Hemopexin
Hemopexin (or haemopexin; Hpx; Hx), also known as beta-1B-glycoprotein, is a glycoprotein that in humans is encoded by the HPX gene[5][6][7] and belongs to the hemopexin family of proteins.[8] Hemopexin is the plasma protein with the highest binding affinity for heme.[9]
Hemoglobin itself circulating alone in the blood plasma (called free hemoglobin, as opposed to the hemoglobin situated in and circulating with the red blood cell.) will soon be oxidized into met-hemoglobin which then further disassociates into free heme along with globin chain. The free heme will then be oxidized into free met-heme and sooner or later the hemopexin will come to bind free met-heme together, forming a complex of met-heme and hemopexin, continuing their journey in the circulation until reaching a receptor, such as LRP1, on hepatocytes or macrophages within the spleen, liver and bone marrow.[10]
Hemopexin's arrival and subsequent binding to the free heme not only prevent heme's pro-oxidant and pro-inflammatory effects but also promotes free heme's detoxification.[10]
Hemopexin is different from haptoglobin, the latter always binds to free hemoglobin.[11][10] (See Haptoglobin § Differentiation with hemopexin)
Cloning, expression, and discovery
[edit]Takahashi et al. (1985) determined that human plasma hemopexin consists of a single polypeptide chain of 439 amino acids residues with six intrachain disulfide bridges and has a molecular mass of approximately 63 kD. The amino-terminal threonine residue is modified by a mucin-type O-linked galactosamine oligosaccharide, and the protein has five N-linked glycan modifications. The 18 tryptophan residues are arranged in four clusters, and 12 of the tryptophans are conserved in homologous positions. Computer-assisted analysis of the internal homology in amino acid sequence suggested duplication of an ancestral gene thus indicating that hemopexin consists of two similar halves.[12]
Altruda et al. (1988) demonstrated that the HPX gene spans approximately 12 kb and is interrupted by 9 exons. The demonstration shows direct correspondence between exons and the 10 repeating units in the protein. The introns were not placed randomly; they fell in the center of the region of amino acid sequence homology in strikingly similar locations in 6 of the 10 units and in a symmetric position in each half of the coding sequence. From these observations, Altruda et al. (1988) concluded that the gene evolved through intron-mediated duplications of a primordial sequence to a 5-exon cluster.[13]
Mapping of hemopexin gene
[edit]Cai and Law (1986) prepared a cDNA clone for hemopexin, by Southern blot analysis of human/hamster hybrids containing different combinations of human chromosomes, assigned the HPX gene to human chromosome 11. Law et al. (1988) assigned the HPX gene to 11p15.5-p15.4, the same location as that of the beta-globin gene complex by in situ hybridization.[14]
Differential transcriptional pattern of hemopexin gene
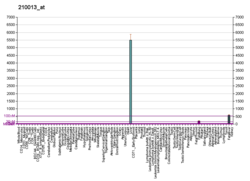
[edit]In 1986, the expression of the human HPX gene in different human tissues and cell lines was carried out by using a specific cDNA probe. From the results obtained it was concluded that this gene was expressed in the liver and it was below the level of detection in other tissues or cell lines examined. By S1 mapping, the transcription initiation site in hepatic cells was located 28 base pairs upstream from the AUG initiation codon of the hemopexin gene.[15]
Function
[edit]Hemopexin binds heme with the highest affinity of any known protein.[9] Its main function is scavenging the heme released or lost by the turnover of heme proteins such as hemoglobin and thus protects the body from the oxidative damage that free heme can cause. In addition, hemopexin releases its bound ligand for internalisation upon interacting with CD91.[16] Hemopexin preserves the body's iron.[17] Hemopexin -dependent uptake of extracellular heme can lead to the deactivation of Bach1 repression which leads to the transcriptional activation of antioxidant heme oxygenase-1 gene. Hemoglobin, haptoglobin (Hp) and Hx associate with high density lipoprotein (HDL) and influence the inflammatory properties of HDL.[18] Hemopexin can downregulate the angiotensin II Type 1 receptor (AT1-R) in vitro.[19]
Clinical significance
[edit]The predominant source of circulating hemopexin is the liver with a plasma concentration of 1–2 mg/ml.[20] Serum hemopexin level reflects how much heme is present in the blood. Therefore, a low hemopexin level indicates that there has been significant degradation of heme containing compounds. A low hemopexin level is one of the diagnostic features of an intravascular hemolytic anemia.[21] Hemopexin has been implicated in cardiovascular disease, septic shock, cerebral ischemic injury, and experimental autoimmune encephalomyelitis.[22] The circulating level of hemopexin is associated with prognosis in patients with septic shock.[22]
HPX is produced in the brain.[23] Deletion of the HPX gene can aggravate brain injury followed by stroma-free hemoglobin-induced intracerebral haemorrhage.[24] High hemopexin level in the cerebrospinal fluid is associated with poor outcome after subarachnoid hemorrhage.[23]
Circulating hemopexin can modulate in patients and in mice anthracycline-induced cardiotoxicity (e.g. heart failure).[25]
Relation to haptoglobin
[edit]In past there have been reports showing that in patients with sickle cell disease, spherocytosis, autoimmune hemolytic anemia, erythropoietic protoporphyria and pyruvate kinase deficiency, a decline in hemopexin concentration occurs in situations when haptoglobin (Hp) concentrations are low or depleted as a result of severe or prolonged hemolysis.[20] Both haptoglobin and hemopexin are acute-phase proteins, the synthesis of which are induced during infection and after inflammatory states to minimize tissue injury and facilitate tissue repair.[9] Hp and hemopexin prevent heme toxicity by binding themselves to heme prior to monocyte or macrophage's arrivals and ensuing clearances,[9] which may explain their effects on outcome in several diseases, and underlies the rationale for exogenous haptoglobin and hemopexin as therapeutic proteins in hemolytic or hemorrhagic conditions.[26] Hemopexin is the major vehicle for the transportation of heme in the plasma.[9]
See also
[edit]References
[edit]- ^ a b c GRCh38: Ensembl release 89: ENSG00000110169 – Ensembl, May 2017
- ^ a b c GRCm38: Ensembl release 89: ENSMUSG00000030895 – Ensembl, May 2017
- ^ "Human PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ "Mouse PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ "Entrez Gene: HPX hemopexin".
- ^ Altruda F, Poli V, Restagno G, Silengo L (1988). "Structure of the human hemopexin gene and evidence for intron-mediated evolution". Journal of Molecular Evolution. 27 (2): 102–8. Bibcode:1988JMolE..27..102A. doi:10.1007/BF02138368. PMID 2842511. S2CID 11271490.
- ^ Altruda F, Poli V, Restagno G, Argos P, Cortese R, Silengo L (June 1985). "The primary structure of human hemopexin deduced from cDNA sequence: evidence for internal, repeating homology". Nucleic Acids Research. 13 (11): 3841–59. doi:10.1093/nar/13.11.3841. PMC 341281. PMID 2989777.
- ^ Bode W (June 1995). "A helping hand for collagenases: the haemopexin-like domain". Structure. 3 (6): 527–30. doi:10.1016/s0969-2126(01)00185-x. PMID 8590012.
- ^ a b c d e Tolosano E, Altruda F (April 2002). "Hemopexin: structure, function, and regulation". DNA and Cell Biology. 21 (4): 297–306. doi:10.1089/104454902753759717. PMID 12042069.
- ^ a b c "Intravascular hemolysis". eClinpath. Retrieved 2019-05-08.
- ^ "Bilirubin and hemolytic anemia". eClinpath. Retrieved 2019-05-08.
- ^ Online Mendelian Inheritance in Man (OMIM): Orthosatic intolerance - 604715
- ^ Takahashi N, Takahashi Y, Putnam FW (January 1985). "Complete amino acid sequence of human hemopexin, the heme-binding protein of serum". Proceedings of the National Academy of Sciences of the United States of America. 82 (1): 73–7. Bibcode:1985PNAS...82...73T. doi:10.1073/pnas.82.1.73. PMC 396973. PMID 3855550.
- ^ Online Mendelian Inheritance in Man (OMIM): Hemopexin - 142290
- ^ Poli V, Altruda F, Silengo L (1986). "Differential transcriptional pattern of the hemopexin gene". The Italian Journal of Biochemistry. 35 (5): 355–60. PMID 3026994.
- ^ Hvidberg V, Maniecki MB, Jacobsen C, Højrup P, Møller HJ, Moestrup SK (October 2005). "Identification of the receptor scavenging hemopexin-heme complexes". Blood. 106 (7): 2572–9. doi:10.1182/blood-2005-03-1185. PMID 15947085.
- ^ Tolosano E, Altruda F (April 2002). "Hemopexin: structure, function, and regulation". DNA and Cell Biology. 21 (4): 297–306. doi:10.1089/104454902753759717. PMID 12042069.
- ^ Watanabe J, Grijalva V, Hama S, Barbour K, Berger FG, Navab M, Fogelman AM, Reddy ST (July 2009). "Hemoglobin and its scavenger protein haptoglobin associate with apoA-1-containing particles and influence the inflammatory properties and function of high density lipoprotein". The Journal of Biological Chemistry. 284 (27): 18292–301. doi:10.1074/jbc.m109.017202. PMC 2709397. PMID 19433579.
- ^ Krikken JA, Lely AT, Bakker SJ, Borghuis T, Faas MM, van Goor H, Navis G, Bakker WW (March 2013). "Hemopexin activity is associated with angiotensin II responsiveness in humans". Journal of Hypertension. 31 (3): 537–41, discussion 542. doi:10.1097/HJH.0b013e32835c1727. PMID 23254305. S2CID 23501030.
- ^ a b Muller-Eberhard U, Javid J, Liem HH, Hanstein A, Hanna M (November 1968). "Plasma concentrations of hemopexin, haptoglobin and heme in patients with various hemolytic diseases". Blood. 32 (5): 811–5. doi:10.1182/blood.V32.5.811.811. PMID 5687939.
- ^ Hoffbrand A, Moss P, Pettit J (2006). Essential Haematology (5th ed.). Oxford: Blackwell Publishing. p. 60. ISBN 978-1-4051-3649-5.
- ^ a b Mehta NU, Reddy ST (October 2015). "Role of hemoglobin/heme scavenger protein hemopexin in atherosclerosis and inflammatory diseases". Current Opinion in Lipidology. 26 (5): 384–7. doi:10.1097/MOL.0000000000000208. PMC 4826275. PMID 26339767.
- ^ a b Garland P, Durnford AJ, Okemefuna AI, Dunbar J, Nicoll JA, Galea J, Boche D, Bulters DO, Galea I (March 2016). "Heme-Hemopexin Scavenging Is Active in the Brain and Associates With Outcome After Subarachnoid Hemorrhage" (PDF). Stroke. 47 (3): 872–6. doi:10.1161/strokeaha.115.011956. PMID 26768209. S2CID 11532383.
- ^ Ma B, Day JP, Phillips H, Slootsky B, Tolosano E, Doré S (February 2016). "Deletion of the hemopexin or heme oxygenase-2 gene aggravates brain injury following stroma-free hemoglobin-induced intracerebral hemorrhage". Journal of Neuroinflammation. 13: 26. doi:10.1186/s12974-016-0490-1. PMC 4736638. PMID 26831741.
- ^ Liu J, Lane S, Lall R, Russo M, Farrell L, Debreli Coskun M, et al. (December 2022). "Circulating hemopexin modulates anthracycline cardiac toxicity in patients and in mice". Science Advances. 8 (51): eadc9245. Bibcode:2022SciA....8C9245L. doi:10.1126/sciadv.adc9245. PMC 9788780. PMID 36563141.
- ^ Schaer DJ, Vinchi F, Ingoglia G, Tolosano E, Buehler PW (2014). "Haptoglobin, hemopexin, and related defense pathways-basic science, clinical perspectives, and drug development". Frontiers in Physiology. 5: 415. doi:10.3389/fphys.2014.00415. PMC 4211382. PMID 25389409.
Further reading
[edit]- Piccard H, Van den Steen PE, Opdenakker G (April 2007). "Hemopexin domains as multifunctional liganding modules in matrix metalloproteinases and other proteins". Journal of Leukocyte Biology. 81 (4): 870–92. doi:10.1189/jlb.1006629. PMID 17185359. S2CID 16210789.
- Morgan WT, Muller-Eberhard U, Lamola AA (January 1978). "Interaction of rabbit hemopexin with bilirubin". Biochimica et Biophysica Acta (BBA) - Protein Structure. 532 (1): 57–64. doi:10.1016/0005-2795(78)90447-6. PMID 620056.
- Liu HM, Atack JR, Rapoport SI (1989). "Immunohistochemical localization of intracellular plasma proteins in the human central nervous system". Acta Neuropathologica. 78 (1): 16–21. doi:10.1007/BF00687397. PMID 2735186. S2CID 24415663.
- Smith A, Tatum FM, Muster P, Burch MK, Morgan WT (April 1988). "Importance of ligand-induced conformational changes in hemopexin for receptor-mediated heme transport". The Journal of Biological Chemistry. 263 (11): 5224–9. doi:10.1016/S0021-9258(18)60703-3. PMID 2833500.
- Altruda F, Poli V, Restagno G, Silengo L (1988). "Structure of the human hemopexin gene and evidence for intron-mediated evolution". Journal of Molecular Evolution. 27 (2): 102–8. Bibcode:1988JMolE..27..102A. doi:10.1007/BF02138368. PMID 2842511. S2CID 11271490.
- Altruda F, Poli V, Restagno G, Argos P, Cortese R, Silengo L (June 1985). "The primary structure of human hemopexin deduced from cDNA sequence: evidence for internal, repeating homology". Nucleic Acids Research. 13 (11): 3841–59. doi:10.1093/nar/13.11.3841. PMC 341281. PMID 2989777.
- Taketani S, Kohno H, Naitoh Y, Tokunaga R (June 1987). "Isolation of the hemopexin receptor from human placenta". The Journal of Biological Chemistry. 262 (18): 8668–71. doi:10.1016/S0021-9258(18)47465-0. PMID 3036819.
- Law ML, Cai GY, Hartz JA, Jones C, Kao FT (July 1988). "The hemopexin gene maps to the same location as the beta-globin gene cluster on human chromosome 11". Genomics. 3 (1): 48–52. doi:10.1016/0888-7543(88)90158-9. PMID 3220477.
- Morgan WT, Alam J, Deaciuc V, Muster P, Tatum FM, Smith A (June 1988). "Interaction of hemopexin with Sn-protoporphyrin IX, an inhibitor of heme oxygenase. Role for hemopexin in hepatic uptake of Sn-protoporphyrin IX and induction of mRNA for heme oxygenase". The Journal of Biological Chemistry. 263 (17): 8226–31. doi:10.1016/S0021-9258(18)68467-4. PMID 3372522.
- Takahashi N, Takahashi Y, Putnam FW (January 1985). "Complete amino acid sequence of human hemopexin, the heme-binding protein of serum". Proceedings of the National Academy of Sciences of the United States of America. 82 (1): 73–7. Bibcode:1985PNAS...82...73T. doi:10.1073/pnas.82.1.73. PMC 396973. PMID 3855550.
- Takahashi N, Takahashi Y, Putnam FW (April 1984). "Structure of human hemopexin: O-glycosyl and N-glycosyl sites and unusual clustering of tryptophan residues". Proceedings of the National Academy of Sciences of the United States of America. 81 (7): 2021–5. Bibcode:1984PNAS...81.2021T. doi:10.1073/pnas.81.7.2021. PMC 345428. PMID 6371807.
- Frantíková V, Borvák J, Kluh I, Morávek L (December 1984). "Amino acid sequence of the N-terminal region of human hemopexin". FEBS Letters. 178 (2): 213–6. doi:10.1016/0014-5793(84)80603-1. PMID 6510521. S2CID 45009902.
- Smith A, Alam J, Escriba PV, Morgan WT (April 1993). "Regulation of heme oxygenase and metallothionein gene expression by the heme analogs, cobalt-, and tin-protoporphyrin". The Journal of Biological Chemistry. 268 (10): 7365–71. doi:10.1016/S0021-9258(18)53184-7. PMID 8463269.
- Morris CM, Candy JM, Edwardson JA, Bloxham CA, Smith A (January 1993). "Evidence for the localization of haemopexin immunoreactivity in neurones in the human brain". Neuroscience Letters. 149 (2): 141–4. doi:10.1016/0304-3940(93)90756-B. PMID 8474687. S2CID 24743139.
- Hrkal Z, Kuzelová K, Muller-Eberhard U, Stern R (March 1996). "Hyaluronan-binding properties of human serum hemopexin". FEBS Letters. 383 (1–2): 72–4. doi:10.1016/0014-5793(96)00225-6. PMID 8612795. S2CID 21283343.
- Hunt RC, Hunt DM, Gaur N, Smith A (July 1996). "Hemopexin in the human retina: protection of the retina against heme-mediated toxicity". Journal of Cellular Physiology. 168 (1): 71–80. doi:10.1002/(SICI)1097-4652(199607)168:1<71::AID-JCP9>3.0.CO;2-5. PMID 8647924. S2CID 10105569.
- Miller YI, Smith A, Morgan WT, Shaklai N (October 1996). "Role of hemopexin in protection of low-density lipoprotein against hemoglobin-induced oxidation". Biochemistry. 35 (40): 13112–7. doi:10.1021/bi960737u. PMID 8855948.
- Grinberg LN, O'Brien PJ, Hrkal Z (July 1999). "The effects of heme-binding proteins on the peroxidative and catalatic activities of hemin". Free Radical Biology & Medicine. 27 (1–2): 214–9. doi:10.1016/S0891-5849(99)00082-9. PMID 10443938.
- Nakajima S, Moriyama T, Hayashi H, Sakata I, Nakae Y, Takemura T (February 2000). "Hemopexin as a carrier protein of tumor-localizing Ga-metalloporphyrin-ATN-2". Cancer Letters. 149 (1–2): 221–6. doi:10.1016/S0304-3835(99)00367-5. PMID 10737728.
- Shipulina N, Smith A, Morgan WT (April 2000). "Heme binding by hemopexin: evidence for multiple modes of binding and functional implications". Journal of Protein Chemistry. 19 (3): 239–48. doi:10.1023/A:1007016105813. PMID 10981817. S2CID 45510572.
External links
[edit]- Hemopexin at the U.S. National Library of Medicine Medical Subject Headings (MeSH)