Fructose 1,6-bisphosphatase
| fructose-1,6-bisphosphatase 1 | |||||||
|---|---|---|---|---|---|---|---|
 Fructose-1,6-bisphosphatase and its fructose 2,6-bisphosphate complex. Rendered from PDB 3FBP. | |||||||
| Identifiers | |||||||
| Symbol | FBP1 | ||||||
| Alt. symbols | FBP | ||||||
| NCBI gene | 2203 | ||||||
| HGNC | 3606 | ||||||
| OMIM | 229700 | ||||||
| RefSeq | NM_000507 | ||||||
| UniProt | P09467 | ||||||
| Other data | |||||||
| EC number | 3.1.3.11 | ||||||
| Locus | Chr. 9 q22.3 | ||||||
| |||||||
| Fructose-1-6-bisphosphatase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
 crystal structure of rabbit liver fructose-1,6-bisphosphatase at 2.3 angstrom resolution | |||||||||
| Identifiers | |||||||||
| Symbol | FBPase | ||||||||
| Pfam | PF00316 | ||||||||
| Pfam clan | CL0171 | ||||||||
| InterPro | IPR000146 | ||||||||
| PROSITE | PDOC00114 | ||||||||
| SCOP2 | 1frp / SCOPe / SUPFAM | ||||||||
| |||||||||
| Firmicute fructose-1,6-bisphosphatase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Identifiers | |||||||||
| Symbol | FBPase_2 | ||||||||
| Pfam | PF06874 | ||||||||
| Pfam clan | CL0163 | ||||||||
| InterPro | IPR009164 | ||||||||
| |||||||||
| Fructose-1,6-bisphosphatase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
 crystal structure of fructose-1,6-bisphosphatase | |||||||||
| Identifiers | |||||||||
| Symbol | FBPase_3 | ||||||||
| Pfam | PF01950 | ||||||||
| InterPro | IPR002803 | ||||||||
| SCOP2 | 1umg / SCOPe / SUPFAM | ||||||||
| |||||||||
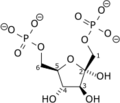
The enzyme fructose bisphosphatase (EC 3.1.3.11; systematic name D-fructose-1,6-bisphosphate 1-phosphohydrolase) catalyses the conversion of fructose-1,6-bisphosphate to fructose 6-phosphate in gluconeogenesis and the Calvin cycle, which are both anabolic pathways:[1][2]
- D-fructose 1,6-bisphosphate + H2O = D-fructose 6-phosphate + phosphate
Phosphofructokinase (EC 2.7.1.11) catalyses the reverse conversion of fructose 6-phosphate to fructose-1,6-bisphosphate, but this is not just the reverse reaction, because the co-substrates are different (and so thermodynamic requirements are not violated). The two enzymes each catalyse the conversion in one direction only, and are regulated by metabolites such as fructose 2,6-bisphosphate so that high activity of one of them is accompanied by low activity of the other. More specifically, fructose 2,6-bisphosphate allosterically inhibits fructose 1,6-bisphosphatase, but activates phosphofructokinase-I. Fructose 1,6-bisphosphatase is involved in many different metabolic pathways and found in most organisms. FBPase requires metal ions for catalysis (Mg2+ and Mn2+ being preferred) and the enzyme is potently inhibited by Li+.
Structure[edit]
The fold of fructose-1,6-bisphosphatase from pigs was noted to be identical to that of inositol-1-phosphatase (IMPase).[3] Inositol polyphosphate 1-phosphatase (IPPase), IMPase and FBPase share a sequence motif (Asp-Pro-Ile/Leu-Asp-Gly/Ser-Thr/Ser) which has been shown to bind metal ions and participate in catalysis. This motif is also found in the distantly-related fungal, bacterial and yeast IMPase homologues. It has been suggested that these proteins define an ancient structurally conserved family involved in diverse metabolic pathways, including inositol signalling, gluconeogenesis, sulphate assimilation and possibly quinone metabolism.[4]
Species distribution[edit]
Three different groups of FBPases have been identified in eukaryotes and bacteria (FBPase I-III).[5] None of these groups have been found in Archaea so far, though a new group of FBPases (FBPase IV) which also show inositol monophosphatase activity has recently been identified in Archaea.[6]
A new group of FBPases (FBPase V) is found in thermophilic archaea and the hyperthermophilic bacterium Aquifex aeolicus.[7] The characterised members of this group show strict substrate specificity for FBP and are suggested to be the true FBPase in these organisms.[7][8] A structural study suggests that FBPase V has a novel fold for a sugar phosphatase, forming a four-layer alpha-beta-beta-alpha sandwich, unlike the more usual five-layered alpha-beta-alpha-beta-alpha arrangement.[8] The arrangement of the catalytic side chains and metal ligands was found to be consistent with the three-metal ion assisted catalysis mechanism proposed for other FBPases.
The fructose 1,6-bisphosphatases found within the Bacillota (low GC Gram-positive bacteria) do not show any significant sequence similarity to the enzymes from other organisms. The Bacillus subtilis enzyme is inhibited by AMP, though this can be overcome by phosphoenolpyruvate, and is dependent on Mn(2+).[9][10] Mutants lacking this enzyme are apparently still able to grow on gluconeogenic growth substrates such as malate and glycerol.
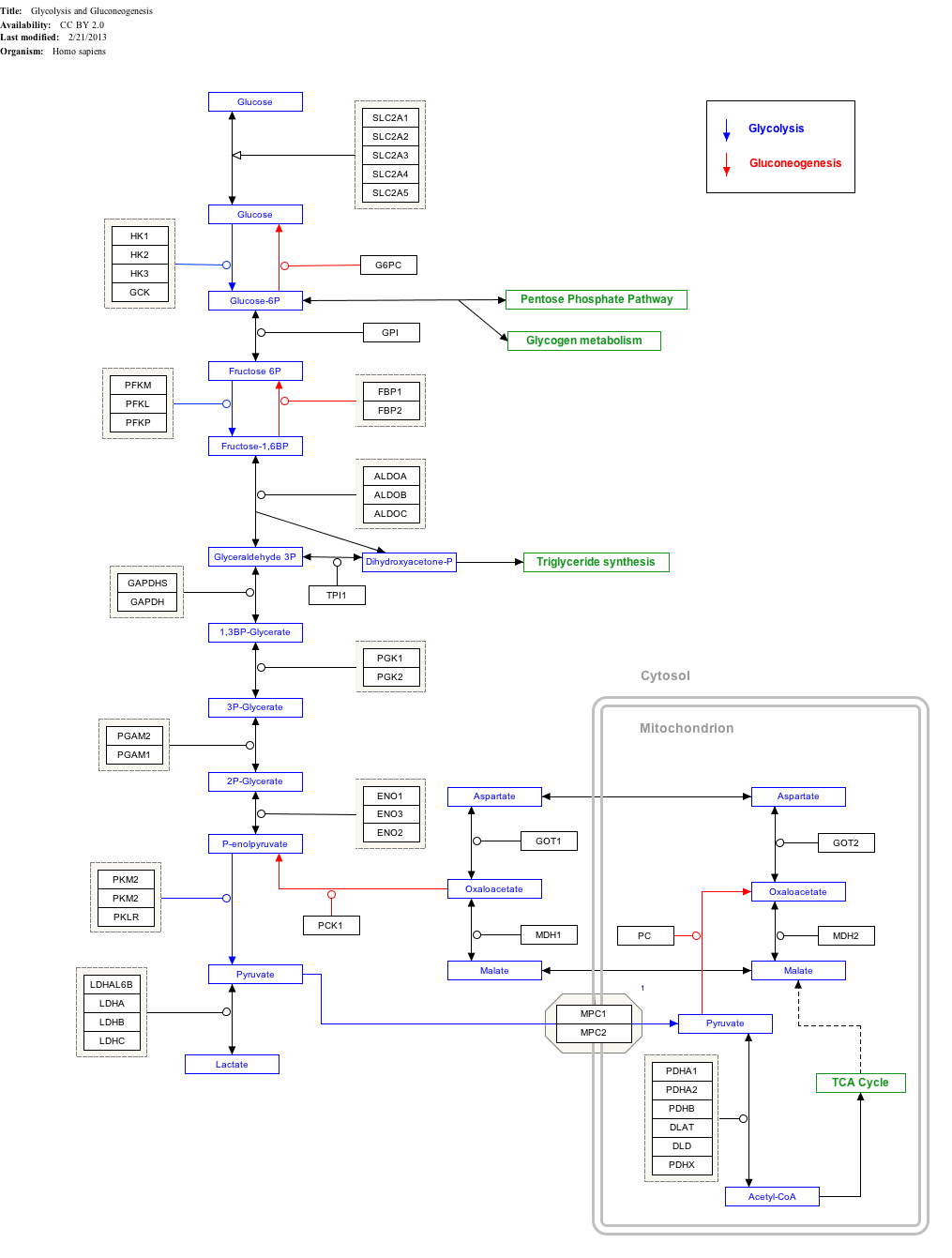
Interactive pathway map[edit]
Click on genes, proteins and metabolites below to link to respective articles.[§ 1]
- ^ The interactive pathway map can be edited at WikiPathways: "GlycolysisGluconeogenesis_WP534".
Hibernation and cold adaptation[edit]
Fructose 1,6-bisphosphatase also plays a key role in hibernation, which requires strict regulation of metabolic processes to facilitate entry into hibernation, maintenance, arousal from hibernation, and adjustments to allow long-term dormancy.[11][12][13] During hibernation, an animal's metabolic rate may decrease to around 1/25 of its euthermic resting metabolic rate.[12][13][14] FBPase is modified in hibernating animals to be much more temperature sensitive than it is in euthermic animals.[11][13][14] FBPase in the liver of a hibernating bat showed a 75% decrease in Km for its substrate FBP at 5 °C than at 37 °C.[11] However, in a euthermic bat this decrease was only 25%, demonstrating the difference in temperature sensitivity between hibernating and euthermic bats.[11] When sensitivity to allosteric inhibitors such as AMP, ADP, inorganic phosphate, and fructose-2,6-bisphosphate were examined, FBPase from hibernating bats was much more sensitive to inhibitors at low temperature than in euthermic bats.[11][15][16]
During hibernation, respiration also dramatically decreases, resulting in conditions of relative anoxia in the tissues. Anoxic conditions inhibit gluconeogenesis, and therefore FBPase, while stimulating glycolysis, and this is another reason for reduced FBPase activity in hibernating animals.[17] The substrate of FBPase, fructose 1,6-bisphosphate, has also been shown to activate pyruvate kinase in glycolysis, linking increased glycolysis to decreased gluconeogenesis when FBPase activity is decreased during hibernation.[13]
In addition to hibernation, there is evidence that FBPase activity varies significantly between warm and cold seasons even for animals that do not hibernate.[18] In rabbits exposed to cold temperatures, FBPase activity decreased throughout the duration of cold exposure, increasing when temperatures became warmer again.[18] The mechanism of this FBPase inhibition is thought to be digestion of FBPase by lysosomal proteases, which are released at higher levels during colder periods.[18] Inhibition of FBPase through proteolytic digestion decreases gluconeogenesis relative to glycolysis during cold periods, similar to hibernation.[18]
Fructose 1,6-bisphosphate aldolase is another temperature dependent enzyme that plays an important role in the regulation of glycolysis and gluconeogenesis during hibernation.[14] Its main role is in glycolysis instead of gluconeogenesis, but its substrate is the same as FBPase's, so its activity affects that of FBPase in gluconeogenesis. Aldolase shows similar changes in activity to FBPase at colder temperatures, such as an upward shift in optimum pH at colder temperatures. This adaptation allows enzymes such as FBPase and fructose-1,6-bisphosphate aldolase to track intracellular pH changes in hibernating animals and match their activity ranges to these shifts.[14] Aldolase also complements the activity of FBPase in anoxic conditions (discussed above) by increasing glycolytic output while FBPase inhibition decreases gluconeogenesis activity.[19]
Diabetes[edit]

Fructose 1,6-bisphosphatase is also a key player in treating type 2 diabetes. In this disease, hyperglycemia causes many serious problems, and treatments often focus on lowering blood sugar levels.[20][21][22] Gluconeogenesis in the liver is a major cause of glucose overproduction in these patients, and so inhibition of gluconeogenesis is a reasonable way to treat type 2 diabetes. FBPase is a good enzyme to target in the gluconeogenesis pathway because it is rate-limiting and controls the incorporation of all three-carbon substrates into glucose but is not involved in glycogen breakdown and is removed from mitochondrial steps in the pathway.[20][21][22] This means that altering its activity can have a large effect on gluconeogenesis while reducing the risk of hypoglycemia and other potential side effects from altering other enzymes in gluconeogenesis.[20][21]
Drug candidates have been developed that mimic the inhibitory activity of AMP on FBPase.[20][22] Efforts were made to mimic the allosteric inhibitory effects of AMP while making the drug as structurally different from it as possible.[22] Second-generation FBPase inhibitors have now been developed and have had good results in clinical trials with non-human mammals and now humans.[20][23]
See also[edit]
References[edit]
- ^ Marcus F, Harrsch PB (May 1990). "Amino acid sequence of spinach chloroplast fructose-1,6-bisphosphatase". Archives of Biochemistry and Biophysics. 279 (1): 151–7. doi:10.1016/0003-9861(90)90475-E. PMID 2159755.
- ^ Marcus F, Gontero B, Harrsch PB, Rittenhouse J (Mar 1986). "Amino acid sequence homology among fructose-1,6-bisphosphatases". Biochemical and Biophysical Research Communications. 135 (2): 374–81. doi:10.1016/0006-291X(86)90005-7. PMID 3008716.
- ^ Zhang Y, Liang JY, Lipscomb WN (Feb 1993). "Structural similarities between fructose-1,6-bisphosphatase and inositol monophosphatase". Biochemical and Biophysical Research Communications. 190 (3): 1080–3. doi:10.1006/bbrc.1993.1159. PMID 8382485.
- ^ York JD, Ponder JW, Majerus PW (May 1995). "Definition of a metal-dependent/Li+-inhibited phosphomonoesterase protein family based upon a conserved three-dimensional core structure". Proceedings of the National Academy of Sciences of the United States of America. 92 (11): 5149–53. Bibcode:1995PNAS...92.5149Y. doi:10.1073/pnas.92.11.5149. PMC 41866. PMID 7761465.
- ^ Donahue JL, Bownas JL, Niehaus WG, Larson TJ (Oct 2000). "Purification and characterization of glpX-encoded fructose 1, 6-bisphosphatase, a new enzyme of the glycerol 3-phosphate regulon of Escherichia coli". Journal of Bacteriology. 182 (19): 5624–7. doi:10.1128/jb.182.19.5624-5627.2000. PMC 111013. PMID 10986273.
- ^ Stec B, Yang H, Johnson KA, Chen L, Roberts MF (Nov 2000). "MJ0109 is an enzyme that is both an inositol monophosphatase and the 'missing' archaeal fructose-1,6-bisphosphatase". Nature Structural Biology. 7 (11): 1046–50. doi:10.1038/80968. PMID 11062561. S2CID 7617099.
- ^ a b Rashid N, Imanaka H, Kanai T, Fukui T, Atomi H, Imanaka T (Aug 2002). "A novel candidate for the true fructose-1,6-bisphosphatase in archaea". The Journal of Biological Chemistry. 277 (34): 30649–55. doi:10.1074/jbc.M202868200. PMID 12065581.
- ^ a b Nishimasu H, Fushinobu S, Shoun H, Wakagi T (Jun 2004). "The first crystal structure of the novel class of fructose-1,6-bisphosphatase present in thermophilic archaea". Structure. 12 (6): 949–59. doi:10.1016/j.str.2004.03.026. PMID 15274916.
- ^ Fujita Y, Freese E (Jun 1979). "Purification and properties of fructose-1,6-bisphosphatase of Bacillus subtilis". The Journal of Biological Chemistry. 254 (12): 5340–9. doi:10.1016/S0021-9258(18)50601-3. PMID 221467.
- ^ Fujita Y, Yoshida K, Miwa Y, Yanai N, Nagakawa E, Kasahara Y (Aug 1998). "Identification and expression of the Bacillus subtilis fructose-1, 6-bisphosphatase gene (fbp)". Journal of Bacteriology. 180 (16): 4309–13. doi:10.1128/JB.180.16.4309-4313.1998. PMC 107433. PMID 9696785.
- ^ a b c d e Storey KB (December 1997). "Metabolic regulation in mammalian hibernation: enzyme and protein adaptations". Comparative Biochemistry and Physiology A. 118 (4): 1115–24. doi:10.1016/S0300-9629(97)00238-7. PMID 9505421.
- ^ a b Heldmaier G, Ortmann S, Elvert R (August 2004). "Natural hypometabolism during hibernation and daily torpor in mammals". Respiratory Physiology & Neurobiology. 141 (3): 317–29. doi:10.1016/j.resp.2004.03.014. PMID 15288602. S2CID 32940046.
- ^ a b c d Brooks SP, Storey KB (January 1992). "Mechanisms of glycolytic control during hibernation in the ground squirrel Spermophilus lateralis". Journal of Comparative Physiology B. 162 (1): 23–28. doi:10.1007/BF00257932. S2CID 1881399.
- ^ a b c d MacDonald JA, Storey KB (December 2002). "Purification and characterization of fructose bisphosphate aldolase from the ground squirrel, Spermophilus lateralis: enzyme role in mammalian hibernation". Archives of Biochemistry and Biophysics. 408 (2): 279–85. doi:10.1016/S0003-9861(02)00579-9. PMID 12464282.
- ^ Ekdahl KN, Ekman P (February 1984). "The effect of fructose 2,6-bisphosphate and AMP on the activity of phosphorylated and unphosphorylated fructose-1,6-bisphosphatase from rat liver". FEBS Letters. 167 (2): 203–9. doi:10.1016/0014-5793(84)80127-1. PMID 6321241. S2CID 22515761.
- ^ Taketa K, Pogell BM (February 1965). "Allosteric Inhibition of Rat Liver Fructose 1,6-Diphosphatase by Adenosine 5'-Monophosphate". The Journal of Biological Chemistry. 240 (2): 651–62. doi:10.1016/S0021-9258(17)45224-0. PMID 14275118.
- ^ Underwood AH, Newsholme EA (July 1967). "Control of glycolysis and gluconeogenesis in rat kidney cortex slices". The Biochemical Journal. 104 (1): 300–5. doi:10.1042/bj1040300. PMC 1270577. PMID 4292000.
- ^ a b c d Fischer EH, Krebs EG, Neurath H, Stadtman ER, eds. (1974). Metabolic Interconversion of Enzymes 1973 Third International Symposium held in Seattle, June 5-8, 1973. Berlin, Heidelberg: Springer. ISBN 978-3-642-80817-3.
- ^ Dawson NJ, Biggar KK, Storey KB (2013). "Characterization of fructose-1,6-bisphosphate aldolase during anoxia in the tolerant turtle, Trachemys scripta elegans: an assessment of enzyme activity, expression and structure". PLOS ONE. 8 (7): e68830. Bibcode:2013PLoSO...868830D. doi:10.1371/journal.pone.0068830. PMC 3715522. PMID 23874782.
- ^ a b c d e Dang Q, Van Poelje PD, Erion MD (2012). "Chapter 11: The Discovery and Development of MB07803, a Second-Generation Fructose-1,6-bisphosphatase Inhibitor with Improved Pharmacokinetic Properties, as a Potential Treatment of Type 2 Diabetes". In Jones RM (ed.). New Therapeutic Strategies for Type 2 Diabetes: Small Molecule Approaches. Cambridge: The Royal Society of Chemistry. doi:10.1039/9781849735322-00306 (inactive 2024-04-11). ISBN 978-1-84973-414-1.
{{cite book}}: CS1 maint: DOI inactive as of April 2024 (link) - ^ a b c Arch JR (2011). "Thermogenesis and Related Metabolic Targets in Anti-Diabetic Therapy". In Schwanstecher M (ed.). Diabetes - Perspectives in Drug Therapy (1st ed.). Berlin, Heidelberg: Springer. p. 203. ISBN 978-3-642-17214-4.
- ^ a b c d van Poelje PD, Potter SC, Chandramouli VC, Landau BR, Dang Q, Erion MD (June 2006). "Inhibition of fructose 1,6-bisphosphatase reduces excessive endogenous glucose production and attenuates hyperglycemia in Zucker diabetic fatty rats". Diabetes. 55 (6): 1747–54. doi:10.2337/db05-1443. PMID 16731838. S2CID 19832318.
- ^ Kaur R, Dahiya L, Kumar M (December 2017). "Fructose-1,6-bisphosphatase inhibitors: A new valid approach for management of type 2 diabetes mellitus". European Journal of Medicinal Chemistry. 141: 473–505. doi:10.1016/j.ejmech.2017.09.029. PMID 29055870.
Further reading[edit]
- Berg JM, Tymoczko JL, Stryer L (2002). "Glycolysis and Gluconeogenesis". In Susan Moran (ed.). Biochemistry (5th ed.). New York: W. H. Freeman and Company. ISBN 0-7167-3051-0.
External links[edit]
- Fructose-1,6-Biphosphatase at the U.S. National Library of Medicine Medical Subject Headings (MeSH)