Single-stranded binding protein
| SSB | |||||||||
|---|---|---|---|---|---|---|---|---|---|
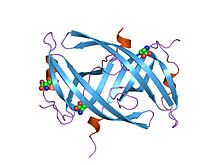 Crystal structure of PriB- a primosomal DNA replication protein of Escherichia coli | |||||||||
| Identifiers | |||||||||
| Symbol | SSB | ||||||||
| Pfam | PF00436 | ||||||||
| Pfam clan | CL0021 | ||||||||
| InterPro | IPR000424 | ||||||||
| PROSITE | PDOC00602 | ||||||||
| SCOP2 | 1kaw / SCOPe / SUPFAM | ||||||||
| TCDB | 3.A.7 | ||||||||
| |||||||||
Single-stranded binding proteins (not to be confused with the E. coli protein, Single-strand DNA-binding protein, SSB) are a class of proteins that have been identified in both viruses and organisms from bacteria to humans.
Function
Single stranded binding proteins prevent premature annealing (binding of complementary DNA sequences), protect the single-stranded DNA from being digested by nucleases, and prevent secondary structure formation, thereby allowing other enzymes to function effectively on the single strand.
Viral SSB
| Viral_DNA_bp | |||||||||
|---|---|---|---|---|---|---|---|---|---|
 Single stranded DNA-binding protein(icp8) from herpes simplex virus-1 | |||||||||
| Identifiers | |||||||||
| Symbol | Viral_DNA_bp | ||||||||
| Pfam | PF00747 | ||||||||
| InterPro | IPR000635 | ||||||||
| |||||||||
Although the overall picture of human cytomegalovirus (HHV-5) DNA synthesis appears typical of the herpesviruses, some novel features are emerging.
Structure
In ICP8, the herpes simplex virus (HSV-1) single-strand DNA-binding protein (ssDNA-binding protein (SSB)), The head consists of the eight alpha helices. The front side of the neck region consists of a five-stranded beta-sheet and two alpha helices, whereas the back side is a three-stranded beta-sheet The shoulder part of the N-terminal domain contains an alpha-helical and beta-sheet region.[1] The herpes simplex virus (HSV-1) SSB, ICP8, is a nuclear protein that, along other replication proteins is required for viral DNA replication during lytic infection.[1]
Mechanism
Six herpes virus-group-common genes encode proteins that likely constitute the replication fork machinery, including a two-subunit DNA polymerase, a Helicase-primase complex and a single-stranded DNA-binding protein.[2] The human herpesvirus 1 (HHV-1) single-strand DNA-binding protein ICP8 is a 128kDa zinc metalloprotein. Photoaffinity labeling has shown that the region encompassing amino acid residues 368-902 contains the single-strand DNA-binding site of ICP8.[3] The HHHV-1 UL5, UL8, and UL52 genes encode an essential heterotrimeric DNA helicase-primase that is responsible for concomitant DNA unwinding and primer synthesis at the viral DNA replication fork. ICP8 may stimulate DNA unwinding and enable bypass of cisplatin damaged DNA by recruiting the helicase-primase to the DNA.[4]
Bacterial SSB
In molecular biology, SSB protein domains in bacteria are important maintaining DNA metabolism, more specifically DNA replication, repair and recombination.[5] It has a structure of three beta-strands to a single six-stranded beta-sheet to form a dimer.[6]
Eukaryotic Replication Protein A
| Replication protein A | |||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| (heterotrimer) | |||||||||||||
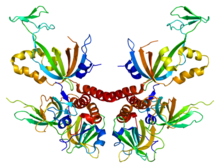 This is an image of human Replication protein A. From PDB: 1L1O Proteopedia protein A Replication protein A | |||||||||||||
| Function | damaged DNA binding, single-stranded DNA binding | ||||||||||||
| |||||||||||||
Replication protein A is the equivalent to SSB in eukaryotic cells.
See also
References
- ^ a b Mapelli M, Panjikar S, Tucker PA (2005). "The crystal structure of the herpes simplex virus 1 ssDNA-binding protein suggests the structural basis for flexible, cooperative single-stranded DNA binding". J Biol Chem. 280 (4): 2990–7. doi:10.1074/jbc.M406780200. PMID 15507432.
{{cite journal}}: CS1 maint: unflagged free DOI (link) - ^ Anders DG, McCue LA (1996). "The human cytomegalovirus genes and proteins required for DNA synthesis". Intervirology. 39 (5–6): 378–88. PMID 9130047.
- ^ White EJ, Boehmer PE (October 1999). "Photoaffinity labeling of the herpes simplex virus type-1 single-strand DNA-binding protein (ICP8) with oligodeoxyribonucleotides". Biochem. Biophys. Res. Commun. 264 (2): 493–7. doi:10.1006/bbrc.1999.1566. PMID 10529391.
- ^ Tanguy Le Gac N, Villani G, Boehmer PE (May 1998). "Herpes simplex virus type-1 single-strand DNA-binding protein (ICP8) enhances the ability of the viral DNA helicase-primase to unwind cisplatin-modified DNA". J. Biol. Chem. 273 (22): 13801–7. doi:10.1074/jbc.273.22.13801. PMID 9593724.
{{cite journal}}: CS1 maint: unflagged free DOI (link) - ^ Meyer RR, Laine PS (December 1990). "The single-stranded DNA-binding protein of Escherichia coli". Microbiol. Rev. 54 (4): 342–80. PMC 372786. PMID 2087220.
- ^ Raghunathan S, Ricard CS, Lohman TM, Waksman G (June 1997). "Crystal structure of the homo-tetrameric DNA binding domain of Escherichia coli single-stranded DNA-binding protein determined by multiwavelength x-ray diffraction on the selenomethionyl protein at 2.9-A resolution". Proc. Natl. Acad. Sci. U.S.A. 94 (13): 6652–7. doi:10.1073/pnas.94.13.6652. PMC 21213. PMID 9192620.
External links
- Single-Stranded+DNA+Binding+Proteins at the U.S. National Library of Medicine Medical Subject Headings (MeSH)
- SSB in PFAM
