Synthetic biology

| Part of a series of articles on |
| Synthetic biology |
|---|
| Synthetic biological circuits |
| Genome editing |
| Artificial cells |
| Xenobiology |
| Other topics |
Synthetic biology (SynBio) is a multidisciplinary field of science that focuses on living systems and organisms, and it applies engineering principles to develop new biological parts, devices, and systems or to redesign existing systems found in nature.[1]
It is a branch of science that encompasses a broad range of methodologies from various disciplines, such as biochemistry, biotechnology, biomaterials, material science/engineering, genetic engineering, molecular biology, molecular engineering, systems biology, membrane science, biophysics, chemical and biological engineering, electrical and computer engineering, control engineering and evolutionary biology.
It includes designing and constructing biological modules, biological systems, and biological machines, or re-designing existing biological systems for useful purposes.[2]
Additionally, it is the branch of science that focuses on the new abilities of engineering into existing organisms to redesign them for useful purposes.[3]
In order to produce predictable and robust systems with novel functionalities that do not already exist in nature, it is also necessary to apply the engineering paradigm of systems design to biological systems. According to the European Commission, this possibly involves a molecular assembler based on biomolecular systems such as the ribosome.[4]
History
[edit]1910: First identifiable use of the term synthetic biology in Stéphane Leduc's publication Théorie physico-chimique de la vie et générations spontanées.[5] He also noted this term in another publication, La Biologie Synthétique in 1912.[6]
1944: Canadian-American scientist Oswald Avery shows that DNA is the material of which genes and chromosomes are made. This becomes the bedrock on which all subsequent genetic research is built.[7]
1953: Francis Crick and James Watson publish the structure of the DNA in Nature.
1961: Jacob and Monod postulate cellular regulation by molecular networks from their study of the lac operon in E. coli and envisioned the ability to assemble new systems from molecular components.[8]
1973: First molecular cloning and amplification of DNA in a plasmid is published in P.N.A.S. by Cohen, Boyer et al. constituting the dawn of synthetic biology.[9]
1978: Arber, Nathans and Smith win the Nobel Prize in Physiology or Medicine for the discovery of restriction enzymes, leading Szybalski to offer an editorial comment in the journal Gene:
The work on restriction nucleases not only permits us easily to construct recombinant DNA molecules and to analyze individual genes, but also has led us into the new era of synthetic biology where not only existing genes are described and analyzed but also new gene arrangements can be constructed and evaluated.[10]
1988: First DNA amplification by the polymerase chain reaction (PCR) using a thermostable DNA polymerase is published in Science by Mullis et al.[11] This obviated adding new DNA polymerase after each PCR cycle, thus greatly simplifying DNA mutagenesis and assembly.
2000: Two papers in Nature report synthetic biological circuits, a genetic toggle switch and a biological clock, by combining genes within E. coli cells.[12][13]
2003: The most widely used standardized DNA parts, BioBrick plasmids, are invented by Tom Knight.[14] These parts will become central to the International Genetically Engineered Machine (iGEM) competition founded at MIT in the following year.
2003: Researchers engineer an artemisinin precursor pathway in E. coli.[15]
2004: First international conference for synthetic biology, Synthetic Biology 1.0 (SB1.0) is held at MIT.
2005: Researchers develop a light-sensing circuit in E. coli.[16] Another group designs circuits capable of multicellular pattern formation.[17]
2006: Researchers engineer a synthetic circuit that promotes bacterial invasion of tumour cells.[18]
2010: Researchers publish in Science the first synthetic bacterial genome, called M. mycoides JCVI-syn1.0.[19][20] The genome is made from chemically-synthesized DNA using yeast recombination.
2011: Functional synthetic chromosome arms are engineered in yeast.[21]
2012: Charpentier and Doudna labs publish in Science the programming of CRISPR-Cas9 bacterial immunity for targeting DNA cleavage.[22] This technology greatly simplified and expanded eukaryotic gene editing.
2019: Scientists at ETH Zurich report the creation of the first bacterial genome, named Caulobacter ethensis-2.0, made entirely by a computer, although a related viable form of C. ethensis-2.0 does not yet exist.[23][24]
2019: Researchers report the production of a new synthetic (possibly artificial) form of viable life, a variant of the bacteria Escherichia coli, by reducing the natural number of 64 codons in the bacterial genome to 59 codons instead, in order to encode 20 amino acids.[25][26]
2020: Scientists created the first xenobot, a programmable synthetic organism derived from frog cells and designed by AI.[27]
2021: Scientists reported that xenobots are able to self-replicate by gathering loose cells in the environment and then forming new xenobot.[28]
Perspectives
[edit]It is a field whose scope is expanding in terms of systems integration, engineered organisms, and practical findings.[1]
Engineers view biology as technology (in other words, a given system includes biotechnology or its biological engineering).[29] Synthetic biology includes the broad redefinition and expansion of biotechnology, with the ultimate goal of being able to design and build engineered live biological systems that process information, manipulate chemicals, fabricate materials and structures, produce energy, provide food, and maintain and enhance human health, as well as advance fundamental knowledge of biological systems and our environment.[30]
Researchers and companies working in synthetic biology are using nature's power to solve issues in agriculture, manufacturing, and medicine.[3]
Due to more powerful genetic engineering capabilities and decreased DNA synthesis and sequencing costs, the field of synthetic biology is rapidly growing. In 2016, more than 350 companies across 40 countries were actively engaged in synthetic biology applications; all these companies had an estimated net worth of $3.9 billion in the global market.[31] Synthetic biology currently has no generally accepted definition. Here are a few examples:
It is the science of emerging genetic and physical engineering to produce new (and, therefore, synthetic) life forms. To develop organisms with novel or enhanced characteristics, this emerging field of study combines biology, engineering, and related disciplines' knowledge and techniques to design chemically synthesised DNA.[32][33]
Biomolecular engineering includes approaches that aim to create a toolkit of functional units that can be introduced to present new technological functions in living cells. Genetic engineering includes approaches to construct synthetic chromosomes or minimal organisms like Mycoplasma laboratorium.
Biomolecular design refers to the general idea of de novo design and additive combination of biomolecular components. Each of these approaches shares a similar task: to develop a more synthetic entity at a higher level of complexity by inventively manipulating a simpler part at the preceding level.[34][35] Optimizing these exogenous pathways in unnatural systems takes iterative fine-tuning of the individual biomolecular components to select the highest concentrations of the desired product.[36]
On the other hand, "re-writers" are synthetic biologists interested in testing the irreducibility of biological systems. Due to the complexity of natural biological systems, it would be simpler to rebuild the natural systems of interest from the ground up; to provide engineered surrogates that are easier to comprehend, control and manipulate.[37] Re-writers draw inspiration from refactoring, a process sometimes used to improve computer software.
Categories
[edit]Bioengineering, synthetic genomics, protocell synthetic biology, unconventional molecular biology, and in silico techniques are the five categories of synthetic biology.[38]
It is necessary to review the distinctions and analogies between the categories of synthetic biology for its social and ethical assessment, to distinguish between issues affecting the whole field and particular to a specific one.[38]
Bioengineering
[edit]The subfield of bioengineering concentrates on creating novel metabolic and regulatory pathways, and is currently the one that likely draws the attention of most researchers and funding. It is primarily motivated by the desire to establish biotechnology as a legitimate engineering discipline. When referring to this area of synthetic biology, the word "bioengineering" should not be confused with "traditional genetic engineering", which involves introducing a single transgene into the intended organism. Bioengineers adapted synthetic biology to provide a substantially more integrated perspective on how to alter organisms or metabolic systems.[38]
A typical example of single-gene genetic engineering is the insertion of the human insulin gene into bacteria to create transgenic proteins. The creation of whole new signalling pathways, containing numerous genes and regulatory components (such as an oscillator circuit to initiate the periodic production of green fluorescent protein (GFP) in mammalian cells), is known as bioengineering as part of synthetic biology.[38]
By utilising simplified and abstracted metabolic and regulatory modules as well as other standardized parts that may be freely combined to create new pathways or creatures, bioengineering aims to create innovative biological systems. In addition to creating infinite opportunities for novel applications, this strategy is anticipated to make bioengineering more predictable and controllable than traditional biotechnology.[38]
Synthetic genomics
[edit]The formation of animals with a chemically manufactured (minimal) genome is another facet of synthetic biology that is highlighted by synthetic genomics. This area of synthetic biology has been made possible by ongoing advancements in DNA synthesis technology, which now makes it feasible to produce DNA molecules with thousands of base pairs at a reasonable cost. The goal is to combine these molecules into complete genomes and transplant them into living cells, replacing the host cell's genome and reprogramming its metabolism to perform different functions.[38]
Scientists have previously demonstrated the potential of this approach by creating infectious viruses by synthesising the genomes of multiple viruses. These significant advances in science and technology triggered the initial public concerns concerning the risks associated with this technology.[38]
A simple genome might also work as a "chassis genome" that could be enlarged quickly by gene inclusion created for particular tasks. Such "chassis creatures" would be more suited for the insertion of new functions than wild organisms since they would have fewer biological pathways that could potentially conflict with the new functionalities in addition to having specific insertion sites. Synthetic genomics strives to create creatures with novel "architectures," much like the bioengineering method. It adopts an integrative or holistic perspective of the organism. In this case, the objective is the creation of chassis genomes based on necessary genes and other required DNA sequences rather than the design of metabolic or regulatory pathways based on abstract criteria.[38]
Protocell synthetic biology
[edit]The in vitro generation of synthetic cells is the protocell branch of synthetic biology. Lipid vesicles, which have all the necessary components to function as a complete system, can be used to create these artificial cells. In the end, these synthetic cells should meet the requirements for being deemed alive, namely the capacity for self-replication, self-maintenance, and evolution. The protocell technique has this as its end aim, however there are other intermediary steps that fall short of meeting all the criteria for a living cell. In order to carry out a specific function, these lipid vesicles contain cell extracts or more specific sets of biological macromolecules and complex structures, such as enzymes, nucleic acids, or ribosomes. For instance, liposomes may carry out particular polymerase chain reactions or synthesise a particular protein.[38]
Protocell synthetic biology takes artificial life one step closer to reality by eventually synthesizing not only the genome but also every component of the cell in vitro, as opposed to the synthetic genomics approach, which relies on coercing a natural cell to carry out the instructions encoded by the introduced synthetic genome. Synthetic biologists in this field view their work as basic study into the conditions necessary for life to exist and its origin more than in any of the other techniques. The protocell technique, however, also lends itself well to applications; similar to other synthetic biology byproducts, protocells could be employed for the manufacture of biopolymers and medicines.[38]
Unconventional molecular biology
[edit]The objective of the "unnatural molecular biology" strategy is to create new varieties of life that are based on a different kind of molecular biology, such as new types of nucleic acids or a new genetic code. The creation of new types of nucleotides that can be built into unique nucleic acids could be accomplished by changing certain DNA or RNA constituents, such as the bases or the backbone sugars.[38]
The normal genetic code is being altered by inserting quadruplet codons or changing some codons to encode new amino acids, which would subsequently permit the use of non-natural amino acids with unique features in protein production. It is a scientific and technological problem to adjust the enzymatic machinery of the cell for both approaches.[38]
A new sort of life would be formed by organisms with a genome built on synthetic nucleic acids or on a totally new coding system for synthetic amino acids. This new style of life would have some benefits but also some new dangers. On release into the environment, there would be no horizontal gene transfer or outcrossing of genes with natural species. Furthermore, these kinds of synthetic organisms might be created to require non-natural materials for protein or nucleic acid synthesis, rendering them unable to thrive in the wild if they accidentally escaped.[38]
On the other hand, if these organisms ultimately were able to survive outside of controlled space, they might have a particular benefit over natural organisms because they would be resistant to predatory living organisms or natural viruses, that could lead to an unmanaged spread of the synthetic organisms.[38]
In silico technique
[edit]Synthetic biology in silico and the various strategies are interconnected. The development of complex designs, whether they are metabolic pathways, fundamental cellular processes, or chassis genomes, is one of the major difficulties faced by the four synthetic-biology methods outlined above. Because of this, synthetic biology has a robust in silico branch, similar to systems biology, that aims to create computational models for the design of common biological components or synthetic circuits, which are essentially simulations of synthetic organisms.[38]
The practical application of simulations and models through bioengineering or other fields of synthetic biology is the long-term goal of in silico synthetic biology. Many of the computational simulations of synthetic organisms up to this point possess little to no direct analogy to living things. Due to this, in silico synthetic biology is regarded as a separate group in this article.[38]
It is sensible to integrate the five areas under the umbrella of synthetic biology as an unified area of study. Even though they focus on various facets of life, such as metabolic regulation, essential elements, or biochemical makeup, these five strategies all work toward the same end: creating new types of living organisms. Additionally, the varied methodologies begin with numerous methodological approaches, which leads to the diversity of synthetic biology approaches.[38]
Synthetic biology is an interdisciplinary field that draws from and is inspired by many different scientific disciplines, not one single field or technique. Synthetic biologists all have the same underlying objective of designing and producing new forms of life, despite the fact that they may employ various methodologies, techniques, and research instruments. Any evaluation of synthetic biology, whether it examines ethical, legal, or safety considerations, must take into account the fact that while some questions, risks, and issues are unique to each technique, in other circumstances, synthetic biology as a whole must be taken into consideration.[38]
Four engineering approaches
[edit]Synthetic biology has traditionally been divided into four different engineering approaches: top down, parallel, orthogonal and bottom up.[39]
To replicate emergent behaviours from natural biology and build artificial life, unnatural chemicals are used. The other looks for interchangeable components from biological systems to put together and create systems that do not work naturally. In either case, a synthetic objective compels researchers to venture into new area in order to engage and resolve issues that cannot be readily resolved by analysis. Due to this, new paradigms are driven to arise in ways that analysis cannot easily do. In addition to equipments that oscillate, creep, and play tic-tac-toe, synthetic biology has produced diagnostic instruments that enhance the treatment of patients with infectious diseases.[40]
Top-down approach
[edit]It involves using metabolic and genetic engineering techniques to impart new functions to living cells.[41] By comparing universal genes and eliminating non-essential ones to create a basic genome, this method seeks to lessen the complexity of existing cells. These initiatives are founded on the hypothesis of a single genesis for cellular life, the so-called Last Universal Common Ancestor, which supports the presence of a universal minimal genome that gave rise to all living things. Recent studies, however, raise the possibility that the eukaryotic and prokaryotic cells that make up the tree of life may have evolved from a group of primordial cells rather than from a single cell. As a result, even while the Holy Grail-like pursuit of the "minimum genome" has grown elusive, cutting out a number of non-essential functions impairs an organism's fitness and leads to "fragile" genomes.[39]
Bottom-up approach
[edit]This approach involves creating new biological systems in vitro by bringing together 'non-living' biomolecular components,[42] often with the aim of constructing an artificial cell.
Reproduction, replication, and assembly are three crucial self-organizational principles that are taken into account in order to accomplish this. Cells, which are made up of a container and a metabolism, are considered "hardware" in the definition of reproduction, whereas replication occurs when a system duplicates a perfect copy of itself, as in the case of DNA, which is considered "software." When vesicles or containers (such as Oparin's coacervates) formed of tiny droplets of molecules that are organic like lipids or liposomes, membrane-like structures comprising phospholipids, aggregate, assembly occur.[39]
The study of protocells exists along with other in vitro synthetic biology initiatives that seek to produce minimum cells, metabolic pathways, or "never-born proteins" as well as to mimic physiological functions including cell division and growth. The in vitro enhancement of synthetic pathways does have the potential to have an effect on some other synthetic biology sectors, including metabolic engineering, despite the fact that it no longer classified as synthetic biology research. This research, which is primarily essential, deserves proper recognition as synthetic biology research.[39]
Parallel approach
[edit]Parallel engineering is also known as bioengineering. The basic genetic code is the foundation for parallel engineering research, which uses conventional biomolecules like nucleic acids and the 20 amino acids to construct biological systems. For a variety of applications in biocomputing, bioenergy, biofuels, bioremediation, optogenetics, and medicine, it involves the standardisation of DNA components, engineering of switches, biosensors, genetic circuits, logic gates, and cellular communication operators. For directing the expression of two or more genes and/or proteins, the majority of these applications often rely on the use of one or more vectors (or plasmids). Small, circular, double-strand DNA units known as plasmids, which are primarily found in prokaryotic but can also occasionally be detected in eukaryotic cells, may replicate autonomously of chromosomal DNA.[39]
Orthogonal approach
[edit]It is also known as perpendicular engineering. This strategy, also referred to as "chemical synthetic biology," principally seeks to alter or enlarge the genetic codes of living systems utilising artificial DNA bases and/or amino acids. This subfield is also connected to xenobiology, a newly developed field that combines systems chemistry, synthetic biology, exobiology, and research into the origins of life. In recent decades, researchers have created compounds that are structurally similar to the DNA canonical bases to see if those "alien" or xeno (XNA) molecules may be employed as genetic information carriers. Similar to this, noncanonical moieties have taken the place of the DNA sugar (deoxyribose).[39] In order to express information other than the 20 conventional amino acids of proteins, the genetic code can be altered or enlarged. One method involves incorporating a specified unnatural, noncanonical, or xeno amino acid (XAA) into one or more proteins at one or more precise places using orthogonal enzymes and a transfer RNA adaptor from an other organism. By using "directed evolution," which entails repeated cycles of gene mutagenesis (genotypic diversity production), screening or selection (of a specific phenotypic trait), and amplification of a better variant for the following iterative round, orthogonal enzymes are produced Numerous XAAs have been effectively incorporated into proteins in more complex creatures like worms and flies as well as in bacteria, yeast, and human cell lines. As a result of canonical DNA sequence changes, directed evolution also enables the development of orthogonal ribosomes, which make it easier to incorporate XAAs into proteins or create "mirror life," or biological systems that contain biomolecules made up of enantiomers with different chiral orientations.[39]
Enabling technologies
[edit]Several novel enabling technologies were critical to the success of synthetic biology. Concepts include standardization of biological parts and hierarchical abstraction to permit using those parts in synthetic systems.[43] DNA serves as the guide for how biological processes should function, like the score to a complex symphony of life. Our ability to comprehend and design biological systems has undergone significant modifications as a result of developments in the previous few decades in both reading (sequencing) and writing (synthesis) DNA sequences. These developments have produced ground-breaking techniques for designing, assembling, and modifying DNA-encoded genes, materials, circuits, and metabolic pathways, enabling an ever-increasing amount of control over biological systems and even entire organisms.[44]
Basic technologies include reading and writing DNA (sequencing and fabrication). Measurements under multiple conditions are needed for accurate modeling and computer-aided design (CAD).
DNA and gene synthesis
[edit]Driven by dramatic decreases in costs of oligonucleotide ("oligos") synthesis and the advent of PCR, the sizes of DNA constructions from oligos have increased to the genomic level.[45] In 2000, researchers reported synthesis of the 9.6 kbp (kilo bp) Hepatitis C virus genome from chemically synthesized 60 to 80-mers.[46] In 2002, researchers at Stony Brook University succeeded in synthesizing the 7741 bp poliovirus genome from its published sequence, producing the second synthetic genome, spanning two years.[47] In 2003, the 5386 bp genome of the bacteriophage Phi X 174 was assembled in about two weeks.[48] In 2006, the same team, at the J. Craig Venter Institute, constructed and patented a synthetic genome of a novel minimal bacterium, Mycoplasma laboratorium and were working on getting it functioning in a living cell.[49][50][51]
In 2007, it was reported that several companies were offering synthesis of genetic sequences up to 2000 base pairs (bp) long, for a price of about $1 per bp and a turnaround time of less than two weeks.[52] Oligonucleotides harvested from a photolithographic- or inkjet-manufactured DNA chip combined with PCR and DNA mismatch error-correction allows inexpensive large-scale changes of codons in genetic systems to improve gene expression or incorporate novel amino-acids (see George M. Church's and Anthony Forster's synthetic cell projects.[53][54]). This favors a synthesis-from-scratch approach.
Additionally, the CRISPR/Cas system has emerged as a promising technique for gene editing. It was described as "the most important innovation in the synthetic biology space in nearly 30 years".[55] While other methods take months or years to edit gene sequences, CRISPR speeds that time up to weeks.[55] Due to its ease of use and accessibility, however, it has raised ethical concerns, especially surrounding its use in biohacking.[56][57][58]
Sequencing
[edit]DNA sequencing determines the order of nucleotide bases in a DNA molecule. Synthetic biologists use DNA sequencing in their work in several ways. First, large-scale genome sequencing efforts continue to provide information on naturally occurring organisms. This information provides a rich substrate from which synthetic biologists can construct parts and devices. Second, sequencing can verify that the fabricated system is as intended. Third, fast, cheap, and reliable sequencing can facilitate rapid detection and identification of synthetic systems and organisms.[59]
Modularity
[edit]This is the ability of a system or component to operate without reference to its context.[60]
The most used[61]: 22–23 standardized DNA parts are BioBrick plasmids, invented by Tom Knight in 2003.[14] Biobricks are stored at the Registry of Standard Biological Parts in Cambridge, Massachusetts. The BioBrick standard has been used by tens of thousands of students worldwide in the international Genetically Engineered Machine (iGEM) competition. BioBrick Assembly Standard 10 promotes modularity by allowing BioBrick coding sequences to be spliced out and exchanged using restriction enzymes EcoRI or XbaI (BioBrick prefix) and SpeI and PstI (BioBrick suffix).[61]: 22–23
Sequence overlap between two genetic elements (genes or coding sequences), called overlapping genes, can prevent their individual manipulation.[62] To increase genome modularity, the practice of genome refactoring or improving "the internal structure of an existing system for future use, while simultaneously maintaining external system function"[63] has been adopted across synthetic biology disciplines.[62] Some notable examples of refactoring including the nitrogen fixation cluster[64] and type III secretion system[65] along with bacteriophages T7[63] and ΦX174.[66]
While DNA is most important for information storage, a large fraction of the cell's activities are carried out by proteins. Tools can send proteins to specific regions of the cell and to link different proteins together. The interaction strength between protein partners should be tunable between a lifetime of seconds (desirable for dynamic signaling events) up to an irreversible interaction (desirable for device stability or resilient to harsh conditions). Interactions such as coiled coils,[67] SH3 domain-peptide binding[68] or SpyTag/SpyCatcher[69] offer such control. In addition, it is necessary to regulate protein-protein interactions in cells, such as with light (using light-oxygen-voltage-sensing domains) or cell-permeable small molecules by chemically induced dimerization.[70]
In a living cell, molecular motifs are embedded in a bigger network with upstream and downstream components. These components may alter the signaling capability of the modeling module. In the case of ultrasensitive modules, the sensitivity contribution of a module can differ from the sensitivity that the module sustains in isolation.[71][72]
Modeling
[edit]Models inform the design of engineered biological systems by better predicting system behavior prior to fabrication. Synthetic biology benefits from better models of how biological molecules bind substrates and catalyze reactions, how DNA encodes the information needed to specify the cell and how multi-component integrated systems behave. Multiscale models of gene regulatory networks focus on synthetic biology applications. Simulations can model all biomolecular interactions in transcription, translation, regulation and induction of gene regulatory networks.[73] [74] [75][76]
Only extensive modelling can enable the exploration of dynamic gene expression in a form suitable for research and design due to the numerous involved species and the intricacy of their relationships. Dynamic simulations of the entire biomolecular interconnection involved in regulation, transport, transcription, induction, and translation enable the molecular level detailing of designs. As opposed to modelling artificial networks a posteriori, this is contrasted.[77]
Microfluidics
[edit]Microfluidics, in particular droplet microfluidics, is an emerging tool used to construct new components, and to analyze and characterize them.[78][79] It is widely employed in screening assays.[80]
Synthetic transcription factors
[edit]Studies have considered the components of the DNA transcription mechanism. One desire of scientists creating synthetic biological circuits is to be able to control the transcription of synthetic DNA in unicellular organisms (prokaryotes) and in multicellular organisms (eukaryotes). One study tested the adjustability of synthetic transcription factors (sTFs) in areas of transcription output and cooperative ability among multiple transcription factor complexes.[81] Researchers were able to mutate functional regions called zinc fingers, the DNA specific component of sTFs, to decrease their affinity for specific operator DNA sequence sites, and thus decrease the associated site-specific activity of the sTF (usually transcriptional regulation). They further used the zinc fingers as components of complex-forming sTFs, which are the eukaryotic translation mechanisms.[81]
Applications
[edit]Synthetic biology initiatives frequently aim to redesign organisms so that they can create a material, such as a drug or fuel, or acquire a new function, such as the ability to sense something in the environment. Examples of what researchers are creating using synthetic biology include:
- Utilizing microorganisms for bioremediation to remove contaminants from our water, soil, and air.
- Production of complex natural products that are usually extracted from plants but cannot be obtained in sufficient amounts, e.g. drugs of natural origin, such as artemisinin and paclitaxel.
- Beta-carotene, a substance typically associated with carrots that prevents vitamin A deficiency, is produced by rice that has been modified. Every year, between 250,000 and 500,000 children lose their vision due to vitamin A deficiency, which also significantly raises their chance of dying from infectious infections.
- As a sustainable and environmentally benign alternative to the fresh roses that perfumers use to create expensive smells, yeast has been created to produce rose oil.[82]
Biosensors
[edit]A biosensor refers to an engineered organism, usually a bacterium, that is capable of reporting some ambient phenomenon such as the presence of heavy metals or toxins. One such system is the Lux operon of Aliivibrio fischeri,[83] which codes for the enzyme that is the source of bacterial bioluminescence, and can be placed after a respondent promoter to express the luminescence genes in response to a specific environmental stimulus.[84] One such sensor created, consisted of a bioluminescent bacterial coating on a photosensitive computer chip to detect certain petroleum pollutants. When the bacteria sense the pollutant, they luminesce.[85] Another example of a similar mechanism is the detection of landmines by an engineered E.coli reporter strain capable of detecting TNT and its main degradation product DNT, and consequently producing a green fluorescent protein (GFP).[86]
Modified organisms can sense environmental signals and send output signals that can be detected and serve diagnostic purposes. Microbe cohorts have been used.[87]
Biosensors could also be used to detect pathogenic signatures—such as of SARS-CoV-2—and can be wearable.[88][89]
For the purpose of detecting and reacting to various and temporary environmental factors, cells have developed a wide range of regulatory circuits, ranging from transcriptional to post-translational. These circuits are made up of transducer modules that filter the signals and activate a biological response, as well as carefully designed sensitive sections that attach analytes and regulate signal-detection thresholds. Modularity and selectivity are programmed to biosensor circuits at the transcriptional, translational, and post-translational levels, to achieve the delicate balancing of the two basic sensing modules.[90]
Food and drink
[edit]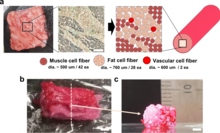
However, not all synthetic nutrition products are animal food products – for instance, as of 2021, there are also products of synthetic coffee that are reported to be close to commercialization.[98][99][100] Similar fields of research and production based on synthetic biology that can be used for the production of food and drink are:
- Genetically engineered microbial food cultures (e.g. for solar-energy-based protein powder)[101][102]
- Cell-free artificial synthesis (e.g. synthetic starch;[103][104] )
Materials
[edit]Photosynthetic microbial cells have been used as a step to synthetic production of spider silk.[105][106]
Biological computers
[edit]A biological computer refers to an engineered biological system that can perform computer-like operations, which is a dominant paradigm in synthetic biology. Researchers built and characterized a variety of logic gates in a number of organisms,[107] and demonstrated both analog and digital computation in living cells. They demonstrated that bacteria can be engineered to perform both analog and/or digital computation.[108][109] In 2007, in human cells, research demonstrated a universal logic evaluator that operates in mammalian cells.[110] Subsequently, researchers utilized this paradigm to demonstrate a proof-of-concept therapy that uses biological digital computation to detect and kill human cancer cells in 2011.[111] In 2016, another group of researchers demonstrated that principles of computer engineering can be used to automate digital circuit design in bacterial cells.[112] In 2017, researchers demonstrated the 'Boolean logic and arithmetic through DNA excision' (BLADE) system to engineer digital computation in human cells.[113] In 2019, researchers implemented a perceptron in biological systems opening the way for machine learning in these systems.[114]
Cell transformation
[edit]Cells use interacting genes and proteins, which are called gene circuits, to implement diverse function, such as responding to environmental signals, decision making and communication. Three key components are involved: DNA, RNA and Synthetic biologist designed gene circuits that can control gene expression from several levels including transcriptional, post-transcriptional and translational levels.
Traditional metabolic engineering has been bolstered by the introduction of combinations of foreign genes and optimization by directed evolution. This includes engineering E. coli and yeast for commercial production of a precursor of the antimalarial drug, Artemisinin.[115]
Entire organisms have yet to be created from scratch, although living cells can be transformed with new DNA. Several ways allow constructing synthetic DNA components and even entire synthetic genomes, but once the desired genetic code is obtained, it is integrated into a living cell that is expected to manifest the desired new capabilities or phenotypes while growing and thriving.[116] Cell transformation is used to create biological circuits, which can be manipulated to yield desired outputs.[12][13]
By integrating synthetic biology with materials science, it would be possible to use cells as microscopic molecular foundries to produce materials whose properties were genetically encoded. Re-engineering has produced Curli fibers, the amyloid component of extracellular material of biofilms, as a platform for programmable nanomaterial. These nanofibers were genetically constructed for specific functions, including adhesion to substrates, nanoparticle templating and protein immobilization.[117]
Designed proteins
[edit]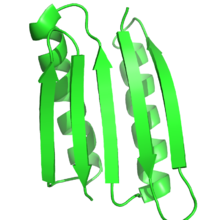
Natural proteins can be engineered, for example, by directed evolution, novel protein structures that match or improve on the functionality of existing proteins can be produced. One group generated a helix bundle that was capable of binding oxygen with similar properties as hemoglobin, yet did not bind carbon monoxide.[119] A similar protein structure was generated to support a variety of oxidoreductase activities[120] while another formed a structurally and sequentially novel ATPase.[121] Another group generated a family of G-protein coupled receptors that could be activated by the inert small molecule clozapine N-oxide but insensitive to the native ligand, acetylcholine; these receptors are known as DREADDs.[122] Novel functionalities or protein specificity can also be engineered using computational approaches. One study was able to use two different computational methods: a bioinformatics and molecular modeling method to mine sequence databases, and a computational enzyme design method to reprogram enzyme specificity. Both methods resulted in designed enzymes with greater than 100 fold specificity for production of longer chain alcohols from sugar.[123]
Another common investigation is expansion of the natural set of 20 amino acids. Excluding stop codons, 61 codons have been identified, but only 20 amino acids are coded generally in all organisms. Certain codons are engineered to code for alternative amino acids including: nonstandard amino acids such as O-methyl tyrosine; or exogenous amino acids such as 4-fluorophenylalanine. Typically, these projects make use of re-coded nonsense suppressor tRNA-Aminoacyl tRNA synthetase pairs from other organisms, though in most cases substantial engineering is required.[124]
Other researchers investigated protein structure and function by reducing the normal set of 20 amino acids. Limited protein sequence libraries are made by generating proteins where groups of amino acids may be replaced by a single amino acid.[125] For instance, several non-polar amino acids within a protein can all be replaced with a single non-polar amino acid.[126] One project demonstrated that an engineered version of Chorismate mutase still had catalytic activity when only nine amino acids were used.[127]
Researchers and companies practice synthetic biology to synthesize industrial enzymes with high activity, optimal yields and effectiveness. These synthesized enzymes aim to improve products such as detergents and lactose-free dairy products, as well as make them more cost effective.[128] The improvements of metabolic engineering by synthetic biology is an example of a biotechnological technique utilized in industry to discover pharmaceuticals and fermentive chemicals. Synthetic biology may investigate modular pathway systems in biochemical production and increase yields of metabolic production. Artificial enzymatic activity and subsequent effects on metabolic reaction rates and yields may develop "efficient new strategies for improving cellular properties ... for industrially important biochemical production".[129]
Designed nucleic acid systems
[edit]Scientists can encode digital information onto a single strand of synthetic DNA. In 2012, George M. Church encoded one of his books about synthetic biology in DNA. The 5.3 Mb of data was more than 1000 times greater than the previous largest amount of information to be stored in synthesized DNA.[130] A similar project encoded the complete sonnets of William Shakespeare in DNA.[131] More generally, algorithms such as NUPACK,[132] ViennaRNA,[133] Ribosome Binding Site Calculator,[134] Cello,[135] and Non-Repetitive Parts Calculator[136] enables the design of new genetic systems.
Many technologies have been developed for incorporating unnatural nucleotides and amino acids into nucleic acids and proteins, both in vitro and in vivo. For example, in May 2014, researchers announced that they had successfully introduced two new artificial nucleotides into bacterial DNA. By including individual artificial nucleotides in the culture media, they were able to exchange the bacteria 24 times; they did not generate mRNA or proteins able to use the artificial nucleotides.[137][138][139]
Space exploration
[edit]Synthetic biology raised NASA's interest as it could help to produce resources for astronauts from a restricted portfolio of compounds sent from Earth.[140][141][142] On Mars, in particular, synthetic biology could lead to production processes based on local resources, making it a powerful tool in the development of occupied outposts with less dependence on Earth.[140] Work has gone into developing plant strains that are able to cope with the harsh Martian environment, using similar techniques to those employed to increase resilience to certain environmental factors in agricultural crops.[143]
Synthetic life
[edit]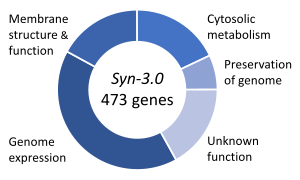
One important topic in synthetic biology is synthetic life, that is concerned with hypothetical organisms created in vitro from biomolecules and/or chemical analogues thereof. Synthetic life experiments attempt to either probe the origins of life, study some of the properties of life, or more ambitiously to recreate life from non-living (abiotic) components. Synthetic life biology attempts to create living organisms capable of carrying out important functions, from manufacturing pharmaceuticals to detoxifying polluted land and water.[145] In medicine, it offers prospects of using designer biological parts as a starting point for new classes of therapies and diagnostic tools.[145]
A living "artificial cell" has been defined as a completely synthetic cell that can capture energy, maintain ion gradients, contain macromolecules as well as store information and have the ability to mutate.[146] Nobody has been able to create such a cell.[146]
A completely synthetic bacterial chromosome was produced in 2010 by Craig Venter, and his team introduced it to genomically emptied bacterial host cells.[19] The host cells were able to grow and replicate.[147][148] The Mycoplasma laboratorium is the only living organism with completely engineered genome.
The first living organism with 'artificial' expanded DNA code was presented in 2014; the team used E. coli that had its genome extracted and replaced with a chromosome with an expanded genetic code. The nucleosides added are d5SICS and dNaM.[139]
In May 2019, in a milestone effort, researchers reported the creation of a new synthetic (possibly artificial) form of viable life, a variant of the bacteria Escherichia coli, by reducing the natural number of 64 codons in the bacterial genome to 59 codons instead, in order to encode 20 amino acids.[25][26]
In 2017, the international Build-a-Cell large-scale open-source research collaboration for the construction of synthetic living cells was started,[149] followed by national synthetic cell organizations in several countries, including FabriCell,[150] MaxSynBio[151] and BaSyC.[152] The European synthetic cell efforts were unified in 2019 as SynCellEU initiative.[153]
In 2023, researchers were able to create the first synthetically made human embryos derived from stem cells.[154]
Drug delivery platforms
[edit]In therapeutics, synthetic biology has achieved significant advancements in altering and simplifying the therapeutics scope in a relatively short period of time. In fact, new therapeutic platforms, from the discovery of disease mechanisms and drug targets to the manufacture and transport of small molecules, are made possible by the logical and model-guided design construction of biological components.[60]
Synthetic biology devices have been designed to act as therapies in therapeutic treatment. It is possible to control complete created viruses and organisms to target particular pathogens and diseased pathways. Thus, in two independent studies 91,92, researchers utilised genetically modified bacteriophages to fight antibiotic-resistant bacteria by giving them genetic features that specifically target and hinder bacterial defences against antibiotic activity.[155]
In the therapy of cancer, since conventional medicines frequently indiscriminately target tumours and normal tissues, artificially created viruses and organisms that can identify and connect their therapeutic action to pathological signals may be helpful. For example, p53 pathway activity in human cells was put into adenoviruses to control how they replicated.[155]
Engineered bacteria-based platform
[edit]Bacteria have long been used in cancer treatment. Bifidobacterium and Clostridium selectively colonize tumors and reduce their size.[156] Recently synthetic biologists reprogrammed bacteria to sense and respond to a particular cancer state. Most often bacteria are used to deliver a therapeutic molecule directly to the tumor to minimize off-target effects. To target the tumor cells, peptides that can specifically recognize a tumor were expressed on the surfaces of bacteria. Peptides used include an affibody molecule that specifically targets human epidermal growth factor receptor 2[157] and a synthetic adhesin.[158] The other way is to allow bacteria to sense the tumor microenvironment, for example hypoxia, by building an AND logic gate into bacteria.[159] Then the bacteria only release target therapeutic molecules to the tumor through either lysis[160] or the bacterial secretion system.[161] Lysis has the advantage that it can stimulate the immune system and control growth. Multiple types of secretion systems can be used and other strategies as well. The system is inducible by external signals. Inducers include chemicals, electromagnetic or light waves.
Multiple species and strains are applied in these therapeutics. Most commonly used bacteria are Salmonella typhimurium, Escherichia coli, Bifidobacteria, Streptococcus, Lactobacillus, Listeria and Bacillus subtilis. Each of these species have their own property and are unique to cancer therapy in terms of tissue colonization, interaction with immune system and ease of application.
Engineered yeast-based platform
Synthetic biologists are developing genetically modified live yeast that can deliver therapeutic biologic medicines. When orally delivered, these live yeast act like micro-factories and will make therapeutic molecules directly in the gastrointestinal tract. Because yeast are eukaryotic, a key benefit is that they can be administered together with antibiotics. Probiotic yeast expressing human P2Y2 purinergic receptor suppressed intestinal inflammation in mouse models of inflammatory bowel disease.[162] A live S. boulardii yeast delivering a tetra-specific anti-toxin that potently neutralizes Toxin A and Toxin B of Clostridioides difficile has been developed. This therapeutic anti-toxin is a fusion of four single-domain antibodies (nanobodies) that potently and broadly neutralize the two major virulence factors of C. difficile at the site of infection in preclinical models.[163] The first in human clinical trial of engineered live yeast for the treatment of Clostridium difficile infection is anticipated in 2024 and will be sponsored by the developer Fzata, Inc.
Cell-based platform
[edit]The immune system plays an important role in cancer and can be harnessed to attack cancer cells. Cell-based therapies focus on immunotherapies, mostly by engineering T cells.
T cell receptors were engineered and 'trained' to detect cancer epitopes. Chimeric antigen receptors (CARs) are composed of a fragment of an antibody fused to intracellular T cell signaling domains that can activate and trigger proliferation of the cell. Multiple second generation CAR-based therapies have been approved by FDA.[164]
Gene switches were designed to enhance safety of the treatment. Kill switches were developed to terminate the therapy should the patient show severe side effects.[165] Mechanisms can more finely control the system and stop and reactivate it.[166][167] Since the number of T-cells are important for therapy persistence and severity, growth of T-cells is also controlled to dial the effectiveness and safety of therapeutics.[168]
Although several mechanisms can improve safety and control, limitations include the difficulty of inducing large DNA circuits into the cells and risks associated with introducing foreign components, especially proteins, into cells.
Biofuels, pharmaceuticals and biomaterials
[edit]The most popular biofuel is ethanol produced from corn or sugar cane, but this method of producing biofuels is troublesome and constrained due to the high agricultural cost and inadequate fuel characteristics of ethanol. An substitute and potential source of renewable energy is microbes that have had their metabolic pathways altered to be more efficient at converting biomass into biofuels. Only if their production costs could be made to match or even exceed those of present fuel production can these techniques be expected to be successful. Related to this, there are several medicines whose pricey manufacturing procedures prevent them from having a larger therapeutic range. The creation of new materials and the microbiological manufacturing of biomaterials would both benefit substantially from novel artificial biology tools.[155]
CRISPR/Cas9
[edit]The clustered frequently interspaced short palindromic repetitions (CRISPR)/CRISPR associated (Cas) system is a powerful method of genome engineering in a range of organisms because of its simplicity, modularity, and scalability. In this technique, a guide RNA (gRNA) attracts the CRISPR nuclease Cas9 to a particular spot in the genome, causing a double strand break. Several DNA repair processes, including homology-directed recombination and non-homology end joining, can be used to accomplish the desired genome change (i.e., gene deletion or insertion). Additionally, dCas9 (dead Cas9 or nuclease-deficient Cas9), a Cas9 double mutant (H840A, D10A), has been utilised to control gene expression in bacteria or when linked to a stimulation of suppression site in yeast.[169]
Regulatory elements
[edit]To build and develop biological systems, regulating components including regulators, ribosome-binding sites (RBSs), and terminators are crucial. Despite years of study, there are many various varieties and numbers of promoters and terminators for Escherichia coli, but also for the well-researched model organism Saccharomyces cerevisiae, as well as for other organisms of interest, these tools are quite scarce. Numerous techniques have been invented for the finding and identification of promoters and terminators in order to overcome this constraint, including genome mining, random mutagenesis, hybrid engineering, biophysical modelling, combinatorial design, and rational design.[169]
Organoids
[edit]Synthetic biology has been used for organoids, which are lab-grown organs with application to medical research and transplantation.[170]
Bioprinted organs
[edit]3D bioprinting can be used to reconstruct tissue from various regions of the body. The precursor to the adoption of 3D printing in healthcare was a series of trials conducted by researchers at Boston Children's Hospital. The team built replacement urinary bladders by hand for seven patients by constructing scaffolds, then layering the scaffolds with cells from the patients and allowing them to grow. The trials were a success as the patients remained in good health 7 years after implantation, which led a research fellow named Anthony Atala, MD, to search or ways to automate the process.[171] Patients with end-stage bladder disease can now be treated by using bio-engineered bladder tissues to rebuild the damaged organ.[172] This technology can also potentially be applied to bone, skin, cartilage and muscle tissue.[173] Though one long-term goal of 3D bioprinting technology is to reconstruct an entire organ as well as minimize the problem of the lack of organs for transplantation.[174] There has been little success in bioprinting of fully functional organs e.g. liver, skin, meniscus or pancreas.[175][176][177] Unlike implantable stents, organs have complex shapes and are significantly harder to bioprint. A bioprinted heart, for example, must not only meet structural requirements, but also vascularization, mechanical load, and electrical signal propagation requirements.[178] In 2022, the first success of a clinical trial for a 3D bioprinted transplant that is made from the patient's own cells, an external ear to treat microtia,[179] was reported.[180]
3D bioprinting contributes to significant advances in the medical field of tissue engineering by allowing for research to be done on innovative materials called biomaterials. Some of the most notable bioengineered substances are usually stronger than the average bodily materials, including soft tissue and bone. These constituents can act as future substitutes, even improvements, for the original body materials. In addition, the Defense Threat Reduction Agency aims to print mini organs such as hearts, livers, and lungs as the potential to test new drugs more accurately and perhaps eliminate the need for testing in animals.[181]
Other transplants and induced regeneration
[edit]There is ongoing research and development into synthetic biology based methods for inducing regeneration in humans[relevant?] as well the creation of transplantable artificial organs.
Nanoparticles, artificial cells and micro-droplets
[edit]Synthetic biology can be used for creating nanoparticles which can be used for drug-delivery as well as for other purposes.[182] Complementing research and development seeks to and has created synthetic cells that mimics functions of biological cells. Applications include medicine such as designer-nanoparticles that make blood cells eat away—from the inside out—portions of atherosclerotic plaque that cause heart attacks.[183][184][185] Synthetic micro-droplets for algal cells or synergistic algal-bacterial multicellular spheroid microbial reactors, for example, could be used to produce hydrogen as hydrogen economy biotechnology.[186][187]
Electrogenetics
[edit]Mammalian designer cells are engineered by humans to behave a specific way, such as an immune cell that expresses a synthetic receptor designed to combat a specific disease.[188][189] Electrogenetics is an application of synthetic biology that involves utilizing electrical fields to stimulate a response in engineered cells.[190] Controlling the designer cells can be done with relative ease through the use of common electronic devices, such as smartphones. Additionally, electrogenetics allows for the possibility of creating devices that are much smaller and compact than devices that use other stimulus through the use of microscopic electrodes.[190] One example of how electrogenetics is used to benefit public health is through stimulating designer cells that are able to produce/deliver therapeutics.[191] This was implemented in ElectroHEK cells, cells that contain voltage-gated calcium channels that are electrosensitive, meaning that the ion channel can be controlled by electrical conduction between electrodes and the ElectroHEK cells.[190] The expression levels of the artificial gene that these ElectroHEK cells contained was shown to be able to be controlled by changing the voltage or electrical pulse length. Further studies have expanded on this robust system, one of which is a beta cell line system designed to control the release of insulin based on electric signals.[192]
Ethics
[edit]This section needs to be updated. (January 2019) |
The creation of new life and the tampering of existing life has raised ethical concerns in the field of synthetic biology and are actively being discussed.[193][194]
Common ethical questions include:
- Is it morally right to tamper with nature?
- Is one playing God when creating new life?
- What happens if a synthetic organism accidentally escapes?
- What if an individual misuses synthetic biology and creates a harmful entity (e.g., a biological weapon)?
- Who will have control of and access to the products of synthetic biology?
- Who will gain from these innovations? Investors? Medical patients? Industrial farmers?
- Does the patent system allow patents on living organisms? What about parts of organisms, like HIV resistance genes in humans?[195]
- What if a new creation is deserving of moral or legal status?
The ethical aspects of synthetic biology has three main features: biosafety, biosecurity, and the creation of new life forms.[196] Other ethical issues mentioned include the regulation of new creations, patent management of new creations, benefit distribution, and research integrity.[197][193]
Ethical issues have surfaced for recombinant DNA and genetically modified organism (GMO) technologies and extensive regulations of genetic engineering and pathogen research were in place in many jurisdictions. Amy Gutmann, former head of the Presidential Bioethics Commission, argued that we should avoid the temptation to over-regulate synthetic biology in general, and genetic engineering in particular. According to Gutmann, "Regulatory parsimony is especially important in emerging technologies...where the temptation to stifle innovation on the basis of uncertainty and fear of the unknown is particularly great. The blunt instruments of statutory and regulatory restraint may not only inhibit the distribution of new benefits, but can be counterproductive to security and safety by preventing researchers from developing effective safeguards.".[198]
The "creation" of life
[edit]One ethical question is whether or not it is acceptable to create new life forms, sometimes known as "playing God". Currently, the creation of new life forms not present in nature is at small-scale, the potential benefits and dangers remain unknown, and careful consideration and oversight are ensured for most studies.[193] Many advocates express the great potential value—to agriculture, medicine, and academic knowledge, among other fields—of creating artificial life forms. Creation of new entities could expand scientific knowledge well beyond what is currently known from studying natural phenomena. Yet there is concern that artificial life forms may reduce nature's "purity" (i.e., nature could be somehow corrupted by human intervention and manipulation) and potentially influence the adoption of more engineering-like principles instead of biodiversity- and nature-focused ideals. Some are also concerned that if an artificial life form were to be released into nature, it could hamper biodiversity by beating out natural species for resources (similar to how algal blooms kill marine species). Another concern involves the ethical treatment of newly created entities if they happen to sense pain, sentience, and self-perception. There is an ongoing debate as to whether such life forms should be granted moral or legal rights, though no consensus exists as to how these rights would be administered or enforced.
Ethical support for synthetic biology
[edit]Ethics and moral rationales that support certain applications of synthetic biology include their potential mitigation of substantial global problems of detrimental environmental impacts of conventional agriculture (including meat production), animal welfare, food security, and human health,[199][200][201][202] as well as potential reduction of human labor needs and, via therapies of diseases, reduction of human suffering and prolonged life.
Biosafety and biocontainment
[edit]What is most ethically appropriate when considering biosafety measures? How can accidental introduction of synthetic life in the natural environment be avoided? Much ethical consideration and critical thought has been given to these questions. Biosafety not only refers to biological containment; it also refers to strides taken to protect the public from potentially hazardous biological agents. Even though such concerns are important and remain unanswered, not all products of synthetic biology present concern for biological safety or negative consequences for the environment. It is argued that most synthetic technologies are benign and are incapable of flourishing in the outside world due to their "unnatural" characteristics as there is yet to be an example of a transgenic microbe conferred with a fitness advantage in the wild.
In general, existing hazard controls, risk assessment methodologies, and regulations developed for traditional genetically modified organisms (GMOs) are considered to be sufficient for synthetic organisms. "Extrinsic" biocontainment methods in a laboratory context include physical containment through biosafety cabinets and gloveboxes, as well as personal protective equipment. In an agricultural context, they include isolation distances and pollen barriers, similar to methods for biocontainment of GMOs. Synthetic organisms may offer increased hazard control because they can be engineered with "intrinsic" biocontainment methods that limit their growth in an uncontained environment, or prevent horizontal gene transfer to natural organisms. Examples of intrinsic biocontainment include auxotrophy, biological kill switches, inability of the organism to replicate or to pass modified or synthetic genes to offspring, and the use of xenobiological organisms using alternative biochemistry, for example using artificial xeno nucleic acids (XNA) instead of DNA.[203][204] Regarding auxotrophy, bacteria and yeast can be engineered to be unable to produce histidine, an important amino acid for all life. Such organisms can thus only be grown on histidine-rich media in laboratory conditions, nullifying fears that they could spread into undesirable areas.
Biosecurity and bioterrorism
[edit]Some ethical issues relate to biosecurity, where biosynthetic technologies could be deliberately used to cause harm to society and/or the environment. Since synthetic biology raises ethical issues and biosecurity issues, humanity must consider and plan on how to deal with potentially harmful creations, and what kinds of ethical measures could possibly be employed to deter nefarious biosynthetic technologies. With the exception of regulating synthetic biology and biotechnology companies,[205][206] however, the issues are not seen as new because they were raised during the earlier recombinant DNA and genetically modified organism (GMO) debates, and extensive regulations of genetic engineering and pathogen research are already in place in many jurisdictions.[207]
Additionally, the development of synthetic biology tools has made it easier for individuals with less education, training, and access to equipment to modify and use pathogenic organisms as bioweapons. This increases the threat of bioterrorism, especially as terrorist groups become aware of the significant social, economic, and political disruption caused by pandemics like COVID-19. As new techniques are developed in the field of synthetic biology, the risk of bioterrorism is likely to continue to grow.[208] Juan Zarate, who served as Deputy National Security Advisor for Combating Terrorism from 2005 to 2009, noted that "the severity and extreme disruption of a novel coronavirus will likely spur the imagination of the most creative and dangerous groups and individuals to reconsider bioterrorist attacks."[209]
European Union
[edit]The European Union-funded project SYNBIOSAFE[210] has issued reports on how to manage synthetic biology. A 2007 paper identified key issues in safety, security, ethics, and the science-society interface, which the project defined as public education and ongoing dialogue among scientists, businesses, government and ethicists.[211][212] The key security issues that SYNBIOSAFE identified involved engaging companies that sell synthetic DNA and the biohacking community of amateur biologists. Key ethical issues concerned the creation of new life forms.
A subsequent report focused on biosecurity, especially the so-called dual-use challenge. For example, while synthetic biology may lead to more efficient production of medical treatments, it may also lead to synthesis or modification of harmful pathogens (e.g., smallpox).[213] The biohacking community remains a source of special concern, as the distributed and diffuse nature of open-source biotechnology makes it difficult to track, regulate or mitigate potential concerns over biosafety and biosecurity.[214]
COSY, another European initiative, focuses on public perception and communication.[215][216][217] To better communicate synthetic biology and its societal ramifications to a broader public, COSY and SYNBIOSAFE published SYNBIOSAFE, a 38-minute documentary film, in October 2009.[210]
The International Association Synthetic Biology has proposed self-regulation.[218] This proposes specific measures that the synthetic biology industry, especially DNA synthesis companies, should implement. In 2007, a group led by scientists from leading DNA-synthesis companies published a "practical plan for developing an effective oversight framework for the DNA-synthesis industry".[205]
United States
[edit]In January 2009, the Alfred P. Sloan Foundation funded the Woodrow Wilson Center, the Hastings Center, and the J. Craig Venter Institute to examine the public perception, ethics and policy implications of synthetic biology.[219]
On July 9–10, 2009, the National Academies' Committee of Science, Technology & Law convened a symposium on "Opportunities and Challenges in the Emerging Field of Synthetic Biology".[220]
After the publication of the first synthetic genome and the accompanying media coverage about "life" being created, President Barack Obama established the Presidential Commission for the Study of Bioethical Issues to study synthetic biology.[221] The commission convened a series of meetings, and issued a report in December 2010 titled "New Directions: The Ethics of Synthetic Biology and Emerging Technologies." The commission stated that "while Venter's achievement marked a significant technical advance in demonstrating that a relatively large genome could be accurately synthesized and substituted for another, it did not amount to the "creation of life".[222] It noted that synthetic biology is an emerging field, which creates potential risks and rewards. The commission did not recommend policy or oversight changes and called for continued funding of the research and new funding for monitoring, study of emerging ethical issues and public education.[207]
Synthetic biology, as a major tool for biological advances, results in the "potential for developing biological weapons, possible unforeseen negative impacts on human health ... and any potential environmental impact".[223] The proliferation of such technology could also make the production of biological and chemical weapons available to a wider array of state and non-state actors.[224] These security issues may be avoided by regulating industry uses of biotechnology through policy legislation. Federal guidelines on genetic manipulation are being proposed by "the President's Bioethics Commission ... in response to the announced creation of a self-replicating cell from a chemically synthesized genome, put forward 18 recommendations not only for regulating the science ... for educating the public".[223]
Opposition
[edit]On March 13, 2012, over 100 environmental and civil society groups, including Friends of the Earth, the International Center for Technology Assessment, and the ETC Group, issued the manifesto The Principles for the Oversight of Synthetic Biology. This manifesto calls for a worldwide moratorium on the release and commercial use of synthetic organisms until more robust regulations and rigorous biosafety measures are established. The groups specifically call for an outright ban on the use of synthetic biology on the human genome or human microbiome.[225][226] Richard Lewontin wrote that some of the safety tenets for oversight discussed in The Principles for the Oversight of Synthetic Biology are reasonable, but that the main problem with the recommendations in the manifesto is that "the public at large lacks the ability to enforce any meaningful realization of those recommendations".[227]
Health and safety
[edit]The hazards of synthetic biology include biosafety hazards to workers and the public, biosecurity hazards stemming from deliberate engineering of organisms to cause harm, and environmental hazards. The biosafety hazards are similar to those for existing fields of biotechnology, mainly exposure to pathogens and toxic chemicals, although novel synthetic organisms may have novel risks.[203] For biosecurity, there is concern that synthetic or redesigned organisms could theoretically be used for bioterrorism. Potential risks include recreating known pathogens from scratch, engineering existing pathogens to be more dangerous, and engineering microbes to produce harmful biochemicals.[228] Lastly, environmental hazards include adverse effects on biodiversity and ecosystem services, including potential changes to land use resulting from agricultural use of synthetic organisms.[229][230] Synthetic biology is an example of a dual-use technology with the potential to be used in ways that could intentionally or unintentionally harm humans and/or damage the environment. Often "scientists, their host institutions and funding bodies" consider whether the planned research could be misused and sometimes implement measures to reduce the likelihood of misuse.[231]
Existing risk analysis systems for GMOs are generally considered sufficient for synthetic organisms, although there may be difficulties for an organism built "bottom-up" from individual genetic sequences.[204][232] Synthetic biology generally falls under existing regulations for GMOs and biotechnology in general, and any regulations that exist for downstream commercial products, although there are generally no regulations in any jurisdiction that are specific to synthetic biology.[233][234]
See also
[edit]- ACS Synthetic Biology (journal)
- Bioengineering
- Biomimicry
- Carlson Curve
- Chiral life concept
- Computational biology
- Computational biomodeling
- DNA digital data storage
- Engineering biology
- International Genetically Engineered Machine
- Non-cellular life
- Open synthetic biology
- Regenerative medicine
- Synthetic intelligence
- Synthetic morphology
- Synthetic virology
- Systems and Synthetic Biology (journal)
- Tissue engineering
- Xenobiology
- Protocell#Artificial models
- Jeewanu
- Hypothetical types of biochemistry
- Playing God (ethics)
References
[edit]- ^ a b Hanczyc MM (May 2020). "Engineering Life: A Review of Synthetic Biology". Artificial Life. 26 (2): 260–273. doi:10.1162/artl_a_00318. hdl:11572/302757. ISSN 1064-5462. PMID 32271630. S2CID 215550945.
- ^ Nakano T, Eckford AW, Haraguchi T (12 September 2013). Molecular Communication. Cambridge University Press. ISBN 978-1-107-02308-6.
- ^ a b "Synthetic Biology". Genome.gov. Retrieved 2023-01-03.
- ^ "Productive Nanosystems: A Technology Roadmap" (PDF). Foresight Institute. Archived from the original (PDF) on 2016-10-25. Retrieved 2020-05-02.
- ^ Le Duc S (October 26, 1910). Théorie physico-chimique de la vie et générations spontanées. A. Poinat. OL 23348076M – via The Open Library.
- ^ Leduc S (1912). Poinat A (ed.). La biologie synthétique, étude de biophysique. Archived from the original on 2016-09-27. Retrieved 2011-01-23.
- ^ Carey, Nessa (2019). Hacking the Code of Life. London: Icon Books. pp. Page=12. ISBN 978-178578-497-2.
- ^ Jacob, F.ß. & Monod, J. On the regulation of gene activity. Cold Spring Harb. Symp. Quant. Biol. 26, 193–211 (1961).
- ^ Cohen SN, Chang AC, Boyer HW, Helling RB (1973). "Construction of biologically functional bacterial plasmids in vitro". Proc. Natl. Acad. Sci. USA. 70 (11): 3240–3244. Bibcode:1973PNAS...70.3240C. doi:10.1073/pnas.70.11.3240. PMC 427208. PMID 4594039.
- ^ Szybalski W, Skalka A (November 1978). "Nobel prizes and restriction enzymes". Gene. 4 (3): 181–2. doi:10.1016/0378-1119(78)90016-1. PMID 744485.
- ^ Saiki RK, Gelfand DH, Stoffel S, Scharf SJ, Higuchi R, Horn GT, Mullis KB, Erlich HA (1988). "Primer-directed enzymatic amplification of DNA with a thermostable DNA polymerase". Science. 239 (4839): 487–491. doi:10.1126/science.239.4839.487. PMID 2448875.
- ^ a b Elowitz MB, Leibler S (January 2000). "A synthetic oscillatory network of transcriptional regulators". Nature. 403 (6767): 335–8. Bibcode:2000Natur.403..335E. doi:10.1038/35002125. PMID 10659856. S2CID 41632754.
- ^ a b Gardner TS, Cantor CR, Collins JJ (January 2000). "Construction of a genetic toggle switch in Escherichia coli". Nature. 403 (6767): 339–42. Bibcode:2000Natur.403..339G. doi:10.1038/35002131. PMID 10659857. S2CID 345059.
- ^ a b Knight T (2003). Idempotent Vector Design for Standard Assembly of Biobricks (Report). MIT Artificial Intelligence Laboratory. hdl:1721.1/21168.
- ^ Martin VJ, Pitera DJ, Withers ST, Newman JD, Keasling JD (July 2003). "Engineering a mevalonate pathway in Escherichia coli for production of terpenoids". Nature Biotechnology. 21 (7): 796–802. doi:10.1038/nbt833. PMID 12778056. S2CID 17214504.
- ^ Levskaya A, Chevalier AA, Tabor JJ, Simpson ZB, Lavery LA, Levy M, et al. (November 2005). "Synthetic biology: engineering Escherichia coli to see light". Nature. 438 (7067): 441–442. Bibcode:2005Natur.438..441L. doi:10.1038/nature04405. PMID 16306980. S2CID 4428475.
- ^ Basu S, Gerchman Y, Collins CH, Arnold FH, Weiss R (April 2005). "A synthetic multicellular system for programmed pattern formation". Nature. 434 (7037): 1130–4. Bibcode:2005Natur.434.1130B. doi:10.1038/nature03461. PMID 15858574. S2CID 4370309.
- ^ Anderson JC, Clarke EJ, Arkin AP, Voigt CA (January 2006). "Environmentally controlled invasion of cancer cells by engineered bacteria". Journal of Molecular Biology. 355 (4): 619–627. doi:10.1016/j.jmb.2005.10.076. PMID 16330045.
- ^ a b Gibson DG, Glass JI, Lartigue C, Noskov VN, Chuang RY, Algire MA, Benders GA, Montague MG, Ma L, Moodie MM, Merryman C, Vashee S, Krishnakumar R, Assad-Garcia N, Andrews-Pfannkoch C, Denisova EA, Young L, Qi ZQ, Segall-Shapiro TH, Calvey CH, Parmar PP, Hutchison CA, Smith HO, Venter JC (July 2010). "Creation of a bacterial cell controlled by a chemically synthesized genome". Science. 329 (5987): 52–6. Bibcode:2010Sci...329...52G. doi:10.1126/science.1190719. PMID 20488990.
- ^ "American scientist who created artificial life denies 'playing God'". The Telegraph. May 2010. Archived from the original on 2022-01-12.
- ^ Dymond JS, Richardson SM, Coombes CE, Babatz T, Muller H, Annaluru N, et al. (September 2011). "Synthetic chromosome arms function in yeast and generate phenotypic diversity by design". Nature. 477 (7365): 471–476. Bibcode:2011Natur.477..471D. doi:10.1038/nature10403. PMC 3774833. PMID 21918511.
- ^ Jinek M, Chylinski K, Fonfara I, Hauer M, Doudna JA, Charpentier E (2012). "A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity". Science. 337 (6096): 816–821. Bibcode:2012Sci...337..816J. doi:10.1126/science.1225829. PMC 6286148. PMID 22745249.
- ^ ETH Zurich (1 April 2019). "First bacterial genome created entirely with a computer". EurekAlert!. Retrieved 2 April 2019.
- ^ Venetz JE, Del Medico L, Wölfle A, Schächle P, Bucher Y, Appert D, et al. (April 2019). "Chemical synthesis rewriting of a bacterial genome to achieve design flexibility and biological functionality". Proceedings of the National Academy of Sciences of the United States of America. 116 (16): 8070–8079. Bibcode:2019PNAS..116.8070V. doi:10.1073/pnas.1818259116. PMC 6475421. PMID 30936302.
- ^ a b Zimmer C (15 May 2019). "Scientists Created Bacteria With a Synthetic Genome. Is This Artificial Life? – In a milestone for synthetic biology, colonies of E. coli thrive with DNA constructed from scratch by humans, not nature". The New York Times. Retrieved 16 May 2019.
- ^ a b Fredens J, Wang K, de la Torre D, Funke LF, Robertson WE, Christova Y, et al. (May 2019). "Total synthesis of Escherichia coli with a recoded genome". Nature. 569 (7757): 514–518. Bibcode:2019Natur.569..514F. doi:10.1038/s41586-019-1192-5. PMC 7039709. PMID 31092918.
- ^ Sokol J (2020-04-03). "Meet the Xenobots: Virtual Creatures Brought to Life". The New York Times.
- ^ Kriegman S, Blackiston D, Levin M, Bongard J (December 2021). "Kinematic self-replication in reconfigurable organisms". Proceedings of the National Academy of Sciences of the United States of America. 118 (49). Bibcode:2021PNAS..11812672K. doi:10.1073/pnas.2112672118. PMC 8670470. PMID 34845026.
- ^ Zeng J. "On the concept of systems bio-engineering". Coomunication on Transgenic Animals, June 1994, CAS, PRC. 6.
- ^ Chopra P, Kamma A. "Engineering life through Synthetic Biology". In Silico Biology. 6.
- ^ Flores Bueso Y, Tangney M (May 2017). "Synthetic Biology in the Driving Seat of the Bioeconomy". Trends in Biotechnology. 35 (5): 373–378. doi:10.1016/j.tibtech.2017.02.002. PMID 28249675.
- ^ Hunter D (October 2013). "How to object to radically new technologies on the basis of justice: the case of synthetic biology". Bioethics. 27 (8): 426–434. doi:10.1111/bioe.12049. PMID 24010854. S2CID 23895514.
- ^ Gutmann A (2011). "The ethics of synthetic biology: guiding principles for emerging technologies". The Hastings Center Report. 41 (4): 17–22. doi:10.1002/j.1552-146x.2011.tb00118.x. PMID 21845917. S2CID 20662786.
- ^ Channon K, Bromley EH, Woolfson DN (August 2008). "Synthetic biology through biomolecular design and engineering". Current Opinion in Structural Biology. 18 (4): 491–498. doi:10.1016/j.sbi.2008.06.006. PMID 18644449.
- ^ Duran-Nebreda S, Pla J, Vidiella B, Piñero J, Conde-Pueyo N, Solé R (February 2021). "Synthetic Lateral Inhibition in Periodic Pattern Forming Microbial Colonies". ACS Synthetic Biology. 10 (2): 277–285. doi:10.1021/acssynbio.0c00318. PMC 8486170. PMID 33449631. S2CID 231624252.
- ^ van Warmerdam T (December 25, 2021). "Exploring biotechnological advancements in the exogenous biological production of industrial small molecules and pharmaceuticals". YourBioHelper.com.
- ^ Stone M (2006). "Life Redesigned to Suit the Engineering Crowd" (PDF). Microbe. 1 (12): 566–570. S2CID 7171812. Archived from the original (PDF) on 2018-06-12.
- ^ a b c d e f g h i j k l m n o p q r Deplazes, Anna (May 2009). "Piecing together a puzzle: An exposition of synthetic biology". EMBO Reports. 10 (5): 428–432. doi:10.1038/embor.2009.76. ISSN 1469-221X. PMC 2680885. PMID 19415076.
- ^ a b c d e f g Acevedo-Rocha, Carlos G. (2016), Hagen, Kristin; Engelhard, Margret; Toepfer, Georg (eds.), "The Synthetic Nature of Biology", Ambivalences of Creating Life: Societal and Philosophical Dimensions of Synthetic Biology, Ethics of Science and Technology Assessment, vol. 45, Cham: Springer International Publishing, pp. 9–53, doi:10.1007/978-3-319-21088-9_2, ISBN 978-3-319-21088-9, PMC 7123144, retrieved 2023-01-23
- ^ Benner SA, Sismour AM (July 2005). "Synthetic biology". Nature Reviews. Genetics. 6 (7): 533–543. doi:10.1038/nrg1637. PMC 7097405. PMID 15995697.
- ^ Schwille, Petra (2011-09-02). "Bottom-Up Synthetic Biology: Engineering in a Tinkerer's World". Science. 333 (6047): 1252–1254. Bibcode:2011Sci...333.1252S. doi:10.1126/science.1211701. ISSN 0036-8075. PMID 21885774. S2CID 43354332.
- ^ Schwille P (September 2011). "Bottom-up synthetic biology: engineering in a tinkerer's world". Science. 333 (6047): 1252–4. Bibcode:2011Sci...333.1252S. doi:10.1126/science.1211701. PMID 21885774. S2CID 43354332.
- ^ Baker D, Church G, Collins J, Endy D, Jacobson J, Keasling J, et al. (June 2006). "Engineering life: building a fab for biology". Scientific American. 294 (6): 44–51. Bibcode:2006SciAm.294f..44B. doi:10.1038/scientificamerican0606-44. PMID 16711359.
- ^ Hughes RA, Ellington AD (January 2017). "Synthetic DNA Synthesis and Assembly: Putting the Synthetic in Synthetic Biology". Cold Spring Harbor Perspectives in Biology. 9 (1): a023812. doi:10.1101/cshperspect.a023812. PMC 5204324. PMID 28049645.
- ^ Kosuri S, Church GM (May 2014). "Large-scale de novo DNA synthesis: technologies and applications". Nature Methods. 11 (5): 499–507. doi:10.1038/nmeth.2918. PMC 7098426. PMID 24781323.
- ^ Blight KJ, Kolykhalov AA, Rice CM (December 2000). "Efficient initiation of HCV RNA replication in cell culture". Science. 290 (5498): 1972–4. Bibcode:2000Sci...290.1972B. doi:10.1126/science.290.5498.1972. PMID 11110665.
- ^ Couzin J (July 2002). "Virology. Active poliovirus baked from scratch". Science. 297 (5579): 174–5. doi:10.1126/science.297.5579.174b. PMID 12114601. S2CID 83531627.
- ^ Smith HO, Hutchison CA, Pfannkoch C, Venter JC (December 2003). "Generating a synthetic genome by whole genome assembly: phiX174 bacteriophage from synthetic oligonucleotides". Proceedings of the National Academy of Sciences of the United States of America. 100 (26): 15440–5. Bibcode:2003PNAS..10015440S. doi:10.1073/pnas.2237126100. PMC 307586. PMID 14657399.
- ^ Wade N (2007-06-29). "Scientists Transplant Genome of Bacteria". The New York Times. ISSN 0362-4331. Retrieved 2007-12-28.
- ^ Gibson DG, Benders GA, Andrews-Pfannkoch C, Denisova EA, Baden-Tillson H, Zaveri J, Stockwell TB, Brownley A, Thomas DW, Algire MA, Merryman C, Young L, Noskov VN, Glass JI, Venter JC, Hutchison CA, Smith HO (February 2008). "Complete chemical synthesis, assembly, and cloning of a Mycoplasma genitalium genome". Science. 319 (5867): 1215–20. Bibcode:2008Sci...319.1215G. doi:10.1126/science.1151721. PMID 18218864. S2CID 8190996.
- ^ Ball P (2016). "Man Made: A History of Synthetic Life". Distillations. 2 (1): 15–23. Retrieved 22 March 2018.
- ^ Pollack A (2007-09-12). "How Do You Like Your Genes? Biofabs Take Orders". The New York Times. ISSN 0362-4331. Retrieved 2007-12-28.
- ^ "Synthetic Biology Projects". arep.med.harvard.edu. Retrieved 2018-02-17.
- ^ Forster AC, Church GM (2006-08-22). "Towards synthesis of a minimal cell". Molecular Systems Biology. 2 (1): 45. doi:10.1038/msb4100090. PMC 1681520. PMID 16924266.
- ^ a b Basulto D (November 4, 2015). "Everything you need to know about why CRISPR is such a hot technology". Washington Post. Retrieved 5 December 2015.
- ^ Kahn J (November 9, 2015). "The Crispr Quandary". The New York Times. Retrieved 5 December 2015.
- ^ Ledford H (June 3, 2015). "CRISPR, the disruptor". Nature. 522 (7554): 20–4. Bibcode:2015Natur.522...20L. doi:10.1038/522020a. PMID 26040877.
{{cite journal}}: Unknown parameter|agency=ignored (help) - ^ Higginbotham S (4 December 2015). "Top VC Says Gene Editing Is Riskier Than Artificial Intelligence". Fortune. Retrieved 5 December 2015.
- ^ Rollie; et al. (2012). "Designing biological systems: Systems Engineering meets Synthetic Biology". Chemical Engineering Science. 69 (1): 1–29. Bibcode:2012ChEnS..69....1R. doi:10.1016/j.ces.2011.10.068.
- ^ a b Khalil AS, Collins JJ (May 2010). "Synthetic biology: applications come of age". Nature Reviews. Genetics. 11 (5): 367–379. doi:10.1038/nrg2775. PMC 2896386. PMID 20395970.
- ^ a b Freemont PS, Kitney TI (2012). Synthetic Biology – A Primer. World Scientific. doi:10.1142/p837. ISBN 978-1-84816-863-3.
- ^ a b Wright BW, Molloy MP, Jaschke PR (March 2022). "Overlapping genes in natural and engineered genomes". Nature Reviews. Genetics. 23 (3): 154–168. doi:10.1038/s41576-021-00417-w. PMC 8490965. PMID 34611352.
- ^ a b Chan LY, Kosuri S, Endy D (2005). "Refactoring bacteriophage T7". Molecular Systems Biology. 1: 2005.0018. doi:10.1038/msb4100025. PMC 1681472. PMID 16729053.
- ^ Temme K, Zhao D, Voigt CA (May 2012). "Refactoring the nitrogen fixation gene cluster from Klebsiella oxytoca". Proceedings of the National Academy of Sciences of the United States of America. 109 (18): 7085–7090. Bibcode:2012PNAS..109.7085T. doi:10.1073/pnas.1120788109. PMC 3345007. PMID 22509035.
- ^ Song M, Sukovich DJ, Ciccarelli L, Mayr J, Fernandez-Rodriguez J, Mirsky EA, et al. (May 2017). "Control of type III protein secretion using a minimal genetic system". Nature Communications. 8 (1): 14737. Bibcode:2017NatCo...814737S. doi:10.1038/ncomms14737. PMC 5436071. PMID 28485369.
- ^ Jaschke PR, Lieberman EK, Rodriguez J, Sierra A, Endy D (December 2012). "A fully decompressed synthetic bacteriophage øX174 genome assembled and archived in yeast". Virology. 434 (2): 278–284. doi:10.1016/j.virol.2012.09.020. PMID 23079106.
- ^ Woolfson DN, Bartlett GJ, Bruning M, Thomson AR (August 2012). "New currency for old rope: from coiled-coil assemblies to α-helical barrels". Current Opinion in Structural Biology. 22 (4): 432–41. doi:10.1016/j.sbi.2012.03.002. PMID 22445228.
- ^ Dueber JE, Wu GC, Malmirchegini GR, Moon TS, Petzold CJ, Ullal AV, Prather KL, Keasling JD (August 2009). "Synthetic protein scaffolds provide modular control over metabolic flux". Nature Biotechnology. 27 (8): 753–9. doi:10.1038/nbt.1557. PMID 19648908. S2CID 2756476.
- ^ Reddington SC, Howarth M (December 2015). "Secrets of a covalent interaction for biomaterials and biotechnology: SpyTag and SpyCatcher". Current Opinion in Chemical Biology. 29: 94–9. doi:10.1016/j.cbpa.2015.10.002. PMID 26517567.
- ^ Bayle JH, Grimley JS, Stankunas K, Gestwicki JE, Wandless TJ, Crabtree GR (January 2006). "Rapamycin analogs with differential binding specificity permit orthogonal control of protein activity". Chemistry & Biology. 13 (1): 99–107. doi:10.1016/j.chembiol.2005.10.017. PMID 16426976.
- ^ Altszyler E, Ventura A, Colman-Lerner A, Chernomoretz A (October 2014). "Impact of upstream and downstream constraints on a signaling module's ultrasensitivity". Physical Biology. 11 (6): 066003. Bibcode:2014PhBio..11f6003A. doi:10.1088/1478-3975/11/6/066003. PMC 4233326. PMID 25313165.
- ^ Altszyler E, Ventura AC, Colman-Lerner A, Chernomoretz A (2017). "Ultrasensitivity in signaling cascades revisited: Linking local and global ultrasensitivity estimations". PLOS ONE. 12 (6): e0180083. arXiv:1608.08007. Bibcode:2017PLoSO..1280083A. doi:10.1371/journal.pone.0180083. PMC 5491127. PMID 28662096.
- ^ Carbonell-Ballestero M, Duran-Nebreda S, Montañez R, Solé R, Macía J, Rodríguez-Caso C (December 2014). "A bottom-up characterization of transfer functions for synthetic biology designs: lessons from enzymology". Nucleic Acids Research. 42 (22): 14060–14069. doi:10.1093/nar/gku964. PMC 4267673. PMID 25404136.
- ^ Kaznessis YN (November 2007). "Models for synthetic biology". BMC Systems Biology. 1 (1): 47. doi:10.1186/1752-0509-1-47. PMC 2194732. PMID 17986347.
- ^ Tuza ZA, Singhal V, Kim J, Murray RM (December 2013). "An in silico modeling toolbox for rapid prototyping of circuits in a biomolecular "breadboard" system.". 52nd IEEE Conference on Decision and Control. doi:10.1109/CDC.2013.6760079.
- ^ LaFleur TL, Hossain A, Salis HM (September 2022). "Automated model-predictive design of synthetic promoters to control transcriptional profiles in bacteria". Nature Communications. 13 (1): 5159. Bibcode:2022NatCo..13.5159L. doi:10.1038/s41467-022-32829-5. PMC 9440211. PMID 36056029.
- ^ Kaznessis, Yiannis N (2007). "Models for synthetic biology". BMC Systems Biology. 1 (1): 47. doi:10.1186/1752-0509-1-47. ISSN 1752-0509. PMC 2194732. PMID 17986347.
- ^ Elani Y (June 2016). "Construction of membrane-bound artificial cells using microfluidics: a new frontier in bottom-up synthetic biology". Biochemical Society Transactions. 44 (3): 723–30. doi:10.1042/BST20160052. PMC 4900754. PMID 27284034.
- ^ Gach PC, Iwai K, Kim PW, Hillson NJ, Singh AK (October 2017). "Droplet microfluidics for synthetic biology" (PDF). Lab on a Chip. 17 (20): 3388–3400. doi:10.1039/C7LC00576H. OSTI 1421856. PMID 28820204. S2CID 206094816.
- ^ Vinuselvi P, Park S, Kim M, Park JM, Kim T, Lee SK (2011-06-03). "Microfluidic technologies for synthetic biology". International Journal of Molecular Sciences. 12 (6): 3576–93. doi:10.3390/ijms12063576. PMC 3131579. PMID 21747695.
- ^ a b Khalil AS, Lu TK, Bashor CJ, Ramirez CL, Pyenson NC, Joung JK, Collins JJ (August 2012). "A synthetic biology framework for programming eukaryotic transcription functions". Cell. 150 (3): 647–58. doi:10.1016/j.cell.2012.05.045. PMC 3653585. PMID 22863014.
- ^ "Synthetic Biology". Genome.gov. Retrieved 2023-01-23.
- ^ de Almeida PE, van Rappard JR, Wu JC (September 2011). "In vivo bioluminescence for tracking cell fate and function". American Journal of Physiology. Heart and Circulatory Physiology. 301 (3): H663–71. doi:10.1152/ajpheart.00337.2011. PMC 3191083. PMID 21666118.
- ^ Close DM, Xu T, Sayler GS, Ripp S (2011). "In vivo bioluminescent imaging (BLI): noninvasive visualization and interrogation of biological processes in living animals". Sensors. 11 (1): 180–206. doi:10.3390/s110100180. PMC 3274065. PMID 22346573.
- ^ Gibbs WW (1997). "Critters on a Chip". Scientific American. Retrieved 2 Mar 2009.
- ^ Belkin S, Yagur-Kroll S, Kabessa Y, Korouma V, Septon T, Anati Y, et al. (April 2017). "Remote detection of buried landmines using a bacterial sensor". Nature Biotechnology. 35 (4): 308–310. doi:10.1038/nbt.3791. PMID 28398330. S2CID 3645230.
- ^ Danino T, Prindle A, Kwong GA, Skalak M, Li H, Allen K, Hasty J, Bhatia SN (May 2015). "Programmable probiotics for detection of cancer in urine". Science Translational Medicine. 7 (289): 289ra84. doi:10.1126/scitranslmed.aaa3519. PMC 4511399. PMID 26019220.
- ^ "Face masks that can diagnose COVID-19". medicalxpress.com. Retrieved 11 July 2021.
- ^ Nguyen PQ, Soenksen LR, Donghia NM, Angenent-Mari NM, de Puig H, Huang A, et al. (November 2021). "Wearable materials with embedded synthetic biology sensors for biomolecule detection". Nature Biotechnology. 39 (11): 1366–1374. doi:10.1038/s41587-021-00950-3. hdl:1721.1/131278. PMID 34183860. S2CID 235673261.
- ^ Khalil, Ahmad S.; Collins, James J. (May 2010). "Synthetic biology: applications come of age". Nature Reviews Genetics. 11 (5): 367–379. doi:10.1038/nrg2775. ISSN 1471-0064. PMC 2896386. PMID 20395970.
- ^ "A Closer Look at Cellular Agriculture and the Processes Defining It - AgFunderNews". 2016-07-05. Retrieved 2016-08-05.
- ^ Bryant, Christopher J (3 August 2020). "Culture, meat, and cultured meat". Journal of Animal Science. 98 (8): skaa172. doi:10.1093/jas/skaa172. ISSN 0021-8812. PMC 7398566. PMID 32745186.
- ^ Hong, Tae Kyung; Shin, Dong-Min; Choi, Joonhyuk; Do, Jeong Tae; Han, Sung Gu (May 2021). "Current Issues and Technical Advances in Cultured Meat Production: AReview". Food Science of Animal Resources. 41 (3): 355–372. doi:10.5851/kosfa.2021.e14. ISSN 2636-0772. PMC 8112310. PMID 34017947.
- ^ Treich, Nicolas (1 May 2021). "Cultured Meat: Promises and Challenges". Environmental and Resource Economics. 79 (1): 33–61. doi:10.1007/s10640-021-00551-3. ISSN 1573-1502. PMC 7977488. PMID 33758465.
- ^ Mattick, CS (January 2018). "Cellular agriculture: The coming revolution in food production". Bulletin of the Atomic Scientists. 74 (1): 32–35. Bibcode:2018BuAtS..74a..32M. doi:10.1080/00963402.2017.1413059. S2CID 149404346.
- ^ "Japanese scientists produce first 3D-bioprinted, marbled Wagyu beef". New Atlas. 25 August 2021. Retrieved 21 September 2021.
- ^ Kang DH, Louis F, Liu H, Shimoda H, Nishiyama Y, Nozawa H, et al. (August 2021). "Engineered whole cut meat-like tissue by the assembly of cell fibers using tendon-gel integrated bioprinting". Nature Communications. 12 (1): 5059. Bibcode:2021NatCo..12.5059K. doi:10.1038/s41467-021-25236-9. PMC 8385070. PMID 34429413.
- ^ Lavars N (20 September 2021). "Lab-grown coffee cuts out the beans and deforestation". New Atlas. Retrieved 18 October 2021.
- ^ "Eco-friendly, lab-grown coffee is on the way, but it comes with a catch". The Guardian. 16 October 2021. Retrieved 26 October 2021.
- ^ "Sustainable coffee grown in Finland". VTT News. 15 September 2021. Retrieved 18 October 2021.
- ^ "Growing food with air and solar power: More efficient than planting crops". phys.org. Retrieved 11 July 2021.
- ^ Leger D, Matassa S, Noor E, Shepon A, Milo R, Bar-Even A (June 2021). "Photovoltaic-driven microbial protein production can use land and sunlight more efficiently than conventional crops". Proceedings of the National Academy of Sciences of the United States of America. 118 (26): e2015025118. Bibcode:2021PNAS..11815025L. doi:10.1073/pnas.2015025118. PMC 8255800. PMID 34155098. S2CID 235595143.
- ^ "World-first artificial synthesis of starch from CO2 outperforms nature". New Atlas. 28 September 2021. Retrieved 18 October 2021.
- ^ Cai T, Sun H, Qiao J, Zhu L, Zhang F, Zhang J, et al. (September 2021). "Cell-free chemoenzymatic starch synthesis from carbon dioxide". Science. 373 (6562): 1523–1527. Bibcode:2021Sci...373.1523C. doi:10.1126/science.abh4049. PMID 34554807. S2CID 237615280.
- ^ "Spider silk made by photosynthetic bacteria". phys.org. Archived from the original on 7 August 2020. Retrieved 16 August 2020.
- ^ Foong CP, Higuchi-Takeuchi M, Malay AD, Oktaviani NA, Thagun C, Numata K (July 2020). "A marine photosynthetic microbial cell factory as a platform for spider silk production". Communications Biology. 3 (1). Springer Science and Business Media LLC: 357. doi:10.1038/s42003-020-1099-6. PMC 7343832. PMID 32641733.
- ^ Singh V (December 2014). "Recent advances and opportunities in synthetic logic gates engineering in living cells". Systems and Synthetic Biology. 8 (4): 271–82. doi:10.1007/s11693-014-9154-6. PMC 4571725. PMID 26396651.
- ^ Purcell O, Lu TK (October 2014). "Synthetic analog and digital circuits for cellular computation and memory". Current Opinion in Biotechnology. Cell and Pathway Engineering. 29: 146–55. doi:10.1016/j.copbio.2014.04.009. PMC 4237220. PMID 24794536.
- ^ Daniel R, Rubens JR, Sarpeshkar R, Lu TK (May 2013). "Synthetic analog computation in living cells". Nature. 497 (7451): 619–23. Bibcode:2013Natur.497..619D. doi:10.1038/nature12148. PMID 23676681. S2CID 4358570.
- ^ Rinaudo K, Bleris L, Maddamsetti R, Subramanian S, Weiss R, Benenson Y (July 2007). "A universal RNAi-based logic evaluator that operates in mammalian cells". Nature Biotechnology. 25 (7): 795–801. doi:10.1038/nbt1307. PMID 17515909. S2CID 280451.
- ^ Xie Z, Wroblewska L, Prochazka L, Weiss R, Benenson Y (September 2011). "Multi-input RNAi-based logic circuit for identification of specific cancer cells". Science. 333 (6047): 1307–11. Bibcode:2011Sci...333.1307X. doi:10.1126/science.1205527. PMID 21885784. S2CID 13743291.
- ^ Nielsen AA, Der BS, Shin J, Vaidyanathan P, Paralanov V, Strychalski EA, Ross D, Densmore D, Voigt CA (April 2016). "Genetic circuit design automation". Science. 352 (6281): aac7341. doi:10.1126/science.aac7341. PMID 27034378.
- ^ Weinberg BH, Pham NT, Caraballo LD, Lozanoski T, Engel A, Bhatia S, Wong WW (May 2017). "Large-scale design of robust genetic circuits with multiple inputs and outputs for mammalian cells". Nature Biotechnology. 35 (5): 453–462. doi:10.1038/nbt.3805. PMC 5423837. PMID 28346402.
- ^ Pandi A, Koch M, Voyvodic PL, Soudier P, Bonnet J, Kushwaha M, Faulon JL (August 2019). "Metabolic perceptrons for neural computing in biological systems". Nature Communications. 10 (1): 3880. Bibcode:2019NatCo..10.3880P. doi:10.1038/s41467-019-11889-0. PMC 6713752. PMID 31462649.
- ^ Westfall PJ, Pitera DJ, Lenihan JR, Eng D, Woolard FX, Regentin R, Horning T, Tsuruta H, Melis DJ, Owens A, Fickes S, Diola D, Benjamin KR, Keasling JD, Leavell MD, McPhee DJ, Renninger NS, Newman JD, Paddon CJ (January 2012). "Production of amorphadiene in yeast, and its conversion to dihydroartemisinic acid, precursor to the antimalarial agent artemisinin". Proceedings of the National Academy of Sciences of the United States of America. 109 (3): E111–8. Bibcode:2012PNAS..109E.111W. doi:10.1073/pnas.1110740109. PMC 3271868. PMID 22247290.
- ^ Connor S (28 March 2014). "Eureka! Scientists unveil giant leap towards synthetic life". The Independent. Archived from the original on 2022-05-26. Retrieved 2015-08-06.
- ^ Nguyen PQ, Botyanszki Z, Tay PK, Joshi NS (September 2014). "Programmable biofilm-based materials from engineered curli nanofibres". Nature Communications. 5: 4945. Bibcode:2014NatCo...5.4945N. doi:10.1038/ncomms5945. PMID 25229329.
- ^ Kuhlman B, Dantas G, Ireton GC, Varani G, Stoddard BL, Baker D (November 2003). "Design of a novel globular protein fold with atomic-level accuracy". Science. 302 (5649): 1364–8. Bibcode:2003Sci...302.1364K. doi:10.1126/science.1089427. PMID 14631033. S2CID 1939390.
- ^ Koder RL, Anderson JL, Solomon LA, Reddy KS, Moser CC, Dutton PL (March 2009). "Design and engineering of an O(2) transport protein". Nature. 458 (7236): 305–309. Bibcode:2009Natur.458..305K. doi:10.1038/nature07841. PMC 3539743. PMID 19295603.
- ^ Farid TA, Kodali G, Solomon LA, Lichtenstein BR, Sheehan MM, Fry BA, et al. (December 2013). "Elementary tetrahelical protein design for diverse oxidoreductase functions". Nature Chemical Biology. 9 (12): 826–833. doi:10.1038/nchembio.1362. PMC 4034760. PMID 24121554.
- ^ Wang MS, Hecht MH (September 2020). "A Completely De Novo ATPase from Combinatorial Protein Design". Journal of the American Chemical Society. 142 (36): 15230–15234. doi:10.1021/jacs.0c02954. PMID 32833456. S2CID 221308093.
- ^ Armbruster BN, Li X, Pausch MH, Herlitze S, Roth BL (March 2007). "Evolving the lock to fit the key to create a family of G protein-coupled receptors potently activated by an inert ligand". Proceedings of the National Academy of Sciences of the United States of America. 104 (12): 5163–5168. Bibcode:2007PNAS..104.5163A. doi:10.1073/pnas.0700293104. PMC 1829280. PMID 17360345.
- ^ Mak WS, Tran S, Marcheschi R, Bertolani S, Thompson J, Baker D, et al. (November 2015). "Integrative genomic mining for enzyme function to enable engineering of a non-natural biosynthetic pathway". Nature Communications. 6: 10005. Bibcode:2015NatCo...610005M. doi:10.1038/ncomms10005. PMC 4673503. PMID 26598135.
- ^ Wang Q, Parrish AR, Wang L (March 2009). "Expanding the genetic code for biological studies". Chemistry & Biology. 16 (3): 323–36. doi:10.1016/j.chembiol.2009.03.001. PMC 2696486. PMID 19318213.
- ^ Davidson AR, Lumb KJ, Sauer RT (October 1995). "Cooperatively folded proteins in random sequence libraries". Nature Structural Biology. 2 (10): 856–864. doi:10.1038/nsb1095-856. PMID 7552709. S2CID 31781262.
- ^ Kamtekar S, Schiffer JM, Xiong H, Babik JM, Hecht MH (December 1993). "Protein design by binary patterning of polar and nonpolar amino acids". Science. 262 (5140): 1680–1685. Bibcode:1993Sci...262.1680K. doi:10.1126/science.8259512. PMID 8259512.
- ^ Walter KU, Vamvaca K, Hilvert D (November 2005). "An active enzyme constructed from a 9-amino acid alphabet". The Journal of Biological Chemistry. 280 (45): 37742–37746. doi:10.1074/jbc.M507210200. PMID 16144843.
- ^ "Synthetic Biology Applications". www.thermofisher.com. Retrieved 2015-11-12.
- ^ Liu Y, Shin HD, Li J, Liu L (February 2015). "Toward metabolic engineering in the context of system biology and synthetic biology: advances and prospects". Applied Microbiology and Biotechnology. 99 (3): 1109–18. doi:10.1007/s00253-014-6298-y. PMID 25547833. S2CID 954858.
- ^ Church GM, Gao Y, Kosuri S (September 2012). "Next-generation digital information storage in DNA". Science. 337 (6102): 1628. Bibcode:2012Sci...337.1628C. doi:10.1126/science.1226355. PMID 22903519. S2CID 934617.
- ^ "Huge amounts of data can be stored in DNA". Sky News. 23 January 2013. Archived from the original on 2016-05-31. Retrieved 24 January 2013.
- ^ Zadeh JN, Steenberg CD, Bois JS, Wolfe BR, Pierce MB, Khan AR, et al. (January 2011). "NUPACK: Analysis and design of nucleic acid systems". Journal of Computational Chemistry. 32 (1): 170–173. doi:10.1002/jcc.21596. PMID 20645303. S2CID 33709556.
- ^ Lorenz R, Bernhart SH, Höner Zu Siederdissen C, Tafer H, Flamm C, Stadler PF, Hofacker IL (November 2011). "ViennaRNA Package 2.0". Algorithms for Molecular Biology. 6 (1): 26. doi:10.1186/1748-7188-6-26. PMC 3319429. PMID 22115189.
- ^ Salis HM, Mirsky EA, Voigt CA (October 2009). "Automated design of synthetic ribosome binding sites to control protein expression". Nature Biotechnology. 27 (10): 946–950. doi:10.1038/nbt.1568. PMC 2782888. PMID 19801975.
- ^ Nielsen AA, Der BS, Shin J, Vaidyanathan P, Paralanov V, Strychalski EA, et al. (April 2016). "Genetic circuit design automation". Science. 352 (6281): aac7341. doi:10.1126/science.aac7341. PMID 27034378.
- ^ Hossain A, Lopez E, Halper SM, Cetnar DP, Reis AC, Strickland D, et al. (December 2020). "Automated design of thousands of nonrepetitive parts for engineering stable genetic systems". Nature Biotechnology. 38 (12): 1466–1475. doi:10.1038/s41587-020-0584-2. PMID 32661437. S2CID 220506228.
- ^ Pollack A (May 7, 2014). "Researchers Report Breakthrough in Creating Artificial Genetic Code". New York Times. Retrieved May 7, 2014.
- ^ Callaway E (May 7, 2014). "First life with 'alien' DNA". Nature. doi:10.1038/nature.2014.15179. S2CID 86967999. Retrieved May 7, 2014.
- ^ a b Malyshev DA, Dhami K, Lavergne T, Chen T, Dai N, Foster JM, Corrêa IR, Romesberg FE (May 2014). "A semi-synthetic organism with an expanded genetic alphabet". Nature. 509 (7500): 385–8. Bibcode:2014Natur.509..385M. doi:10.1038/nature13314. PMC 4058825. PMID 24805238.
- ^ a b Verseux CN, Paulino-Lima IG, Baqué M, Billi D, Rothschild LJ (2016). "Synthetic Biology for Space Exploration: Promises and Societal Implications". Ambivalences of Creating Life. Ethics of Science and Technology Assessment. Vol. 45. Springer-Verlag. pp. 73–100. doi:10.1007/978-3-319-21088-9_4. ISBN 978-3-319-21087-2.
- ^ Menezes AA, Cumbers J, Hogan JA, Arkin AP (January 2015). "Towards synthetic biological approaches to resource utilization on space missions". Journal of the Royal Society, Interface. 12 (102): 20140715. doi:10.1098/rsif.2014.0715. PMC 4277073. PMID 25376875.
- ^ Montague M, McArthur GH, Cockell CS, Held J, Marshall W, Sherman LA, et al. (December 2012). "The role of synthetic biology for in situ resource utilization (ISRU)". Astrobiology. 12 (12): 1135–1142. Bibcode:2012AsBio..12.1135M. doi:10.1089/ast.2012.0829. PMID 23140229.
- ^ Steigerwald B. "NASA – Designer Plants on Mars". www.nasa.gov. Retrieved 2020-05-29.
- ^ Hutchison CA, Chuang RY, Noskov VN, Assad-Garcia N, Deerinck TJ, Ellisman MH, Gill J, Kannan K, Karas BJ, Ma L, Pelletier JF, Qi ZQ, Richter RA, Strychalski EA, Sun L, Suzuki Y, Tsvetanova B, Wise KS, Smith HO, Glass JI, Merryman C, Gibson DG, Venter JC (March 2016). "Design and synthesis of a minimal bacterial genome". Science. 351 (6280): aad6253. Bibcode:2016Sci...351.....H. doi:10.1126/science.aad6253. PMID 27013737.
- ^ a b Connor S (1 December 2014). "Major synthetic life breakthrough as scientists make the first artificial enzymes". The Independent. London. Archived from the original on 2022-05-26. Retrieved 2015-08-06.
- ^ a b Deamer D (July 2005). "A giant step towards artificial life?". Trends in Biotechnology. 23 (7): 336–338. doi:10.1016/j.tibtech.2005.05.008. PMID 15935500.
- ^ "Scientists Reach Milestone On Way To Artificial Life". NPR. 2010-05-20. Retrieved 2010-06-09.
- ^ Venter JC (25 January 2017). "From Designing Life to Prolonging Healthy Life". YouTube. University of California Television (UCTV). Archived from the original on 2021-12-21. Retrieved 1 February 2017.
- ^ "Build-a-Cell". Retrieved 4 Dec 2019.
- ^ "FabriCell". Retrieved 8 Dec 2019.
- ^ "MaxSynBio – Max Planck Research Network in Synthetic Biology". Retrieved 8 Dec 2019.
- ^ "BaSyC". Retrieved 8 Dec 2019.
- ^ "SynCell EU". Retrieved 8 Dec 2019.
- ^ Devlin, Hannah; correspondent, Hannah Devlin Science (2023-06-14). "Synthetic human embryos created in groundbreaking advance". The Guardian. ISSN 0261-3077. Retrieved 2023-11-30.
{{cite news}}:|last2=has generic name (help) - ^ a b c Khalil, Ahmad S.; Collins, James J. (May 2010). "Synthetic biology: applications come of age". Nature Reviews Genetics. 11 (5): 367–379. doi:10.1038/nrg2775. ISSN 1471-0056. PMC 2896386. PMID 20395970. S2CID 2926016.
- ^ Zu C, Wang J (August 2014). "Tumor-colonizing bacteria: a potential tumor targeting therapy". Critical Reviews in Microbiology. 40 (3): 225–235. doi:10.3109/1040841X.2013.776511. PMID 23964706. S2CID 26498221.
- ^ Gujrati V, Kim S, Kim SH, Min JJ, Choy HE, Kim SC, Jon S (February 2014). "Bioengineered bacterial outer membrane vesicles as cell-specific drug-delivery vehicles for cancer therapy". ACS Nano. 8 (2): 1525–1537. doi:10.1021/nn405724x. PMID 24410085.
- ^ Piñero-Lambea C, Bodelón G, Fernández-Periáñez R, Cuesta AM, Álvarez-Vallina L, Fernández LÁ (April 2015). "Programming controlled adhesion of E. coli to target surfaces, cells, and tumors with synthetic adhesins". ACS Synthetic Biology. 4 (4): 463–473. doi:10.1021/sb500252a. PMC 4410913. PMID 25045780.
- ^ Deyneko IV, Kasnitz N, Leschner S, Weiss S (2016). "Composing a Tumor Specific Bacterial Promoter". PLOS ONE. 11 (5): e0155338. Bibcode:2016PLoSO..1155338D. doi:10.1371/journal.pone.0155338. PMC 4865170. PMID 27171245.
- ^ Rice KC, Bayles KW (March 2008). "Molecular control of bacterial death and lysis". Microbiology and Molecular Biology Reviews. 72 (1): 85–109, table of contents. doi:10.1128/mmbr.00030-07. PMC 2268280. PMID 18322035.
- ^ Ganai S, Arenas RB, Forbes NS (November 2009). "Tumour-targeted delivery of TRAIL using Salmonella typhimurium enhances breast cancer survival in mice". British Journal of Cancer. 101 (10): 1683–1691. doi:10.1038/sj.bjc.6605403. PMC 2778534. PMID 19861961.
- ^ Scott, Benjamin M.; Gutiérrez-Vázquez, Cristina; Sanmarco, Liliana M.; da Silva Pereira, Jessica A.; Li, Zhaorong; Plasencia, Agustín; Hewson, Patrick; Cox, Laura M.; O’Brien, Madelynn; Chen, Steven K.; Moraes-Vieira, Pedro M.; Chang, Belinda S. W.; Peisajovich, Sergio G.; Quintana, Francisco J. (July 2021). "Self-tunable engineered yeast probiotics for the treatment of inflammatory bowel disease". Nature Medicine. 27 (7): 1212–1222. doi:10.1038/s41591-021-01390-x. ISSN 1546-170X. PMID 34183837. S2CID 235673499.
- ^ Chen, Kevin; Zhu, Yixuan; Zhang, Yongrong; Hamza, Therwa; Yu, Hua; Saint Fleur, Ashley; Galen, James; Yang, Zhiyong; Feng, Hanping (2020-10-28). "A probiotic yeast-based immunotherapy against Clostridioides difficile infection". Science Translational Medicine. 12 (567): eaax4905. doi:10.1126/scitranslmed.aax4905. ISSN 1946-6234. PMC 7692727. PMID 33115949.
- ^ Chen, Yi-Ju; Abila, Bams; Mostafa Kamel, Yasser (2023). "CAR-T: What Is Next?". Cancers. 15 (3): 663. doi:10.3390/cancers15030663. ISSN 2072-6694. PMC 9913679. PMID 36765623.
- ^ Jones, B.S., Lamb, L.S., Goldman, F. & Di Stasi, A. Improving the safety of cell therapy products by suicide gene transfer. Front. Pharmacol. 5, 254 (2014).
- ^ Wei P, Wong WW, Park JS, Corcoran EE, Peisajovich SG, Onuffer JJ, et al. (August 2012). "Bacterial virulence proteins as tools to rewire kinase pathways in yeast and immune cells". Nature. 488 (7411): 384–388. Bibcode:2012Natur.488..384W. doi:10.1038/nature11259. PMC 3422413. PMID 22820255.
- ^ Danino T, Mondragón-Palomino O, Tsimring L, Hasty J (January 2010). "A synchronized quorum of genetic clocks". Nature. 463 (7279): 326–330. Bibcode:2010Natur.463..326D. doi:10.1038/nature08753. PMC 2838179. PMID 20090747.
- ^ Chen YY, Jensen MC, Smolke CD (May 2010). "Genetic control of mammalian T-cell proliferation with synthetic RNA regulatory systems". Proceedings of the National Academy of Sciences of the United States of America. 107 (19): 8531–8536. Bibcode:2010PNAS..107.8531C. doi:10.1073/pnas.1001721107. PMC 2889348. PMID 20421500.
- ^ a b Katz, Leonard; Chen, Yvonne Y; Gonzalez, Ramon; Peterson, Todd C; Zhao, Huimin; Baltz, Richard H (2018-07-01). "Synthetic biology advances and applications in the biotechnology industry: a perspective". Journal of Industrial Microbiology and Biotechnology. 45 (7): 449–461. doi:10.1007/s10295-018-2056-y. ISSN 1476-5535. PMID 29915997. S2CID 49296740.
- ^ Hofer M, Lutolf MP (May 2021). "Engineering organoids". Nature Reviews. Materials. 6 (5): 402–420. Bibcode:2021NatRM...6..402H. doi:10.1038/s41578-021-00279-y. PMC 7893133. PMID 33623712.
- ^ Whitaker, Matthew (July 2014). "The history of 3D printing in healthcare". The Bulletin of the Royal College of Surgeons of England. 96 (7): 228–229. doi:10.1308/147363514X13990346756481. ISSN 1473-6357.
- ^ Atala A, Bauer SB, Soker S, Yoo JJ, Retik AB (April 2006). "Tissue-engineered autologous bladders for patients needing cystoplasty". Lancet. 367 (9518): 1241–6. doi:10.1016/S0140-6736(06)68438-9. PMID 16631879. S2CID 17892321.
- ^ Hong N, Yang GH, Lee J, Kim G (January 2018). "3D bioprinting and its in vivo applications". Journal of Biomedical Materials Research Part B: Applied Biomaterials. 106 (1): 444–459. doi:10.1002/jbm.b.33826. PMID 28106947.
- ^ Shinkar, Kalyani; Rhode, Kawal (August 1, 2022). "Could 3D extrusion bioprinting serve to be a real alternative to organ transplantation in the future?". Annals of 3D Printed Medicine. 7: 100066. doi:10.1016/j.stlm.2022.100066. ISSN 2666-9641. S2CID 249083907.
- ^ "90-OR: 3D Bioprinting of a Bionic Pancreas with a Vascular System—Results of Transplantation in Large Animals". diabetesjournals.org. Retrieved October 26, 2023.
- ^ Sommer AC, Blumenthal EZ (September 2019). "Implementations of 3D printing in ophthalmology". Graefe's Archive for Clinical and Experimental Ophthalmology. 257 (9): 1815–1822. doi:10.1007/s00417-019-04312-3. PMID 30993457. S2CID 116884575.
- ^ Klak, Marta; Bryniarski, Tomasz; Kowalska, Patrycja; Gomolka, Magdalena; Tymicki, Grzegorz; Kosowska, Katarzyna; Cywoniuk, Piotr; Dobrzanski, Tomasz; Turowski, Pawel; Wszola, Michal (June 30, 2020). "Novel Strategies in Artificial Organ Development: What Is the Future of Medicine?". Micromachines. 11 (7): 646. doi:10.3390/mi11070646. ISSN 2072-666X. PMC 7408042. PMID 32629779.
- ^ Cui H, Miao S, Esworthy T, Zhou X, Lee SJ, Liu C, Yu ZX, Fisher JP, Mohiuddin M, Zhang LG (July 2018). "3D bioprinting for cardiovascular regeneration and pharmacology". Advanced Drug Delivery Reviews. 132: 252–269. doi:10.1016/j.addr.2018.07.014. PMC 6226324. PMID 30053441.
- ^ "A Multicenter, Single Arm, Prospective, Open-Label, Staged Study of the Safety and Efficacy of the AuriNovo Construct for Auricular Reconstruction in Subjects With Unilateral Microtia". clinicaltrials.gov. October 15, 2021. Retrieved July 19, 2022.
- ^ Rabin, Roni Caryn (June 2, 2022). "Doctors Transplant Ear of Human Cells, Made by 3-D Printer". The New York Times. Retrieved July 19, 2022.
- ^ Cooper-White M (March 1, 2015). "How 3D Printing Could End The Deadly Shortage Of Donor Organs". Huffpost Science. TheHuffingtonPost.com, Inc. Retrieved February 17, 2016.
- ^ Edmundson MC, Capeness M, Horsfall L (December 2014). "Exploring the potential of metallic nanoparticles within synthetic biology". New Biotechnology. 31 (6): 572–578. doi:10.1016/j.nbt.2014.03.004. hdl:20.500.11820/5cd4fa26-dee8-4862-86af-cf6a79546a13. PMID 24681407. S2CID 15790244.
- ^ "Nanoparticle chomps away plaques that cause heart attacks". Michigan State University. 27 January 2020. Archived from the original on 29 January 2020. Retrieved 31 January 2020.
- ^ "Nanoparticle helps eat away deadly arterial plaque". New Atlas. 28 January 2020. Archived from the original on 1 March 2020. Retrieved 13 April 2020.
- ^ Flores AM, Hosseini-Nassab N, Jarr KU, Ye J, Zhu X, Wirka R, et al. (February 2020). "Pro-efferocytic nanoparticles are specifically taken up by lesional macrophages and prevent atherosclerosis". Nature Nanotechnology. 15 (2): 154–161. Bibcode:2020NatNa..15..154F. doi:10.1038/s41565-019-0619-3. PMC 7254969. PMID 31988506.
- ^ "Research creates hydrogen-producing living droplets, paving way for alternative future energy source". phys.org. Archived from the original on 16 December 2020. Retrieved 9 December 2020.
- ^ Xu Z, Wang S, Zhao C, Li S, Liu X, Wang L, et al. (November 2020). "Photosynthetic hydrogen production by droplet-based microbial micro-reactors under aerobic conditions". Nature Communications. 11 (1): 5985. Bibcode:2020NatCo..11.5985X. doi:10.1038/s41467-020-19823-5. PMC 7689460. PMID 33239636.
- ^ Xie, Mingqi; Fussenegger, Martin (July 2015). "Mammalian designer cells: Engineering principles and biomedical applications". Biotechnology Journal. 10 (7): 1005–1018. doi:10.1002/biot.201400642. ISSN 1860-6768. PMID 26010998. S2CID 11838264.
- ^ academic.oup.com https://academic.oup.com/proteincell/article/13/7/476/6747017. Retrieved 2024-01-31.
{{cite web}}: Missing or empty|title=(help) - ^ a b c Mansouri, Maysam; Fussenegger, Martin (2022-06-01). "Electrogenetics: Bridging synthetic biology and electronics to remotely control the behavior of mammalian designer cells". Current Opinion in Chemical Biology. 68: 102151. doi:10.1016/j.cbpa.2022.102151. hdl:20.500.11850/547194. ISSN 1367-5931. PMID 35483127.
- ^ Krawczyk, Krzysztof; Xue, Shuai; Buchmann, Peter; Charpin-El-Hamri, Ghislaine; Saxena, Pratik; Hussherr, Marie-Didiée; Shao, Jiawei; Ye, Haifeng; Xie, Mingqi; Fussenegger, Martin (2020-05-29). "Electrogenetic cellular insulin release for real-time glycemic control in type 1 diabetic mice". Science. 368 (6494): 993–1001. doi:10.1126/science.aau7187. hdl:20.500.11850/418642. ISSN 0036-8075. PMID 32467389. S2CID 218984797.
- ^ McCluskey, Jane T.; Hamid, Muhajir; Guo-Parke, Hong; McClenaghan, Neville H.; Gomis, Ramon; Flatt, Peter R. (2011-06-24). "Development and Functional Characterization of Insulin-releasing Human Pancreatic Beta Cell Lines Produced by Electrofusion*". Journal of Biological Chemistry. 286 (25): 21982–21992. doi:10.1074/jbc.M111.226795. ISSN 0021-9258. PMC 3121343. PMID 21515691.
- ^ a b c Newson AJ (2015). "Synthetic Biology: Ethics, Exceptionalism and Expectations". Macquarie Law Journal. 15: 45.
- ^ "Frontiers 2018/19: Emerging Issues of Environmental Concern". UN Environment Programme. 2019-03-04. Retrieved 2021-01-03.
- ^ "World's first gene-edited babies created in China, claims scientist". The Guardian. November 2018.
- ^ Häyry M (April 2017). "Synthetic Biology and Ethics: Past, Present, and Future". Cambridge Quarterly of Healthcare Ethics. 26 (2): 186–205. doi:10.1017/S0963180116000803. PMID 28361718. S2CID 29992939.
- ^ Jin S, Clark B, Kuznesof S, Lin X, Frewer LJ (September 2019). "Synthetic biology applied in the agrifood sector: Public perceptions, attitudes and implications for future studie". Trends in Food Science and Technology. 91: 454–466. doi:10.1016/j.tifs.2019.07.025. S2CID 199632852.
- ^ Gutmann A (2012). "The ethics of synthetic biology: guiding principles for emerging technologies". The Hastings Center Report. 41 (4): 17–22. doi:10.1002/j.1552-146X.2011.tb00118.x. PMID 21845917. S2CID 20662786.
- ^ Bryant CJ (August 2020). "Culture, meat, and cultured meat". Journal of Animal Science. 98 (8): skaa172. doi:10.1093/jas/skaa172. PMC 7398566. PMID 32745186.
- ^ Hong TK, Shin DM, Choi J, Do JT, Han SG (May 2021). "Current Issues and Technical Advances in Cultured Meat Production: A Review". Food Science of Animal Resources. 41 (3): 355–372. doi:10.5851/kosfa.2021.e14. PMC 8112310. PMID 34017947.
- ^ Treich N (1 May 2021). "Cultured Meat: Promises and Challenges". Environmental & Resource Economics. 79 (1): 33–61. doi:10.1007/s10640-021-00551-3. PMC 7977488. PMID 33758465.
- ^ Treich N (May 2021). "Cultured Meat: Promises and Challenges". Environmental & Resource Economics. 79 (1): 33–61. doi:10.1007/s10640-021-00551-3. PMC 7977488. PMID 33758465.
- ^ a b Howard J, Murashov V, Schulte P (March 2017). "Synthetic biology and occupational risk". Journal of Occupational and Environmental Hygiene. 14 (3): 224–236. doi:10.1080/15459624.2016.1237031. PMID 27754800. S2CID 205893358.
- ^ a b European Commission. Directorate General for Health Consumers (2016-02-12). Opinion on synthetic biology II: Risk assessment methodologies and safety aspects. Publications Office. doi:10.2772/63529. ISBN 9789279439162.
{{cite book}}:|website=ignored (help) - ^ a b Bügl H, Danner JP, Molinari RJ, Mulligan JT, Park HO, Reichert B, Roth DA, Wagner R, Budowle B, Scripp RM, Smith JA, Steele SJ, Church G, Endy D (June 2007). "DNA synthesis and biological security". Nature Biotechnology. 25 (6): 627–629. doi:10.1038/nbt0607-627. PMID 17557094. S2CID 7776829.
- ^ "Ethical Issues in Synthetic Biology: An Overview of the Debates" (PDF).
- ^ a b Presidential Commission for the study of Bioethical Issues, December 2010 NEW DIRECTIONS The Ethics of Synthetic Biology and Emerging Technologies Archived 2017-01-18 at the Wayback Machine Retrieved 2012-04-14.
- ^ Hummel K (2020-08-31). "Engineered Pathogens and Unnatural Biological Weapons: The Future Threat of Synthetic Biology". Combating Terrorism Center at West Point. Retrieved 2023-01-05.
- ^ Hummel K (2020-06-29). "A View from the CT Foxhole: A Virtual Roundtable on COVID-19 and Counterterrorism with Audrey Kurth Cronin, Lieutenant General (Ret) Michael Nagata, Magnus Ranstorp, Ali Soufan, and Juan Zarate". Combating Terrorism Center at West Point. Retrieved 2023-01-05.
- ^ a b "Synbiosafe". Synbiosafe.
- ^ Schmidt M, Ganguli-Mitra A, Torgersen H, Kelle A, Deplazes A, Biller-Andorno N (December 2009). "A priority paper for the societal and ethical aspects of synthetic biology" (PDF). Systems and Synthetic Biology. 3 (1–4): 3–7. doi:10.1007/s11693-009-9034-7. PMC 2759426. PMID 19816794.
- ^ Schmidt M. Kelle A. Ganguli A, de Vriend H. (Eds.) 2009. "Synthetic Biology. The Technoscience and its Societal Consequences". Springer Academic Publishing.
- ^ Kelle A (December 2009). "Ensuring the security of synthetic biology-towards a 5P governance strategy". Systems and Synthetic Biology. 3 (1–4): 85–90. doi:10.1007/s11693-009-9041-8. PMC 2759433. PMID 19816803.
- ^ Schmidt M (June 2008). "Diffusion of synthetic biology: a challenge to biosafety" (PDF). Systems and Synthetic Biology. 2 (1–2): 1–6. doi:10.1007/s11693-008-9018-z. PMC 2671588. PMID 19003431.
- ^ COSY: Communicating Synthetic Biology
- ^ Kronberger N, Holtz P, Kerbe W, Strasser E, Wagner W (December 2009). "Communicating Synthetic Biology: from the lab via the media to the broader public". Systems and Synthetic Biology. 3 (1–4): 19–26. doi:10.1007/s11693-009-9031-x. PMC 2759424. PMID 19816796.
- ^ Cserer A, Seiringer A (December 2009). "Pictures of Synthetic Biology: A reflective discussion of the representation of Synthetic Biology (SB) in the German-language media and by SB experts". Systems and Synthetic Biology. 3 (1–4): 27–35. doi:10.1007/s11693-009-9038-3. PMC 2759430. PMID 19816797.
- ^ Report of IASB "Technical solutions for biosecurity in synthetic biology" Archived July 19, 2011, at the Wayback Machine, Munich, 2008
- ^ Parens E., Johnston J., Moses J. Ethical Issues in Synthetic Biology. 2009.
- ^ "Policy and Global Affairs at the National Academies". sites.nationalacademies.org.
- ^ Presidential Commission for the study of Bioethical Issues, December 2010 FAQ Archived 2015-12-24 at the Wayback Machine
- ^ "Synthetic Biology F.A.Q.'s | Presidential Commission for the Study of Bioethical Issues". Archived from the original on 2015-12-24. Retrieved 2014-10-11.
- ^ a b Erickson B, Singh R, Winters P (September 2011). "Synthetic biology: regulating industry uses of new biotechnologies". Science. 333 (6047): 1254–6. Bibcode:2011Sci...333.1254E. doi:10.1126/science.1211066. PMID 21885775. S2CID 1568198.
- ^ Sayler KM (June 8, 2021). Defense Primer: Emerging Technologies (PDF) (Report). Congressional Research Service. Retrieved July 22, 2021.
- ^ Katherine Xue for Harvard Magazine. September–October 2014 Synthetic Biology's New Menagerie
- ^ Yojana Sharma for Scidev.net March 15, 2012. NGOs call for international regulation of synthetic biology
- ^ The New Synthetic Biology: Who Gains? (2014-05-08), Richard C. Lewontin, New York Review of Books
- ^ National Academies Of Sciences; Division on Earth Life Studies; Board On Life Sciences; Board on Chemical Sciences Technology; Committee on Strategies for Identifying Addressing Potential Biodefense Vulnerabilities Posed by Synthetic Biology (2018-06-19). Biodefense in the Age of Synthetic Biology. National Academies of Sciences, Engineering, and Medicine. doi:10.17226/24890. ISBN 9780309465182. PMID 30629396. S2CID 90767286.
- ^ "Future Brief: Synthetic biology and biodiversity". European Commission. September 2016. pp. 14–15. Retrieved 2019-01-14.
- ^ Final opinion on synthetic biology III: Risks to the environment and biodiversity related to synthetic biology and research priorities in the field of synthetic biology. European Commission. 2016-04-04. pp. 8, 27. ISBN 9789279549731. Retrieved 2019-01-14.
{{cite book}}:|website=ignored (help) - ^ El Karoui M, Hoyos-Flight M, Fletcher L (2019). "Future Trends in Synthetic Biology-A Report". Frontiers in Bioengineering and Biotechnology. 7: 175. doi:10.3389/fbioe.2019.00175. PMC 6692427. PMID 31448268.
- ^ Bailey C, Metcalf H, Crook B (2012). "Synthetic biology: A review of the technology, and current and future needs from the regulatory framework in Great Britain" (PDF). UK Health and Safety Executive. Retrieved 2018-11-29.
- ^ Pei L, Bar-Yam S, Byers-Corbin J, Casagrande R, Eichler F, Lin A, et al. (2012). "Regulatory Frameworks for Synthetic Biology". Synthetic Biology. John Wiley & Sons, Ltd. pp. 157–226. doi:10.1002/9783527659296.ch5. ISBN 9783527659296.
- ^ Trump BD (November 2017). "Synthetic biology regulation and governance: Lessons from TAPIC for the United States, European Union, and Singapore". Health Policy. 121 (11): 1139–1146. doi:10.1016/j.healthpol.2017.07.010. PMID 28807332.
Bibliography
[edit]- Church G, Regis E (2012). Regenesis:How Synthetic Biology will Reinvent Nature and Ourselves. New York, NY: Basic Books. ISBN 978-0465021758.
- Synthetic biology and biodiversity; Science for Environment Policy (PDF). Future Brief 15. Produced for the European Commission DG Environment by the Science Communication Unit, UWE, Bristol (Report). European Commission. 2016.
- Venter C (2013). Life at the Speed of Light: The Double Helix and the Dawn of Digital Life. New York, NY: Penguin Books. ISBN 978-0670025404. OCLC 834432832.
- Rutherford, Adam (2014). Creation : how science is reinventing life itself. Current. ISBN 9781617230110. OCLC 880230551.
External links
[edit]- Engineered Pathogens and Unnatural Biological Weapons: The Future Threat of Synthetic Biology . Threats and considerations
- Synthetic biology books popular science book and textbooks
- Introductory Summary of Synthetic Biology Archived 2018-04-02 at the Wayback Machine. Concise overview of synthetic biology concepts, developments and applications
- Collaborative overview article on Synthetic Biology
- Controversial DNA startup wants to let customers create creatures (2015-01-03), San Francisco Chronicle
- It's Alive, But Is It Life: Synthetic Biology and the Future of Creation (28 September 2016), World Science Festival

