the National Cancer Institute. Monographs. 2010 (41): 139–41. doi:10.1093/jncimonographs/lgq027. PMC 5161058. PMID 20956818.
Lobular carcinoma in situ: 97% - Xie, Ze-Ming; Sun, Jian; Hu, Zhe-Yu; Wu, Yao-Pan; Liu, Peng; Tang, Jun; Xiao, Xiang-Sheng; Wei, Wei-Dong; Wang, Xi; Xie, Xiao-Ming; Yang, Ming-Tian (2017). "Survival outcomes of patients with lobular carcinoma in situ who underwent bilateral mastectomy or partial mastectomy". European Journal of Cancer. 82: 6–15. doi:10.1016/j.ejca.2017.05.030. ISSN 0959-8049. PMID 28646773. - ^ a b Percentage values are from United States statistics 2004. Subtype specific incidences are taken from Table 6 Archived 23 February 2013 at archive.today (invasive) and Table 3 Archived 23 February 2013 at archive.today (in situ) from Eheman CR, Shaw KM, Ryerson AB, Miller JW, Ajani UA, White MC (June 2009). "The changing incidence of in situ and invasive ductal and lobular breast carcinomas: United States, 1999–2004". Cancer Epidemiol. Biomarkers Prev. 18 (6): 1763–9. doi:10.1158/1055-9965.EPI-08-1082. PMID 19454615.. These are divided by total breast cancer incidence (211,300 invasive and 55,700 in situ cases) as reported from Breast Cancer Facts & Figures 2003–2004 "ACS :: Breast Cancer Facts & Figures 2003-2004". Archived from the original on 15 April 2009. Retrieved 15 June 2010.
- ^ NOTE: Number really refers to invasive ductal carcinoma, despite title. Arpino G, Bardou VJ, Clark GM, Elledge RM (2004). "Infiltrating lobular carcinoma of the breast: tumor characteristics and clinical outcome". Breast Cancer Res. 6 (3): R149–56. doi:10.1186/bcr767. PMC 400666. PMID 15084238.
- ^ Evans, A. (2004). "Ductal carcinoma in situ (DCIS): are we overdetecting it?". Breast Cancer Research. 6 (Suppl 1): P23. doi:10.1186/bcr842. PMC 3300383. [1] Archived 8 July 2011 at the Wayback Machine
- ^ World Health Organization: Tumours of the Breast and Female Genital Organs. Oxford [Oxfordshire]: Oxford University Press. 2003. ISBN 978-92-832-2412-9.
- ^ Elston, CW; Ellis, IO (1991). "Pathologic prognostic factors in breast cancer. I. The value of histological grades in breast cancer. Experience from a large study with long-term follow-up". Histopathology. 19 (5): 403–10. doi:10.1111/j.1365-2559.1991.tb00229.x. PMID 1757079. S2CID 17622089. "Republished". Histopathology. 41: 154–161. 2002. doi:10.1046/j.1365-2559.2002.14892.x. S2CID 208083532.
- ^ Oudai Hassan. "What is the Nottingham combined histologic grade (modified Scarff-Bloom-Richardson grade) system for breast tumors?". Medscape. Archived from the original on 30 November 2020. Retrieved 2 October 2019. Updated: 20 March 2019
- ^ Al-Kuraya K, Schraml P, Torhorst J, Tapia C, Zaharieva B, Novotny H, et al. (December 2004). "Prognostic relevance of gene amplifications and coamplifications in breast cancer". Cancer Research. 64 (23): 8534–8540. doi:10.1158/0008-5472.CAN-04-1945. PMID 15574759.
- ^ Elston CW, Ellis IO. "Pathologic prognostic factors in breast cancer. I. The value of histological grades in breast cancer. Experience from a large study with long-term follow-up". Histopathology. 1991 (19): 403–410.
- ^ Bloom, H.J.; Richardson, W.W. (1957). "Histological grading and prognosis in breast cancer; A study of 1409 cases of which 359 have been followed for 15 years". British Journal of Cancer. 11 (3): 359–77. doi:10.1038/bjc.1957.43. PMC 2073885. PMID 13499785.
- ^ Genestie C, Zafrani B, Asselain B, Fourquet A, Rozan S, Validire P, et al. (1998). "Comparison of the prognostic value of Scarff-Bloom-Richardson and Nottingham histological grades in a series of 825 cases of breast cancer: Major importance of the mitotic count as a component of both grading systems". Anticancer Research. 18 (1B): 571–6. PMID 9568179.
- ^ "Medullary Carcinoma of the Breast". Stanford Medicine. Archived from the original on 8 September 2020. Retrieved 31 December 2020.
- ^ "Metaplastic Carcinoma of the Breast". Stanford Medicine. Archived from the original on 8 September 2020. Retrieved 31 December 2020.
- ^ Pujani, Mukta; Sharma, KiranLata; Srivastava, AN; Singh, US; Bansal, Cherry (2014). "Grading systems in the cytological diagnosis of breast cancer: A review". Journal of Cancer Research and Therapeutics. 10 (4): 839–845. doi:10.4103/0973-1482.140979. ISSN 0973-1482. PMID 25579516.
- ^ "Infiltrating Ductal Carcinoma of the Breast (Carcinoma of No Special Type)". Stanford University School of Medicine. Archived from the original on 11 September 2019. Retrieved 2 October 2019.
- ^ a b What is Cancer Staging? American Joint Committee on Cancer 2010 May 5.http://www.cancerstaging.org/mission/whatis.html Archived 27 August 2013 at the Wayback Machine
- ^ a b c d National Cancer Institute. Stage Information for Breast Cancer.http://www.cancer.gov/cancertopics/pdq/treatment/breast/healthprofessional/page3 Archived 8 June 2011 at the Wayback Machine
- ^ a b c Originally copied from Fadi M. Alkabban; Troy Ferguson (2021). "Breast Cancer". Cancer, Breast. StatPearls. PMID 29493913. Archived from the original on 24 December 2020. Retrieved 4 October 2019.
{{cite book}}:|website=ignored (help) Last Update: 4 June 2019. Creative Commons Attribution 4.0 International License Archived 21 November 2015 at the Wayback Machine - ^ Scatarige JC, Fishman EK, Zinreich ES, Brem RF, Almaraz R (April 1988). "Internal mammary lymphadenopathy in breast carcinoma: CT appraisal of anatomic distribution". Radiology. 167 (1): 89–91. doi:10.1148/radiology.167.1.3347753. PMID 3347753.
- ^ Scatarige JC, Boxen I, Smathers RL (September 1990). "Internal mammary lymphadenopathy: imaging of a vital lymphatic pathway in breast cancer". Radiographics. 10 (5): 857–70. doi:10.1148/radiographics.10.5.2217975. PMID 2217975. available as full text article with multiple images at http://radiographics.rsna.org/content/10/5/857.full.pdf
- ^ a b c d Joseph A Sparano, MD (6 October 2021). "Breast Cancer Staging". medscape. Archived from the original on 13 September 2020. Retrieved 21 August 2020. Updated: 8 June 2019
- ^ a b Data from Mangone L, Marinelli F, Bisceglia I, Braghiroli MB, Damato A, Pinto C (2022). "Five-year relative survival by stage of breast and colon cancers in northern Italy". Front Oncol. 12: 982461. doi:10.3389/fonc.2022.982461. PMC 9659859. PMID 36387150.
{{cite journal}}: CS1 maint: multiple names: authors list (link) - ^ a b c d e American Joint Committee on Cancer. Past Editions of the AJCC Cancer Staging Manual available at "American Joint Committee on Cancer: Publications & Electronic Products: Past Editions of the AJCC Cancer Staging Manual". Archived from the original on 6 January 2011. Retrieved 3 January 2011.
- ^ a b c d AJCC Cancer Staging Manual, 7th edition, updated Cancer Staging Posters.http://www.cancerstaging.org/staging/posters/breast8.5x11.pdf Archived 26 June 2011 at the Wayback Machine
- ^ AJCC Cancer Staging Manual 6th edition; Chapter 25; Breast Cancer – original pages 223-240 http://www.cancerstaging.org/products/csmanual6ed-4.pdf Archived 28 September 2011 at the Wayback Machine
- ^ AJCC Cancer Staging Manual 5th edition; Chapter 25; Breast Cancer – original pages 171-180 http://www.cancerstaging.org/products/csmanual5ed_4.pdf Archived 28 September 2011 at the Wayback Machine
- ^ AJCC Cancer Staging Manual 4th edition; Chapter 25; Breast Cancer – original pages 149-154 http://www.cancerstaging.org/products/csmanual4ed_2.pdf Archived 28 September 2011 at the Wayback Machine
- ^ AJCC Cancer Staging Manual 3rd edition; 1988; Chapter 23; Breast Cancer-original pages 145-150 http://www.cancerstaging.org/products/csmanual3ed_2.pdf Archived 28 September 2011 at the Wayback Machine
- ^ AJCC Cancer Staging Manual 2nd edition; 1983; Chapter 21; Breast Cancer-original pages 127-134 http://www.cancerstaging.org/products/csmanual2ed.pdf Archived 28 September 2011 at the Wayback Machine
- ^ AJCC Cancer Staging Manual 1977; Chapter 12; Breast Cancer-original pages 101-108 http://www.cancerstaging.org/products/csmanual1ed.pdf Archived 28 September 2011 at the Wayback Machine
- ^ Feinstein, A. R.; Sosin, D. M.; Wells, C. K. (1985). "The Will Rogers Phenomenon". New England Journal of Medicine. 312 (25): 1604–1608. doi:10.1056/NEJM198506203122504. PMID 4000199.
- ^ Woodward WA, Strom EA, Tucker SL, McNeese MD, Perkins GH, Schechter NR, et al. (September 2003). "Changes in the 2003 American Joint Committee on Cancer staging for breast cancer dramatically affect stage-specific survival". Journal of Clinical Oncology. 21 (17): 3244–3248. doi:10.1200/JCO.2003.03.052. PMID 12947058.
- ^ a b c d e f g h i j k l m n "National Comprehensive Cancer Network (NCCN) guidelines, Breast Cancer Version 2.2011" (PDF). Archived (PDF) from the original on 24 March 2009. Retrieved 3 July 2011.
- ^ a b c Sotirou Christos, Pusztai Lajos (2009). "Molecular origin of cancer: gene-expression signatures in breast cancer". The New England Journal of Medicine. 360 (8): 790–800. doi:10.1056/NEJMra0801289. PMID 19228622. Archived from the original on 5 January 2010. Retrieved 16 June 2010.
- ^ Romond EH, Perez EA, Bryant J, Suman VJ, Geyer CE, Davidson NE, et al. (October 2005). "Trastuzumab plus adjuvant chemotherapy for operable HER2+ breast cancer". The New England Journal of Medicine. 353 (16): 1673–1684. doi:10.1056/NEJMoa052122. PMID 16236738. S2CID 9534884.
- ^ Dent R, Trudeau M, Pritchard KI, Hanna WM, Kahn HK, Sawka CA, et al. (August 2007). "Triple-negative breast cancer: clinical features and patterns of recurrence". Clinical Cancer Research. 13 (15 Pt 1): 4429–4434. doi:10.1158/1078-0432.CCR-06-3045. PMID 17671126.
- ^ "Understanding and Treating Triple-Negative Breast Cancer". Cancer Network. Archived from the original on 9 June 2010. Retrieved 8 May 2010.
- ^ a b Lehmann BD, Bauer JA, Chen X, Sanders ME, Chakravarthy AB, Shyr Y, et al. (July 2011). "Identification of human triple-negative breast cancer subtypes and preclinical models for selection of targeted therapies". Journal of Clinical Investigation. 121 (7): 2750–2767. doi:10.1172/JCI45014. PMC 3127435. PMID 21633166.
- ^ Hu R, Dawood S, Holmes MD, Collins LC, Schnitt SJ, Cole K, et al. (2011). "Androgen receptor expression and breast cancer survival in postmenopausal women". Clinical Cancer Research. 17 (7): 1867–1874. doi:10.1158/1078-0432.CCR-10-2021. PMC 3076683. PMID 21325075.
- ^ a b c Prat, A.; Perou, C. M. (2011). "Deconstructing the molecular portraits of breast cancer". Molecular Oncology. 5 (1): 5–23. doi:10.1016/j.molonc.2010.11.003. PMC 5528267. PMID 21147047.
- ^ Geyer, F. C.; Marchiò, C.; Reis-Filho, J. S. (2009). "The role of molecular analysis in breast cancer". Pathology. 41 (1): 77–88. doi:10.1080/00313020802563536. PMID 19089743. S2CID 39374813.
- ^ a b c Perou, C. M. (2011). "Molecular Stratification of Triple-Negative Breast Cancers". The Oncologist. 16: 61–70. doi:10.1634/theoncologist.2011-S1-61. PMID 21278442.
- ^ a b Ross, J. S. (2009). "Multigene Classifiers, Prognostic Factors, and Predictors of Breast Cancer Clinical Outcome". Advances in Anatomic Pathology. 16 (4): 204–215. doi:10.1097/PAP.0b013e3181a9d4bf. PMID 19546609. S2CID 37465636.
- ^ Herschkowitz JI, Zhao W, Zhang M, Usary J, Murrow G, Edwards D, et al. (February 2012). "Breast Cancer Special Feature: Comparative oncogenomics identifies breast tumors enriched in functional tumor-initiating cells". Proceedings of the National Academy of Sciences. 109 (8): 2778–2783. doi:10.1073/pnas.1018862108. PMC 3286979. PMID 21633010.
- ^ Harrell JC, Prat A, Parker JS, Fan C, He X, Carey L, et al. (April 2012). "Genomic analysis identifies unique signatures predictive of brain, lung, and liver relapse". Breast Cancer Research and Treatment. 132 (2): 523–535. doi:10.1007/s10549-011-1619-7. PMC 3303043. PMID 21671017.
- ^ Tsang, Julia Y.S.; Tse, Gary M. (2020). "Molecular Classification of Breast Cancer". Advances in Anatomic Pathology. 27 (1): 27–35. doi:10.1097/PAP.0000000000000232. ISSN 1072-4109. PMID 31045583. S2CID 143423297.
- ^ Perou CM, Sørlie T, Eisen MB, van de Rijn M, Jeffrey SS, Rees CA, et al. (August 2000). "Molecular portraits of human breast tumours". Nature. 406 (6797): 747–752. Bibcode:2000Natur.406..747P. doi:10.1038/35021093. PMID 10963602. S2CID 1280204.
- ^ Nagasaki K, Miki Y (2006). "Gene expression profiling of breast cancer". Breast Cancer. 13 (1): 2–7. doi:10.2325/jbcs.13.2. PMID 16518056. S2CID 6255927.
- ^ Normanno N, De Luca A, Carotenuto P, Lamura L, Oliva I, D'Alessio A (2009). "Prognostic applications of gene expression signatures in breast cancer". Oncology. 77 (Suppl 1): 2–8. doi:10.1159/000258489. PMID 20130425. S2CID 21011879.
- ^ Jönsson G, Staaf J, Vallon-Christersson J, Ringnér M, Holm K, Hegardt C, et al. (2010). "Genomic subtypes of breast cancer identified by array-comparative genomic hybridization display distinct molecular and clinical characteristics". Breast Cancer Research. 12 (3): R42. doi:10.1186/bcr2596. PMC 2917037. PMID 20576095.
- ^ Sparano JA, Solin LJ (April 2010). "Defining the clinical utility of gene expression assays in breast cancer: the intersection of science and art in clinical decision making". Journal of Clinical Oncology. 28 (10): 1625–7. doi:10.1200/JCO.2009.25.2882. PMID 20065178.
- ^ Mandrekar SJ, Sargent DJ (October 2010). "Predictive biomarker validation in practice: lessons from real trials". Clin Trials. 7 (5): 567–73. doi:10.1177/1740774510368574. PMC 3913192. PMID 20392785.
- ^ Pharoah PD, Caldas C (November 2010). "Genetics: How to validate a breast cancer prognostic signature". Nat Rev Clin Oncol. 7 (11): 615–6. doi:10.1038/nrclinonc.2010.142. PMID 20981123. S2CID 32411967.
- ^ a b c d Ross, J. S.; Hatzis, C.; Symmans, W. F.; Pusztai, L.; Hortobagyi, G. N. (2008). "Commercialized Multigene Predictors of Clinical Outcome for Breast Cancer". The Oncologist. 13 (5): 477–493. doi:10.1634/theoncologist.2007-0248. PMID 18515733. S2CID 19435207.
- ^ a b c Albain, K. S.; Paik, S.; Van't Veer, L. (2009). "Prediction of adjuvant chemotherapy benefit in endocrine responsive, early breast cancer using multigene assays". The Breast. 18: S141–S145. doi:10.1016/S0960-9776(09)70290-5. PMID 19914534.
- ^ a b Fan C, Oh DS, Wessels L, Weigelt B, Nuyten DS, Nobel AB, et al. (August 2006). "Concordance among gene-expression-based predictors for breast cancer". New England Journal of Medicine. 355 (6): 560–569. doi:10.1056/NEJMoa052933. PMID 16899776.
- ^ a b Harris L, Fritsche H, Mennel R, Norton L, Ravdin P, Taube S, et al. (((American Society of Clinical Oncology))) (November 2007). "American Society of Clinical Oncology 2007 update of recommendations for the use of tumor markers in breast cancer". Journal of Clinical Oncology. 25 (33): 5287–5312. doi:10.1200/JCO.2007.14.2364. PMC 2793754. PMID 17954709.
- ^ Armen Hareyanon. MapQuant Dx Genomic Grade Test Identifies Breast Cancer Patients. 2 June 2008 http://www.emaxhealth.com/98/22731.html Archived 29 September 2011 at the Wayback Machine
- ^ Filho, O. M.; Ignatiadis, M.; Sotiriou, C. (2011). "Genomic Grade Index: An important tool for assessing breast cancer tumor grade and prognosis". Critical Reviews in Oncology/Hematology. 77 (1): 20–29. doi:10.1016/j.critrevonc.2010.01.011. PMID 20138540.
- ^ "Select your country: Qiagen Marseille" (PDF). ipsogen.com. Archived from the original (PDF) on 26 March 2012. Retrieved 12 December 2015.
- ^ Gianni L, Zambetti M, Clark K, Baker J, Cronin M, Wu J, et al. (October 2005). "Gene expression profiles in paraffin-embedded core biopsy tissue predict response to chemotherapy in women with locally advanced breast cancer". Journal of Clinical Oncology. 23 (29): 7265–7277. doi:10.1200/JCO.2005.02.0818. PMID 16145055.
- ^ Kelly CM, Krishnamurthy S, Bianchini G, Litton JK, Gonzalez-Angulo AM, Hortobagyi GN, et al. (November 2010). "Utility of oncotype DX risk estimates in clinically intermediate risk hormone receptor-positive, HER2-normal, grade II, lymph node-negative breast cancers". Cancer. 116 (22): 5161–5167. doi:10.1002/cncr.25269. PMID 20665886.
- ^ Lo SS, Mumby PB, Norton J, Rychlik K, Smerage J, Kash J, et al. (April 2010). "Prospective multicenter study of the impact of the 21-gene recurrence score assay on medical oncologist and patient adjuvant breast cancer treatment selection". Journal of Clinical Oncology. 28 (10): 1671–1676. doi:10.1200/JCO.2008.20.2119. PMID 20065191.
- ^ Albanell, J.; González, A.; Ruiz-Borrego, M.; Alba, E.; García-Saenz, J. A.; Corominas, J. M.; Burgues, O.; Furio, V.; Rojo, A.; Palacios, J.; Bermejo, B.; Martínez-García, M.; Limon, M. L.; Muñoz, A. S.; Martín, M.; Tusquets, I.; Rojo, F.; Colomer, R.; Faull, I.; Lluch, A. (2011). "Prospective transGEICAM study of the impact of the 21-gene Recurrence Score assay and traditional clinicopathological factors on adjuvant clinical decision making in women with estrogen receptor-positive (ER+) node-negative breast cancer". Annals of Oncology. 23 (3): 625–631. doi:10.1093/annonc/mdr278. PMID 21652577.
- ^ New Mexico OncologyHematology Consultants, Ltd. Copyright held by CancerConsultants Breast Cancer Information Center. European Study Reports that Oncotype DX Influences Breast Cancer Treatment Decisions. Posted 2010 October 17, accessioned 2010 Dec 19 and 2, 011 July 3 at http://nmcancercenter.org/european-study-reports-that-oncotype-dx%C2%AE-influences-breast-cancer-treatment-decisions/ Archived 28 March 2012 at the Wayback Machine
- ^ a b c d Tice JA. The 70-Gene Signature (MammaPrint) as a Guide for the Management of Early Stage Breast Cancer. California Technology Assessment Forum. 2010 June 2nd. Full text accessioned 2010 Dec 19 at http://www.ctaf.org/content/assessments/detail/?id=1178 Archived 25 July 2011 at the Wayback Machine
- ^ a b NCI Cancer Bulletin FDA Update 2007 February 14, Volume 4, Number 7 as Retrieved 17 October 2010 at http://www.cancer.gov/aboutnci/ncicancerbulletin/archive/2007/021407/page5 Archived 22 December 2010 at the Wayback Machine
- ^ The mission of Agendia. Agendia web isite.http://www.agendia.com/pages/mission/86.php Archived 23 November 2013 at the Wayback Machine
- ^ Ordering Symphony is a simple process. Agendia web site.http://www.agendia.com/pages/ordering_symphony/38.php Archived 5 June 2011 at the Wayback Machine
- ^ MammaPrint Patient Eligibility Internationally (Outside of the USA) http://www.agendia.com/pages/patient_eligibility_internationally/317.php Archived 5 June 2011 at the Wayback Machine
- ^ Introducing BluePrint: A Molecular Subtyping Profile for Breast Cancer. Agendia web site.http://www.agendia.com/pages/blueprint/324.php Archived 3 September 2011 at the Wayback Machine
- ^ MacGrogan G, Rudolph P, Mascarel I, Mauriac L, Durand M, Avril A, et al. (August 2003). "DNA topoisomerase II alfa expression and the response to primary chemotherapy in breast cancer". British Journal of Cancer. 89 (4): 666–671. doi:10.1038/sj.bjc.6601185. PMC 2376904. PMID 12915875.
- ^ Gene Review TOP2A – topoisomerase (DNA) II alpha 170kDa Homo sapiens as Retrieved 18 October 2010 [2]
- ^ Alvarez, R. H.; Valero, V.; Hortobagyi, G. N. (2010). "Emerging Targeted Therapies for Breast Cancer". Journal of Clinical Oncology. 28 (20): 3366–3379. doi:10.1200/JCO.2009.25.4011. PMID 20530283.
- ^ Tutt A J Clin Onc 2009; 27(suppl 15): abst CRA501
- ^ Issaeva N, Thomas HD, Djureinovic T, Djurenovic T, Jaspers JE, Stoimenov I, et al. (August 2010). "6-thioguanine selectively kills BRCA2-defective tumors and overcomes PARP inhibitor resistance". Cancer Research. 70 (15): 6268–6276. doi:10.1158/0008-5472.CAN-09-3416. PMC 2913123. PMID 20631063. Correction published at Correction: 6-Thioguanine Selectively Kills BRCA2-Defective Tumors and Overcomes PARP Inhibitor Resistance Cancer Res 2010 October 1;70:7734
- ^ Baselga J, Semiglazov V, van Dam P, Manikhas A, Bellet M, Mayordomo J, et al. (June 2009). "Phase II randomized study of neoadjuvant everolimus plus letrozole compared with placebo plus letrozole in patients with estrogen receptor-positive breast cancer". Journal of Clinical Oncology. 27 (16): 2630–2637. doi:10.1200/JCO.2008.18.8391. PMID 19380449.
- ^ D'Anello L, Sansone P, Storci G, Mitrugno V, D'Uva G, Chieco P, et al. (November 2010). "Epigenetic control of the basal-like gene expression profile via Interleukin-6 in breast cancer cells". Molecular Cancer. 9: 300. doi:10.1186/1476-4598-9-300. PMC 3002335. PMID 21092249.
- ^ Liu, C. -C.; Prior, J.; Piwnica-Worms, D.; Bu, G. (2010). "LRP6 overexpression defines a class of breast cancer subtype and is a target for therapy". Proceedings of the National Academy of Sciences. 107 (11): 5136–5141. Bibcode:2010PNAS..107.5136L. doi:10.1073/pnas.0911220107. PMC 2841938. PMID 20194742.
- ^ Rae JM, Drury S, Hayes DF et al. Lack of Correlation between Gene Variants in Tamoxifen Metabolizing Enymes with Primary Endpoints in the ATAC Trial. 33rd Annual San Antonio Breast Cancer Symposium (SABCS): abstract S1-7 presented 2010 December 9; accessioned 2010 December 17 at http://www.abstracts2view.com/sabcs10/view.php?nu=SABCS10L_1093&terms= Archived 11 August 2011 at the Wayback Machine
- ^ Leyland-Jones B, Regan MM, Bouzyk M et al. Outcome According to CYP2D6 Genotype among Postmenopausal Women with Endocrine-Responsive Early Invasive Breast Cancer Randomized in the BIG 1-98 Trial. 33rd Annual San Antonio Breast Cancer Symposium (SABCS): abstract S1-8 presented 2010, 9 December; accessioned 2010 December 17 at http://www.abstracts2view.com/sabcs10/view.php?nu=SABCS10L_556&terms= Archived 7 July 2011 at the Wayback Machine
- ^ Ruiter R, Bijl MJ, van Schaik RH, Berns EM, Hofman A, Coebergh JW, et al. (October 2010). "CYP2C19*2 polymorphism is associated with increased survival in breast cancer patients using tamoxifen". Pharmacogenomics. 11 (10): 1367–1375. doi:10.2217/pgs.10.112. PMID 21047200.
- ^ Jerevall PL, Jansson A, Fornander T, Skoog L, Nordenskjöld B, Stål O (2010). "Predictive relevance of HOXB13 protein expression for tamoxifen benefit in breast cancer". Breast Cancer Research. 12 (4): 206. doi:10.1186/bcr2612. PMC 2949642. PMID 20649975.
- ^ "Study sheds new light on tamoxifen resistance". CORDIS : News. 13 November 2008. Archived from the original on 20 February 2009. Retrieved 14 November 2008.
Hurtado A, Holmes KA, Geistlinger TR, Hutcheson IR, Nicholson RI, Brown M, et al. (December 2008). "Regulation of ERBB2 by oestrogen receptor-PAX2 determines response to tamoxifen". Nature. 456 (7222): 663–666. Bibcode:2008Natur.456..663H. doi:10.1038/nature07483. PMC 2920208. PMID 19005469. - ^ Mc Ilroy M, Fleming FJ, Buggy Y, Hill AD, Young LS (December 2006). "Tamoxifen-induced ER-alpha-SRC-3 interaction in HER2 positive human breast cancer; a possible mechanism for ER isoform specific recurrence". Endocr. Relat. Cancer. 13 (4): 1135–45. doi:10.1677/erc.1.01222. PMID 17158759.
- ^ Spears M, Bartlett J (June 2009). "The potential role of estrogen receptors and the SRC family as targets for the treatment of breast cancer". Expert Opin. Ther. Targets. 13 (6): 665–74. doi:10.1517/14728220902911509. PMID 19456271. S2CID 39034059.
- ^ Ravdin PM, Siminoff LA, Davis GJ, Mercer MB, Hewlett J, Gerson N, et al. (February 2001). "Computer program to assist in making decisions about adjuvant therapy for women with early breast cancer". Journal of Clinical Oncology. 19 (4): 980–991. doi:10.1200/JCO.2001.19.4.980. PMID 11181660.
- ^ Olivotto IA, Bajdik CD, Ravdin PM, Speers CH, Coldman AJ, Norris BD, et al. (April 2005). "Population-based validation of the prognostic model ADJUVANT! for early breast cancer". Journal of Clinical Oncology. 23 (12): 2716–2725. doi:10.1200/JCO.2005.06.178. PMID 15837986. S2CID 27080534.
- ^ Mook S, Schmidt MK, Rutgers EJ, van de Velde AO, Visser O, Rutgers SM, et al. (November 2009). "Calibration and discriminatory accuracy of prognosis calculation for breast cancer with the online Adjuvant! program: a hospital-based retrospective cohort study". Lancet Oncology. 10 (11): 1070–1076. doi:10.1016/S1470-2045(09)70254-2. PMID 19801202.
- ^ Campbell HE, Taylor MA, Harris AL, Gray AM (October 2009). "An investigation into the performance of the Adjuvant! Online prognostic programme in early breast cancer for a cohort of patients in the United Kingdom". Br. J. Cancer. 101 (7): 1074–84. doi:10.1038/sj.bjc.6605283. PMC 2768087. PMID 19724274.
- ^ Wishart GC, Azzato EM, Greenberg DC, Rashbass J, Kearins O, Lawrence G, et al. (2010). "PREDICT: a new UK prognostic model that predicts survival following surgery for invasive breast cancer". Breast Cancer Research. 12 (1): R1. doi:10.1186/bcr2464. PMC 2880419. PMID 20053270.
- ^ Dawson SJ, Makretsov N, Blows FM, Driver KE, Provenzano E, Le Quesne J, et al. (August 2010). "BCL2 in breast cancer: a favourable prognostic marker across molecular subtypes and independent of adjuvant therapy received". British Journal of Cancer. 103 (5): 668–75. doi:10.1038/sj.bjc.6605736. PMC 2938244. PMID 20664598.
- ^ Silverstein Melvin J., Lagios Michael D. (2010). "Choosing Treatment for Patients With Ductal Carcinoma In Situ: Fine Tuning the University of Southern California/Van Nuys Prognostic Index". J Natl Cancer Inst Monogr. 2010 (41): 193–196. doi:10.1093/jncimonographs/lgq040. PMC 5161065. PMID 20956828.