Succinate dehydrogenase
| succinate dehydrogenase (succinate-ubiquinone oxidoreductase) | |||||||||
|---|---|---|---|---|---|---|---|---|---|
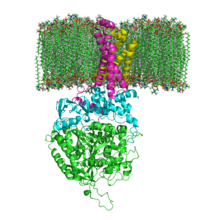 The structure of SQR in a phospholipid membrane. SDHA, SDHB, SDHC and SDHD | |||||||||
| Identifiers | |||||||||
| EC no. | 1.3.5.1 | ||||||||
| CAS no. | 9028-11-9 | ||||||||
| Databases | |||||||||
| IntEnz | IntEnz view | ||||||||
| BRENDA | BRENDA entry | ||||||||
| ExPASy | NiceZyme view | ||||||||
| KEGG | KEGG entry | ||||||||
| MetaCyc | metabolic pathway | ||||||||
| PRIAM | profile | ||||||||
| PDB structures | RCSB PDB PDBe PDBsum | ||||||||
| Gene Ontology | AmiGO / QuickGO | ||||||||
| |||||||||
| Succinate dehydrogenase | |
|---|---|
| Identifiers | |
| Symbol | Respiratory complex II |
| OPM superfamily | 3 |
| OPM protein | 1zoy |
| Membranome | 656 |
Succinate dehydrogenase (SDH) or succinate-coenzyme Q reductase (SQR) or respiratory complex II is an enzyme complex, found in many bacterial cells and in the inner mitochondrial membrane of eukaryotes. It is the only enzyme that participates in both the citric acid cycle and oxidative phosphorylation.[1] Histochemical analysis showing high succinate dehydrogenase in muscle demonstrates high mitochondrial content and high oxidative potential.[2]
In step 6 of the citric acid cycle, SQR catalyzes the oxidation of succinate to fumarate with the reduction of ubiquinone to ubiquinol. This occurs in the inner mitochondrial membrane by coupling the two reactions together.
Structure
[edit]
Subunits
[edit]Mitochondrial and many bacterial SQRs are composed of four structurally different subunits: two hydrophilic and two hydrophobic. The first two subunits, a flavoprotein (SDHA) and an iron-sulfur protein (SDHB), form a hydrophilic head where enzymatic activity of the complex takes place. SDHA contains a covalently attached flavin adenine dinucleotide (FAD) cofactor and the succinate binding site and SDHB contains three iron-sulfur clusters: [2Fe-2S], [4Fe-4S], and [3Fe-4S]. The second two subunits are hydrophobic membrane anchor subunits, SDHC and SDHD. Human mitochondria contain two distinct isoforms of SDHA (Fp subunits type I and type II), these isoforms are also found in Ascaris suum and Caenorhabditis elegans.[3] The subunits form a membrane-bound cytochrome b complex with six transmembrane helices containing one heme b group and a ubiquinone-binding site. Two phospholipid molecules, one cardiolipin and one phosphatidylethanolamine, are also found in the SDHC and SDHD subunits (not shown in the image). They serve to occupy the hydrophobic space below the heme b. These subunits are displayed in the attached image. SDHA is green, SDHB is teal, SDHC is fuchsia, and SDHD is yellow. Around SDHC and SDHD is a phospholipid membrane with the intermembrane space at the top of the image.[4]
| No. | Subunit name | Human protein | Protein description from UniProt | Pfam family with Human protein |
|---|---|---|---|---|
| 1 | SDHA | SDHA_HUMAN | Succinate dehydrogenase [ubiquinone] flavoprotein subunit, mitochondrial | Pfam PF00890, Pfam PF02910 |
| 2 | SDHB | SDHB_HUMAN | Succinate dehydrogenase [ubiquinone] iron-sulfur subunit, mitochondrial | Pfam PF13085, Pfam PF13183 |
| 3 | SDHC | C560_HUMAN | Succinate dehydrogenase cytochrome b560 subunit, mitochondrial | Pfam PF01127 |
| 4 | SDHD | DHSD_HUMAN | Succinate dehydrogenase [ubiquinone] cytochrome b small subunit, mitochondrial | Pfam PF05328 |
Ubiquinone binding site
[edit]Two distinctive ubiquinone binding sites can be recognized on mammalian SDH – matrix-proximal QP and matrix-distal QD. Ubiquinone binding site Qp, which shows higher affinity to ubiquinone, is located in a gap composed of SDHB, SDHC, and SDHD. Ubiquinone is stabilized by the side chains of His207 of subunit B, Ser27 and Arg31 of subunit C, and Tyr83 of subunit D. The quinone ring is surrounded by Ile28 of subunit C and Pro160 of subunit B. These residues, along with Il209, Trp163, and Trp164 of subunit B, and Ser27 (C atom) of subunit C, form the hydrophobic environment of the quinone-binding pocket Qp.[6] In contrast, ubiquinone binding site QD, which lies closer to inter-membrane space, is composed of SDHD only and has lower affinity to ubiquinone.[7]
Succinate binding site
[edit]SDHA provides the binding site for the oxidation of succinate. The side chains Thr254, His354, and Arg399 of subunit A stabilize the molecule while FAD oxidizes and carries the electrons to the first of the iron-sulfur clusters, [2Fe-2S].[8] This can be seen in image 5.
Redox centers
[edit]The succinate-binding site and ubiquinone-binding site are connected by a chain of redox centers including FAD and the iron-sulfur clusters. This chain extends over 40 Å through the enzyme monomer. All edge-to-edge distances between the centers are less than the suggested 14 Å limit for physiological electron transfer.[4] This electron transfer is demonstrated in image 8.
Assembly and maturation
[edit]All subunits of human mitochondrial SDH are nuclear encoded. After translation, SDHA subunit is translocated as apoprotein into the mitochondrial matrix. Subsequently, one of the first steps is covalent attachment of the FAD cofactor (covalent flavinylation). This process is enhanced by succinate dehydrogenase assembly factor 2 (SDHAF2;[9] also called SDH5 in yeast and SDHE in bacteria) and by some of the Krebs cycle intermediates. Fumarate most strongly stimulates covalent flavinylation of SDHA.[10] Through studies of the bacterial system, the mechanism of FAD attachment has been shown to involve a quinone:methide intermediate.[11] In mitochondrial, but not bacterial, assembly, SDHA interacts with a second assembly factor called succinate dehydrogenase assembly factor 4 (SDHAF4; called SDH8 in yeast) before it is inserted into the final complex.[7]
Fe-S prosthetic groups of the subunit SDHB are being preformed in the mitochondrial matrix by protein complex ISU. The complex is also thought to be capable of inserting the iron-sulphur clusters in SDHB during its maturation. The studies suggest that Fe-S cluster insertion precedes SDHA-SDHB dimer forming. Such incorporation requires reduction of cysteine residues within active site of SDHB. Both reduced cysteine residues and already incorporated Fe-S clusters are highly susceptible to ROS damage. Two more SDH assembly factors, SDHAF1 (SDH6) and SDHAF3 (SDH7 in yeast), seem to be involved in SDHB maturation in way of protecting the subunit or dimer SDHA-SDHB from Fe-S cluster damage caused by ROS.[7]
Assembly of the hydrophobic anchor consisting of subunits SDHC and SDHD remains unclear. Especially in case of heme b insertion and even its function. Heme b prosthetic group does not appear to be part of electron transporting pathway within the complex II.[5] The cofactor rather maintains the anchor stability.
Mechanism
[edit]

Succinate oxidation
[edit]Much is known about the succinate oxidation mechanism, which involves the transfer of a proton and a hydride. A combination of mutagenesis and structural analysis identifies Arg-286 of the SDHA subunit (E. coli numbering) as the proton shuttle. Crystal structures of the enzymes from multiple organisms shows that this is well poised for the proton transfer step. Thereafter, there are two possible elimination mechanisms: E2 or E1cb. In the E2 elimination, the mechanism is concerted. The basic residue or cofactor deprotonates the alpha carbon, and FAD accepts the hydride from the beta carbon, oxidizing the bound succinate to fumarate—refer to image 6. In E1cb, an enolate intermediate is formed, shown in image 7, before FAD accepts the hydride. Further research is required to determine which elimination mechanism succinate undergoes in Succinate Dehydrogenase. Oxidized fumarate, now loosely bound to the active site, is free to exit the protein.
Electron tunneling
[edit]After the electrons are derived from succinate oxidation via FAD, they tunnel along the [Fe-S] relay until they reach the [3Fe-4S] cluster. These electrons are subsequently transferred to an awaiting ubiquinone molecule within the active site. The Iron-Sulfur electron tunneling system is shown in image 9.
Ubiquinone reduction
[edit]

The O1 carbonyl oxygen of ubiquinone is oriented at the active site (image 4) by hydrogen bond interactions with Tyr83 of subunit D. The presence of electrons in the [3Fe-4S] iron sulphur cluster induces the movement of ubiquinone into a second orientation. This facilitates a second hydrogen bond interaction between the O4 carbonyl group of ubiquinone and Ser27 of subunit C. Following the first single electron reduction step, a semiquinone radical species is formed. The second electron arrives from the [3Fe-4S] cluster to provide full reduction of the ubiquinone to ubiquinol. This mechanism of the ubiquinone reduction is shown in image 8.
Heme prosthetic group
[edit]Although the functionality of the heme in succinate dehydrogenase is still being researched, some studies[by whom?] have asserted that the first electron delivered to ubiquinone via [3Fe-4S] may tunnel back and forth between the heme and the ubiquinone intermediate. In this way, the heme cofactor acts as an electron sink. Its role is to prevent the interaction of the intermediate with molecular oxygen to produce reactive oxygen species (ROS). The heme group, relative to ubiquinone, is shown in image 4.
It has also been proposed that a gating mechanism may be in place to prevent the electrons from tunneling directly to the heme from the [3Fe-4S] cluster. A potential candidate is residue His207, which lies directly between the cluster and the heme. His207 of subunit B is in direct proximity to the [3Fe-4S] cluster, the bound ubiquinone, and the heme; and could modulate electron flow between these redox centers.[12]
Proton transfer
[edit]To fully reduce the quinone in SQR, two electrons as well as two protons are needed. It has been argued that a water molecule (HOH39) arrives at the active site and is coordinated by His207 of subunit B, Arg31 of subunit C, and Asp82 of subunit D. The semiquinone species is protonated by protons delivered from HOH39, completing the ubiquinone reduction to ubiquinol. His207 and Asp82 most likely facilitate this process. Other studies claim that Tyr83 of subunit D is coordinated to a nearby histidine as well as the O1 carbonyl oxygen of ubiquinone. The histidine residue decreases the pKa of tyrosine, making it more suitable to donate its proton to the reduced ubiquinone intermediate.
Inhibitors
[edit]There are two distinct classes of inhibitors (SDHIs) of complex II: those that bind in the succinate pocket and those that bind in the ubiquinone pocket. Ubiquinone type inhibitors include carboxin and thenoyltrifluoroacetone. Succinate-analogue inhibitors include the synthetic compound malonate as well as the TCA cycle intermediates, malate and oxaloacetate. Indeed, oxaloacetate is one of the most potent inhibitors of Complex II. Why a common TCA cycle intermediate would inhibit Complex II is not entirely understood, though it may exert a protective role in minimizing reverse-electron transfer mediated production of superoxide by Complex I.[13] Atpenin 5a are highly potent Complex II inhibitors mimicking ubiquinone binding.
Ubiquinone type inhibitors have been used as fungicides in agriculture since the 1960s. Carboxin was mainly used to control disease caused by basidiomycetes such as stem rusts and Rhizoctonia diseases. In the 1980s simple benzanilides were found to have comparable activity to carboxin and a number of these were marketed, including benodanil, flutolanil and mepronil.[14] More recently, other compounds with a broader spectrum against a range of plant pathogens have been developed including boscalid, fluopyram, fluxapyroxad, pydiflumetofen and sedaxane.[15][14] Some agriculturally important fungi are not sensitive towards members of the new generation of ubiquinone type inhibitors.[16]
FRAC has a working group[17] for SDHIs and recommends resistance management practices.[18]
Role in disease
[edit]The fundamental role of succinate-coenzyme Q reductase in both oxidative phosphorylation and the citric acid cycle makes it vital in all eukaryotic organisms. Loss of function of SDH via mutations or through toxins can cause a wide range of disease.
When SDH is dysfunctional in the citric acid cycle, it can lead to a buildup of the oncometabolite succinate, which can lead to tumorogenesis. This is well-known to occur in chromaffin cells, causing neuroendocrine tumors such as paraganglioma, renal carcinoma, and Gastrointestinal stromal tumor (GISTs). [19] The penetrance data for SDH mutations causing tumorigenesis is lacking, and international guidelines suggest thorough screening for any carriers. [20] The penetrance of paraganglioma in loss of function mutations of SDH is incomplete and varies by subunit. SDHB mutations have a penetrance between 8% and 37%, SDHD mutations have a penetrance between 38% and 64% with some maternal imprinting effects, and the penetrance for both SDHA and SDHC mutations are poorly studied, but likely between 1% and 30%.[21][22] Mammalian SDH functions not only in energy generation, but also has a role in oxygen sensing. Buildup of succinate due to defective SDH can cause a pseudo-hypoxia and angiogenesis, both of which contribute to the distinctly vascular and characteristic "salt and pepper" appearance of paraganglioma on imaging. [23]
Bi-Allelic loss of function mutations of SDHA, SDHB, SDHD, and SDHAF1 or monoallelic loss of function mutations of SDHA can cause Mitochondrial complex II deficiency. This disruption in oxidative phosphorylation can lead to Leigh syndrome, mitochondrial encephalopathy, optic atrophy, myopathy, and a spectrum of disease. These presentations can range from death within the first year of life or in utero to mild symptoms beginning as an adult.[24]
Reduced levels SDH are observed post mortem in the brains of patients with Huntington's disease, and energy metabolism defects have been identified in both presymptomatic and symptomatic HD patients.[25]
See also
[edit]References
[edit]- ^ Oyedotun KS, Lemire BD (March 2004). "The quaternary structure of the Saccharomyces cerevisiae succinate dehydrogenase. Homology modeling, cofactor docking, and molecular dynamics simulation studies". The Journal of Biological Chemistry. 279 (10): 9424–9431. doi:10.1074/jbc.M311876200. PMID 14672929.
- ^ webmaster (2009-03-04). "Using Histochemistry to Determine Muscle Properties". Succinate Dehydrogenase: Identifying Oxidative Potential. University of California, San Diego. Archived from the original on 2018-10-10. Retrieved 2017-12-27.
- ^ Tomitsuka E, Hirawake H, Goto Y, Taniwaki M, Harada S, Kita K (August 2003). "Direct evidence for two distinct forms of the flavoprotein subunit of human mitochondrial complex II (succinate-ubiquinone reductase)". Journal of Biochemistry. 134 (2): 191–195. doi:10.1093/jb/mvg144. PMID 12966066.
- ^ a b Yankovskaya V, Horsefield R, Törnroth S, Luna-Chavez C, Miyoshi H, Léger C, et al. (January 2003). "Architecture of succinate dehydrogenase and reactive oxygen species generation". Science. 299 (5607): 700–704. Bibcode:2003Sci...299..700Y. doi:10.1126/science.1079605. PMID 12560550. S2CID 29222766.
- ^ a b Sun F, Huo X, Zhai Y, Wang A, Xu J, Su D, et al. (July 2005). "Crystal structure of mitochondrial respiratory membrane protein complex II". Cell. 121 (7): 1043–1057. doi:10.1016/j.cell.2005.05.025. PMID 15989954.
- ^ Horsefield R, Yankovskaya V, Sexton G, Whittingham W, Shiomi K, Omura S, et al. (March 2006). "Structural and computational analysis of the quinone-binding site of complex II (succinate-ubiquinone oxidoreductase): a mechanism of electron transfer and proton conduction during ubiquinone reduction". The Journal of Biological Chemistry. 281 (11): 7309–7316. doi:10.1074/jbc.M508173200. PMID 16407191.
- ^ a b c Van Vranken JG, Na U, Winge DR, Rutter J (December 2014). "Protein-mediated assembly of succinate dehydrogenase and its cofactors". Critical Reviews in Biochemistry and Molecular Biology. 50 (2): 168–180. doi:10.3109/10409238.2014.990556. PMC 4653115. PMID 25488574.
- ^ Kenney WC (April 1975). "The reaction of N-ethylmaleimide at the active site of succinate dehydrogenase". The Journal of Biological Chemistry. 250 (8): 3089–3094. doi:10.1016/S0021-9258(19)41598-6. PMID 235539.
- ^ Sharma P, Maklashina E, Cecchini G, Iverson TM (September 2020). "The roles of SDHAF2 and dicarboxylate in covalent flavinylation of SDHA, the human complex II flavoprotein". Proceedings of the National Academy of Sciences of the United States of America. 117 (38): 23548–23556. Bibcode:2020PNAS..11723548S. doi:10.1073/pnas.2007391117. PMC 7519310. PMID 32887801.
- ^ Maklashina E, Iverson TM, Cecchini G (October 2022). "How an assembly factor enhances covalent FAD attachment to the flavoprotein subunit of complex II". The Journal of Biological Chemistry. 298 (10): 102472. doi:10.1016/j.jbc.2022.102472. PMC 9557727. PMID 36089066.
- ^ Sharma P, Maklashina E, Cecchini G, Iverson TM (January 2018). "Crystal structure of an assembly intermediate of respiratory Complex II". Nature Communications. 9 (1): 274. Bibcode:2018NatCo...9..274S. doi:10.1038/s41467-017-02713-8. PMC 5773532. PMID 29348404.
- ^ Tran QM, Rothery RA, Maklashina E, Cecchini G, Weiner JH (October 2006). "The quinone binding site in Escherichia coli succinate dehydrogenase is required for electron transfer to the heme b". The Journal of Biological Chemistry. 281 (43): 32310–32317. doi:10.1074/jbc.M607476200. PMID 16950775.
- ^ Muller FL, Liu Y, Abdul-Ghani MA, Lustgarten MS, Bhattacharya A, Jang YC, et al. (January 2008). "High rates of superoxide production in skeletal-muscle mitochondria respiring on both complex I- and complex II-linked substrates". The Biochemical Journal. 409 (2): 491–499. doi:10.1042/BJ20071162. PMID 17916065.
- ^ a b Walter H (2016). "Fungicidal Succinate-Dehydrogenase-Inhibiting Carboxamides". Bioactive Carboxylic Compound Classes: Pharmaceuticals and Agrochemicals. pp. 405–425. doi:10.1002/9783527693931.ch31. ISBN 978-3-527-33947-1.
- ^ Avenot HF, Michailides TJ (2010). "Progress in understanding molecular mechanisms and evolution of resistance to succinate dehydrogenase inhibiting (SDHI) fungicides in phytopathogenic fungi". Crop Protection. 29 (7): 643–651. Bibcode:2010CrPro..29..643A. doi:10.1016/j.cropro.2010.02.019.
- ^
- Lucas JA, Hawkins NJ, Fraaije BA (2015). The Evolution of Fungicide Resistance. Advances in Applied Microbiology. Vol. 90. pp. 29–92. doi:10.1016/bs.aambs.2014.09.001. ISBN 978-0-12-802275-7. PMID 25596029.
- Dubos T, Pasquali M, Pogoda F, Casanova A, Hoffmann L, Beyer M (January 2013). "Differences between the succinate dehydrogenase sequences of isopyrazam sensitive Zymoseptoria tritici and insensitive Fusarium graminearum strains". Pesticide Biochemistry and Physiology. 105 (1): 28–35. Bibcode:2013PBioP.105...28D. doi:10.1016/j.pestbp.2012.11.004. PMID 24238287.
- ^ "SDHI Fungicides Working Group". FRAC (Fungicide Resistance Action Committee). 2020-01-31. Retrieved 2022-07-05.
- ^ "Recommendations for SDHI". FRAC. March 2020. Retrieved 2022-07-05.
- ^ Barletta JA, Hornick JL (July 2012). "Succinate dehydrogenase-deficient tumors: diagnostic advances and clinical implications". Advances in Anatomic Pathology. 19 (4): 193–203. doi:10.1097/PAP.0b013e31825c6bc6. PMID 22692282. S2CID 32088940.
- ^ Amar L, Pacak K, Steichen O, Akker SA, Aylwin SJ, Baudin E, et al. (July 2021). "International consensus on initial screening and follow-up of asymptomatic SDHx mutation carriers". Nature Reviews Endocrinology. 17 (7): 435–444. doi:10.1038/s41574-021-00492-3.
- ^ Rijken JA, Niemeijer ND, Jonker MA, Eijkelenkamp K, Jansen JC, van Berkel A, et al. (January 2018). "The penetrance of paraganglioma and pheochromocytoma in SDHB germline mutation carriers". Clinical Genetics. 93 (1): 60–66. doi:10.1111/cge.13055. PMID 28503760.
- ^ Baysal BE (May 2013). "Mitochondrial complex II and genomic imprinting in inheritance of paraganglioma tumors". Biochimica et Biophysica Acta (BBA) - Bioenergetics. 1827 (5): 573–577. doi:10.1016/j.bbabio.2012.12.005. PMID 23291190.
- ^ https://radiopaedia.org/articles/salt-and-pepper-sign-paraganglioma-1?lang=us
- ^ Fullerton M, McFarland R, Taylor RW, Alston CL (September 2020). "The genetic basis of isolated mitochondrial complex II deficiency". Molecular Genetics and Metabolism. 131 (1–2): 53–65. doi:10.1016/j.ymgme.2020.09.009. PMC 7758838. PMID 33162331.
- ^ Skillings EA, Morton AJ (2016). "Delayed Onset and Reduced Cognitive Deficits through Pre-Conditioning with 3-Nitropropionic Acid is Dependent on Sex and CAG Repeat Length in the R6/2 Mouse Model of Huntington's Disease". Journal of Huntington's Disease. 5 (1): 19–32. doi:10.3233/JHD-160189. PMID 27031731.
