Fructose-bisphosphate aldolase
| Fructose-bisphosphate aldolase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
 Fructose-bisphosphate aldolase octamer, Human | |||||||||
| Identifiers | |||||||||
| EC no. | 4.1.2.13 | ||||||||
| CAS no. | 9024-52-6 | ||||||||
| Databases | |||||||||
| IntEnz | IntEnz view | ||||||||
| BRENDA | BRENDA entry | ||||||||
| ExPASy | NiceZyme view | ||||||||
| KEGG | KEGG entry | ||||||||
| MetaCyc | metabolic pathway | ||||||||
| PRIAM | profile | ||||||||
| PDB structures | RCSB PDB PDBe PDBsum | ||||||||
| Gene Ontology | AmiGO / QuickGO | ||||||||
| |||||||||
| Fructose-bisphosphate aldolase class-I | |||||||||
|---|---|---|---|---|---|---|---|---|---|
 fructose 1,6-bisphosphate aldolase from rabbit liver | |||||||||
| Identifiers | |||||||||
| Symbol | Glycolytic | ||||||||
| Pfam | PF00274 | ||||||||
| InterPro | IPR000741 | ||||||||
| PROSITE | PDOC00143 | ||||||||
| SCOP2 | 1ald / SCOPe / SUPFAM | ||||||||
| CDD | cd00344 | ||||||||
| |||||||||
| Fructose-bisphosphate aldolase class-II | |||||||||
|---|---|---|---|---|---|---|---|---|---|
 class II fructose-1,6-bisphosphate aldolase in complex with phosphoglycolohydroxamate | |||||||||
| Identifiers | |||||||||
| Symbol | F_bP_aldolase | ||||||||
| Pfam | PF01116 | ||||||||
| Pfam clan | CL0036 | ||||||||
| InterPro | IPR000771 | ||||||||
| PROSITE | PDOC00523 | ||||||||
| SCOP2 | 1dos / SCOPe / SUPFAM | ||||||||
| CDD | cd00453 | ||||||||
| |||||||||
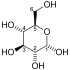
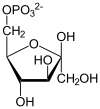
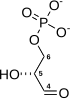
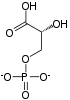
Fructose-bisphosphate aldolase (EC 4.1.2.13), often just aldolase, is an enzyme catalyzing a reversible reaction that splits the aldol, fructose 1,6-bisphosphate, into the triose phosphates dihydroxyacetone phosphate (DHAP) and glyceraldehyde 3-phosphate (G3P). Aldolase can also produce DHAP from other (3S,4R)-ketose 1-phosphates such as fructose 1-phosphate and sedoheptulose 1,7-bisphosphate. Gluconeogenesis and the Calvin cycle, which are anabolic pathways, use the reverse reaction. Glycolysis, a catabolic pathway, uses the forward reaction. Aldolase is divided into two classes by mechanism.
The word aldolase also refers, more generally, to an enzyme that performs an aldol reaction (creating an aldol) or its reverse (cleaving an aldol), such as Sialic acid aldolase, which forms sialic acid. See the list of aldolases.
Mechanism and structure[edit]
Class I proteins form a protonated Schiff base intermediate linking a highly conserved active site lysine with the DHAP carbonyl carbon. Additionally, tyrosine residues are crucial to this mechanism in acting as stabilizing hydrogen acceptors. Class II proteins use a different mechanism which polarizes the carbonyl group with a divalent cation like Zn2+. The Escherichia coli galactitol operon protein, gatY, and N-acetyl galactosamine operon protein, agaY, which are tagatose-bisphosphate aldolase, are homologs of class II fructose-bisphosphate aldolase. Two histidine residues in the first half of the sequence of these homologs have been shown to be involved in binding zinc.[1]
The protein subunits of both classes each have an α/β domain folded into a TIM barrel containing the active site. Several subunits are assembled into the complete protein. The two classes share little sequence identity.
With few exceptions only class I proteins have been found in animals, plants, and green algae.[2] With few exceptions only class II proteins have been found in fungi. Both classes have been found widely in other eukaryotes and in bacteria.[3] The two classes are often present together in the same organism. Plants and algae have plastidal aldolase, sometimes a relic of endosymbiosis, in addition to the usual cytosolic aldolase. A bifunctional fructose-bisphosphate aldolase/phosphatase, with class I mechanism, has been found widely in archaea and in some bacteria.[4] The active site of this archaeal aldolase is also in a TIM barrel.
In gluconeogenesis and glycolysis[edit]
Gluconeogenesis and glycolysis share a series of six reversible reactions. In gluconeogenesis glyceraldehyde-3-phosphate is reduced to fructose 1,6-bisphosphate with aldolase. In glycolysis fructose 1,6-bisphosphate is made into glyceraldehyde-3-phosphate and dihydroxyacetone phosphate through the use of aldolase. The aldolase used in gluconeogenesis and glycolysis is a cytoplasmic protein.
Three forms of class I protein are found in vertebrates. Aldolase A is preferentially expressed in muscle and brain; aldolase B in liver, kidney, and in enterocytes; and aldolase C in brain. Aldolases A and C are mainly involved in glycolysis, while aldolase B is involved in both glycolysis and gluconeogenesis.[5] Some defects in aldolase B cause hereditary fructose intolerance. The metabolism of free fructose in liver exploits the ability of aldolase B to use fructose 1-phosphate as a substrate.[6] Archaeal fructose-bisphosphate aldolase/phosphatase is presumably involved in gluconeogenesis because its product is fructose 6-phosphate.[7]
In the Calvin cycle[edit]
The Calvin cycle is a carbon fixation pathway; it is part of photosynthesis, which convert carbon dioxide and other compounds into glucose. It and gluconeogenesis share a series of four reversible reactions. In both pathways 3-phosphoglycerate (3-PGA or 3-PG) is reduced to fructose 1,6-bisphosphate with aldolase catalyzing the last reaction. A fifth reaction, catalyzed in both pathways by fructose 1,6-bisphosphatase, hydrolyzes the fructose 1-6-bisphosphate to fructose 6-phosphate and inorganic phosphate. The large decrease in free energy makes this reaction irreversible. In the Calvin cycle aldolase also catalyzes the production of sedoheptulose 1,7-bisphosphate from DHAP and erythrose 4-phosphate. The chief products of the Calvin cycle are triose phosphate (TP), which is a mixture of DHAP and G3P, and fructose 6-phosphate. Both are also needed to regenerate RuBP. The aldolase used by plants and algae in the Calvin cycle is usually a plastid-targeted protein encoded by a nuclear gene.
Reactions[edit]
Aldolase catalyzes
- fructose 1,6-bisphosphate ⇌ DHAP + G3P
and also
- sedoheptulose 1,7-bisphosphate ⇌ DHAP + erythrose 4-phosphate
- fructose 1-phosphate ⇌ DHAP + glyceraldehyde
Aldolase is used in the reversible trunk of gluconeogenesis/glycolysis
- 2(PEP + NADH + H+ + ATP + H2O) ⇌ fructose 1,6-bisphosphate + 2(NAD+ + ADP + Pi)
Aldolase is also used in the part of the Calvin cycle shared with gluconeogenesis, with the irreversible phosphate hydrolysis at the end catalyzed by fructose 1,6-bisphosphatase
- 2(3-PG + NADPH + H+ + ATP + H2O) ⇌ fructose 1,6-bisphosphate + 2(NADP+ + ADP + Pi)
- fructose 1,6-bisphosphate + H2O → fructose 6-phosphate + Pi
In gluconeogenesis 3-PG is produced by enolase and phosphoglycerate mutase acting in series
- PEP + H2O ⇌ 2-PG ⇌ 3-PG
In the Calvin cycle 3-PG is produced by RuBisCO
- RuBP + CO2 + H2O → 2(3-PG)
G3P is produced by phosphoglycerate kinase acting in series with glyceraldehyde-3-phosphate dehydrogenase (GAPDH) in gluconeogenesis, and in series with glyceraldehyde-3-phosphate dehydrogenase (NADP+) (phosphorylating) in the Calvin cycle
- 3-PG + ATP ⇌ 1,3-bisphosphoglycerate + ADP
- 1,3-bisphosphoglycerate + NAD(P)H + H+ ⇌ G3P + Pi + NAD(P)+
Triose-phosphate isomerase maintains DHAP and G3P in near equilibrium, producing the mixture called triose phosphate (TP)
- G3P ⇌ DHAP
Thus both DHAP and G3P are available to aldolase.
Moonlighting properties[edit]
Aldolase has also been implicated in many "moonlighting" or non-catalytic functions, based upon its binding affinity for many other proteins including F-actin, α-tubulin, light chain dynein, WASP, Band 3 anion exchanger, phospholipase D (PLD2), glucose transporter GLUT4, inositol trisphosphate, V-ATPase and ARNO (a guanine nucleotide exchange factor of ARF6). These associations are thought to be predominantly involved in cellular structure, however, involvement in endocytosis, parasite invasion, cytoskeleton rearrangement, cell motility, membrane protein trafficking and recycling, signal transduction and tissue compartmentalization have been explored.[8][9][10]
References[edit]
- ^ Zgiby SM, Thomson GJ, Qamar S, Berry A (2000). "Exploring substrate binding and discrimination in fructose1, 6-bisphosphate and tagatose 1,6-bisphosphate aldolases". Eur. J. Biochem. 267 (6): 1858–68. doi:10.1046/j.1432-1327.2000.01191.x. PMID 10712619.
- ^ Patron NJ, Rogers MB, Keeling PJ (2004). "Gene replacement of fructose-1,6-bisphosphate aldolase supports the hypothesis of a single photosynthetic ancestor of chromalveolates". Eukaryotic Cell. 3 (5): 1169–75. doi:10.1128/EC.3.5.1169-1175.2004. PMC 522617. PMID 15470245.
- ^ Trung Hieu Pham, Shreesha Rao, Ta-Chih Cheng, Pei-Chi Wang, Shih-Chu Chen, The moonlighting protein fructose 1,6-bisphosphate aldolase as a potential vaccine candidate against Photobacterium damselae subsp. piscicida in Asian sea bass (Lates calcarifer), Developmental & Comparative Immunology,Volume 124,2021,104187,ISSN 0145-305X,https://doi.org/10.1016/j.dci.2021.104187.
- ^ Siebers B, Brinkmann H, Dörr C, Tjaden B, Lilie H, van der Oost J, Verhees CH (2001). "Archaeal fructose-1,6-bisphosphate aldolases constitute a new family of archaeal type class I aldolase". J. Biol. Chem. 276 (31): 28710–8. doi:10.1074/jbc.M103447200. PMID 11387336.
- ^ Walther EU, Dichgans M, Maricich SM, Romito RR, Yang F, Dziennis S, Zackson S, Hawkes R, Herrup K (1998). "Genomic sequences of aldolase C (Zebrin II) direct lacZ expression exclusively in non-neuronal cells of transgenic mice". Proc. Natl. Acad. Sci. U.S.A. 95 (5): 2615–20. Bibcode:1998PNAS...95.2615W. doi:10.1073/pnas.95.5.2615. PMC 19434. PMID 9482935.
- ^ Gopher A, Vaisman N, Mandel H, Lapidot A (1990). "Determination of fructose metabolic pathways in normal and fructose-intolerant children: a C-13 NMR study using C-13 fructose". Proc. Natl. Acad. Sci. U.S.A. 87 (14): 5449–53. doi:10.1073/pnas.87.14.5449. PMC 54342. PMID 2371280.
- ^ Estelmann S, Hügler M, Eisenreich W, Werner K, Berg IA, Ramos-Vera WH, Say RF, Kockelkorn D, Gad'on N, Fuchs G (2011). "Labeling and enzyme studies of the central carbon metabolism in Metallosphaera sedula". J. Bacteriol. 193 (5): 1191–200. doi:10.1128/JB.01155-10. PMC 3067578. PMID 21169486.
- ^ Rangarajan ES, Park H, Fortin E, Sygusch J, Izard T (2010). "Mechanism of Alolase Control of Sorting Nexin 9 Function in Endocytosis". J. Biol. Chem. 285 (16): 11983–90. doi:10.1074/jbc.M109.092049. PMC 2852936. PMID 20129922.
- ^ Ahn AH, Dziennis S, Hawkes R, Herrup K (1994). "The cloning of zebrin II reveals its identity with aldolase C". Development. 120 (8): 2081–90. doi:10.1242/dev.120.8.2081. PMID 7925012.
- ^ Merkulova M, Hurtado-Lorenzo A, Hosokawa H, Zhuang Z, Brown D, Ausiello DA, Marshansky V (2011). "Aldolase directly interacts with ARNO and modulates cell morphology and acid vesicle distribution". Am J Physiol Cell Physiol. 300 (6): C1442-55. doi:10.1152/ajpcell.00076.2010. PMC 3118619. PMID 21307348.
Further reading[edit]
- Berry A, Marshall KE (February 1993). "Identification of zinc-binding ligands in the class II fructose-1,6-bisphosphate aldolase of Escherichia coli". FEBS Lett. 318 (1): 11–6. doi:10.1016/0014-5793(93)81317-S. PMID 8436219. S2CID 7682431.
- Freemont PS, Dunbar B, Fothergill-Gilmore LA (February 1988). "The complete amino acid sequence of human skeletal-muscle fructose-bisphosphate aldolase". Biochem. J. 249 (3): 779–88. doi:10.1042/bj2490779. PMC 1148774. PMID 3355497.
- Galkin A, Li Z, Li L, Kulakova L, Pal LR, Dunaway-Mariano D, Herzberg O (2009). "Structural insights into the substrate binding and stereoselectivity of giardia fructose-1,6-bisphosphate aldolase". Biochemistry. 48 (14): 3186–96. doi:10.1021/bi9001166. PMC 2666783. PMID 19236002.
- Marsh JJ, Lebherz HG (March 1992). "Fructose-bisphosphate aldolases: an evolutionary history". Trends Biochem. Sci. 17 (3): 110–3. doi:10.1016/0968-0004(92)90247-7. PMID 1412694.
- Perham RN (April 1990). "The fructose-1,6-bisphosphate aldolases: same reaction, different enzymes". Biochem. Soc. Trans. 18 (2): 185–7. doi:10.1042/bst0180185. PMID 2199259.
External links[edit]
 Media related to Fructose-bisphosphate aldolase at Wikimedia Commons
Media related to Fructose-bisphosphate aldolase at Wikimedia Commons- Tolan Laboratory at Boston University